Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
14.66 | LDD0402 | [1] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [2] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [2] | |
|
TH211 Probe Info |
 |
Y60(20.00) | LDD0257 | [3] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [4] | |
|
STPyne Probe Info |
 |
K280(10.00); K39(6.20); K412(7.34); K48(5.56) | LDD0277 | [5] | |
|
DBIA Probe Info |
 |
C589(1.99); C608(2.64); C27(1.89) | LDD3311 | [6] | |
|
BTD Probe Info |
 |
C506(0.83); C525(0.68) | LDD2089 | [7] | |
|
Sulforaphane-probe2 Probe Info |
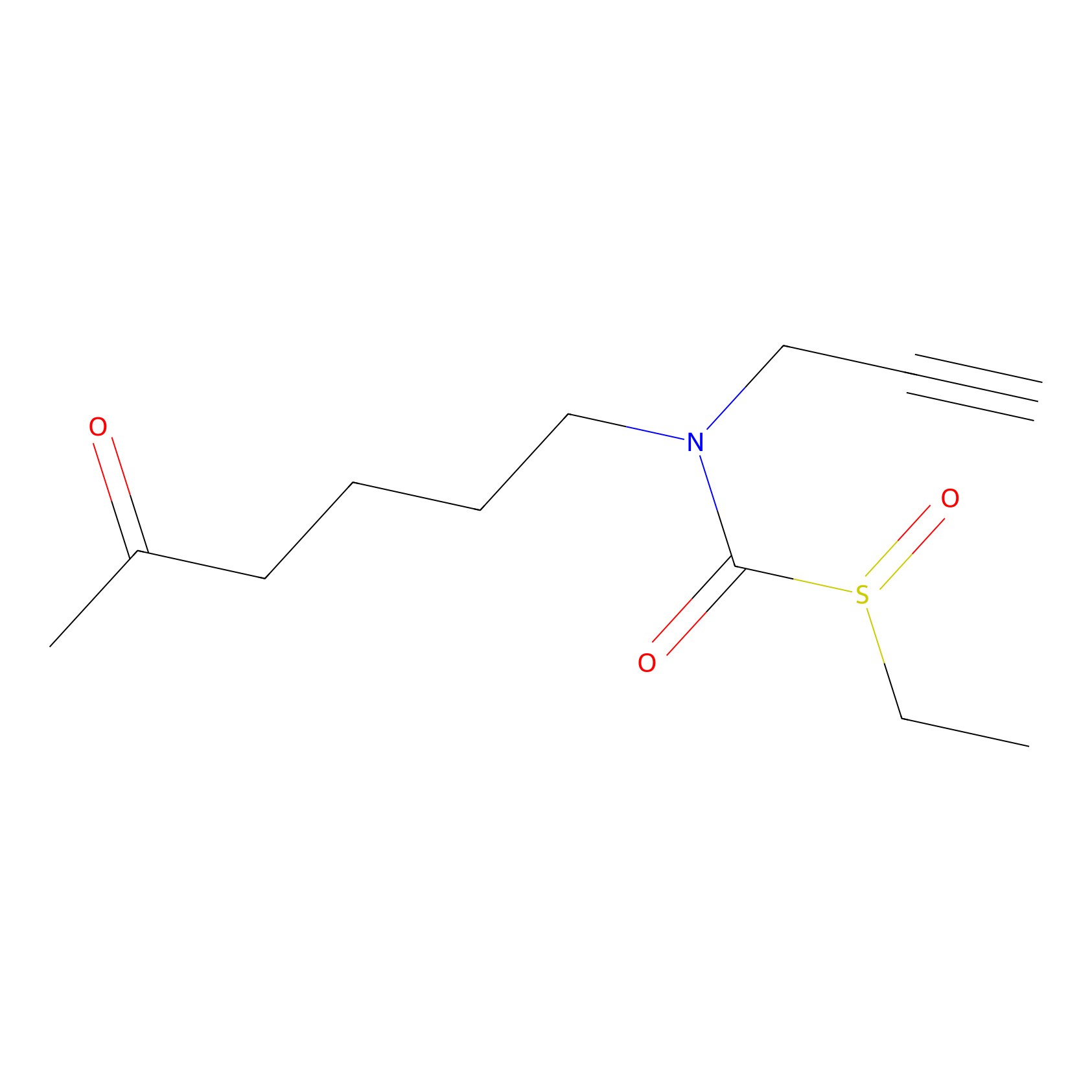 |
1.72 | LDD0160 | [8] | |
|
AHL-Pu-1 Probe Info |
 |
C335(2.41) | LDD0170 | [9] | |
|
HHS-482 Probe Info |
 |
Y53(1.20) | LDD0285 | [10] | |
|
HHS-465 Probe Info |
 |
Y60(10.00) | LDD2237 | [11] | |
|
ATP probe Probe Info |
 |
K148(0.00); K149(0.00); K503(0.00) | LDD0199 | [12] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C506(0.00); C525(0.00); C335(0.00); C204(0.00) | LDD0038 | [13] | |
|
IA-alkyne Probe Info |
 |
C506(0.00); C525(0.00); C335(0.00); C204(0.00) | LDD0036 | [13] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [14] | |
|
Lodoacetamide azide Probe Info |
 |
C506(0.00); C525(0.00); C335(0.00); C204(0.00) | LDD0037 | [13] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [15] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [15] | |
|
Compound 10 Probe Info |
 |
C27(0.00); C335(0.00) | LDD2216 | [16] | |
|
Compound 11 Probe Info |
 |
N.A. | LDD2213 | [16] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [15] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [15] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [15] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [15] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [17] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [18] | |
|
Acrolein Probe Info |
 |
C182(0.00); C405(0.00); C506(0.00); H157(0.00) | LDD0217 | [19] | |
|
Methacrolein Probe Info |
 |
C506(0.00); C27(0.00) | LDD0218 | [19] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [20] | |
|
AOyne Probe Info |
 |
10.90 | LDD0443 | [21] | |
|
NAIA_5 Probe Info |
 |
C525(0.00); C335(0.00); C92(0.00); C506(0.00) | LDD2223 | [22] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C056 Probe Info |
 |
16.11 | LDD1753 | [23] | |
|
C072 Probe Info |
 |
22.01 | LDD1768 | [23] | |
|
C112 Probe Info |
 |
18.25 | LDD1799 | [23] | |
|
C161 Probe Info |
 |
10.13 | LDD1841 | [23] | |
|
C210 Probe Info |
 |
51.98 | LDD1884 | [23] | |
|
FFF probe3 Probe Info |
 |
6.79 | LDD0465 | [24] | |
|
FFF probe4 Probe Info |
 |
18.26 | LDD0466 | [24] | |
|
JN0003 Probe Info |
 |
12.02 | LDD0469 | [24] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0139 | [25] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0070 | [26] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0068 | [27] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C525(0.75) | LDD2130 | [7] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C525(0.91) | LDD2117 | [7] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C506(2.24); C525(0.96) | LDD2152 | [7] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C335(0.73) | LDD2132 | [7] |
| LDCM0025 | 4SU-RNA | DM93 | C335(2.41) | LDD0170 | [9] |
| LDCM0545 | Acetamide | MDA-MB-231 | C525(0.70) | LDD2138 | [7] |
| LDCM0156 | Aniline | NCI-H1299 | C27(0.00); C204(0.00) | LDD0404 | [1] |
| LDCM0108 | Chloroacetamide | HeLa | H157(0.00); C506(0.00); C405(0.00) | LDD0222 | [19] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C27(2.28); C182(1.28) | LDD1702 | [7] |
| LDCM0625 | F8 | Ramos | C27(1.53) | LDD2187 | [28] |
| LDCM0572 | Fragment10 | Ramos | C27(0.89) | LDD2189 | [28] |
| LDCM0573 | Fragment11 | Ramos | C27(0.09) | LDD2190 | [28] |
| LDCM0574 | Fragment12 | Ramos | C27(0.22) | LDD2191 | [28] |
| LDCM0575 | Fragment13 | Ramos | C27(0.52) | LDD2192 | [28] |
| LDCM0576 | Fragment14 | Ramos | C27(0.73) | LDD2193 | [28] |
| LDCM0579 | Fragment20 | Ramos | C27(0.33) | LDD2194 | [28] |
| LDCM0580 | Fragment21 | Ramos | C27(0.77) | LDD2195 | [28] |
| LDCM0582 | Fragment23 | Ramos | C27(0.24) | LDD2196 | [28] |
| LDCM0578 | Fragment27 | Ramos | C27(0.57) | LDD2197 | [28] |
| LDCM0586 | Fragment28 | Ramos | C27(0.30) | LDD2198 | [28] |
| LDCM0588 | Fragment30 | Ramos | C27(0.67) | LDD2199 | [28] |
| LDCM0589 | Fragment31 | Ramos | C27(0.43) | LDD2200 | [28] |
| LDCM0590 | Fragment32 | Ramos | C27(0.29) | LDD2201 | [28] |
| LDCM0468 | Fragment33 | Ramos | C27(1.17) | LDD2202 | [28] |
| LDCM0596 | Fragment38 | Ramos | C27(0.46) | LDD2203 | [28] |
| LDCM0566 | Fragment4 | Ramos | C27(0.74) | LDD2184 | [28] |
| LDCM0610 | Fragment52 | Ramos | C27(0.52) | LDD2204 | [28] |
| LDCM0614 | Fragment56 | Ramos | C27(0.79) | LDD2205 | [28] |
| LDCM0569 | Fragment7 | Ramos | C27(0.85) | LDD2186 | [28] |
| LDCM0571 | Fragment9 | Ramos | C27(0.34) | LDD2188 | [28] |
| LDCM0107 | IAA | HeLa | H157(0.00); C506(0.00) | LDD0221 | [19] |
| LDCM0123 | JWB131 | DM93 | Y53(1.20) | LDD0285 | [10] |
| LDCM0124 | JWB142 | DM93 | Y53(0.50) | LDD0286 | [10] |
| LDCM0125 | JWB146 | DM93 | Y53(2.12) | LDD0287 | [10] |
| LDCM0126 | JWB150 | DM93 | Y53(1.59) | LDD0288 | [10] |
| LDCM0127 | JWB152 | DM93 | Y53(1.16) | LDD0289 | [10] |
| LDCM0128 | JWB198 | DM93 | Y53(0.74) | LDD0290 | [10] |
| LDCM0130 | JWB211 | DM93 | Y53(0.77) | LDD0292 | [10] |
| LDCM0022 | KB02 | HEK-293T | C27(1.05); C405(0.97); C389(1.10); C335(0.87) | LDD1492 | [29] |
| LDCM0023 | KB03 | Jurkat | C525(4.13) | LDD0315 | [30] |
| LDCM0024 | KB05 | G361 | C589(1.99); C608(2.64); C27(1.89) | LDD3311 | [6] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C525(0.57) | LDD2121 | [7] |
| LDCM0109 | NEM | HeLa | H157(0.00); H224(0.00) | LDD0223 | [19] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C506(0.83); C525(0.68) | LDD2089 | [7] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C506(5.44) | LDD2093 | [7] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C525(0.88) | LDD2099 | [7] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C525(0.91) | LDD2107 | [7] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C27(0.70); C525(0.73) | LDD2109 | [7] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C525(1.04) | LDD2111 | [7] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C525(1.67) | LDD2119 | [7] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C525(1.02) | LDD2122 | [7] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C525(0.92) | LDD2123 | [7] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C525(1.29) | LDD2124 | [7] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C525(0.96) | LDD2125 | [7] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C525(1.01) | LDD2127 | [7] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C506(6.03); C27(0.97); C525(0.95) | LDD2129 | [7] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C525(0.96) | LDD2134 | [7] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C27(1.07) | LDD2135 | [7] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C525(1.10) | LDD2136 | [7] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C27(0.99); C525(0.95) | LDD2137 | [7] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C525(0.83) | LDD2140 | [7] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C525(1.67) | LDD2144 | [7] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C525(0.94) | LDD2146 | [7] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C506(0.44); C525(0.56); C335(0.46) | LDD2150 | [7] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C27(0.86); C27(0.85) | LDD2206 | [31] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C27(0.56); C27(0.35) | LDD2207 | [31] |
| LDCM0131 | RA190 | MM1.R | C27(1.15) | LDD0304 | [32] |
| LDCM0003 | Sulforaphane | MDA-MB-231 | 1.72 | LDD0160 | [8] |
| LDCM0112 | W16 | Hep-G2 | C27(0.74) | LDD0239 | [20] |
The Interaction Atlas With This Target
References
