Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
P3 Probe Info |
 |
10.00 | LDD0450 | [2] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [3] | |
|
TH211 Probe Info |
 |
Y97(20.00) | LDD0257 | [4] | |
|
TH214 Probe Info |
 |
Y97(18.78) | LDD0258 | [4] | |
|
TH216 Probe Info |
 |
Y97(20.00) | LDD0259 | [4] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [5] | |
|
ONAyne Probe Info |
 |
N.A. | LDD0273 | [6] | |
|
OPA-S-S-alkyne Probe Info |
 |
K91(4.21) | LDD3494 | [7] | |
|
Probe 1 Probe Info |
 |
Y53(42.28) | LDD3495 | [8] | |
|
DA-P3 Probe Info |
 |
4.46 | LDD0179 | [9] | |
|
HHS-482 Probe Info |
 |
Y53(1.22); Y85(1.63); Y97(0.92) | LDD0285 | [10] | |
|
HHS-475 Probe Info |
 |
Y85(0.79); Y53(0.80); Y97(0.96) | LDD0264 | [11] | |
|
HHS-465 Probe Info |
 |
Y53(0.63); Y85(10.00); Y97(9.45) | LDD2237 | [12] | |
|
Acrolein Probe Info |
 |
H58(0.00); H92(0.00) | LDD0221 | [13] | |
|
5E-2FA Probe Info |
 |
H18(0.00); H75(0.00); H58(0.00) | LDD2235 | [14] | |
|
ATP probe Probe Info |
 |
K78(0.00); K39(0.00) | LDD0199 | [15] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0162 | [16] | |
|
ATP probe Probe Info |
 |
K44(0.00); K30(0.00); K78(0.00) | LDD0035 | [17] | |
|
BTD Probe Info |
 |
N.A. | LDD0004 | [18] | |
|
Dyn-2 Probe Info |
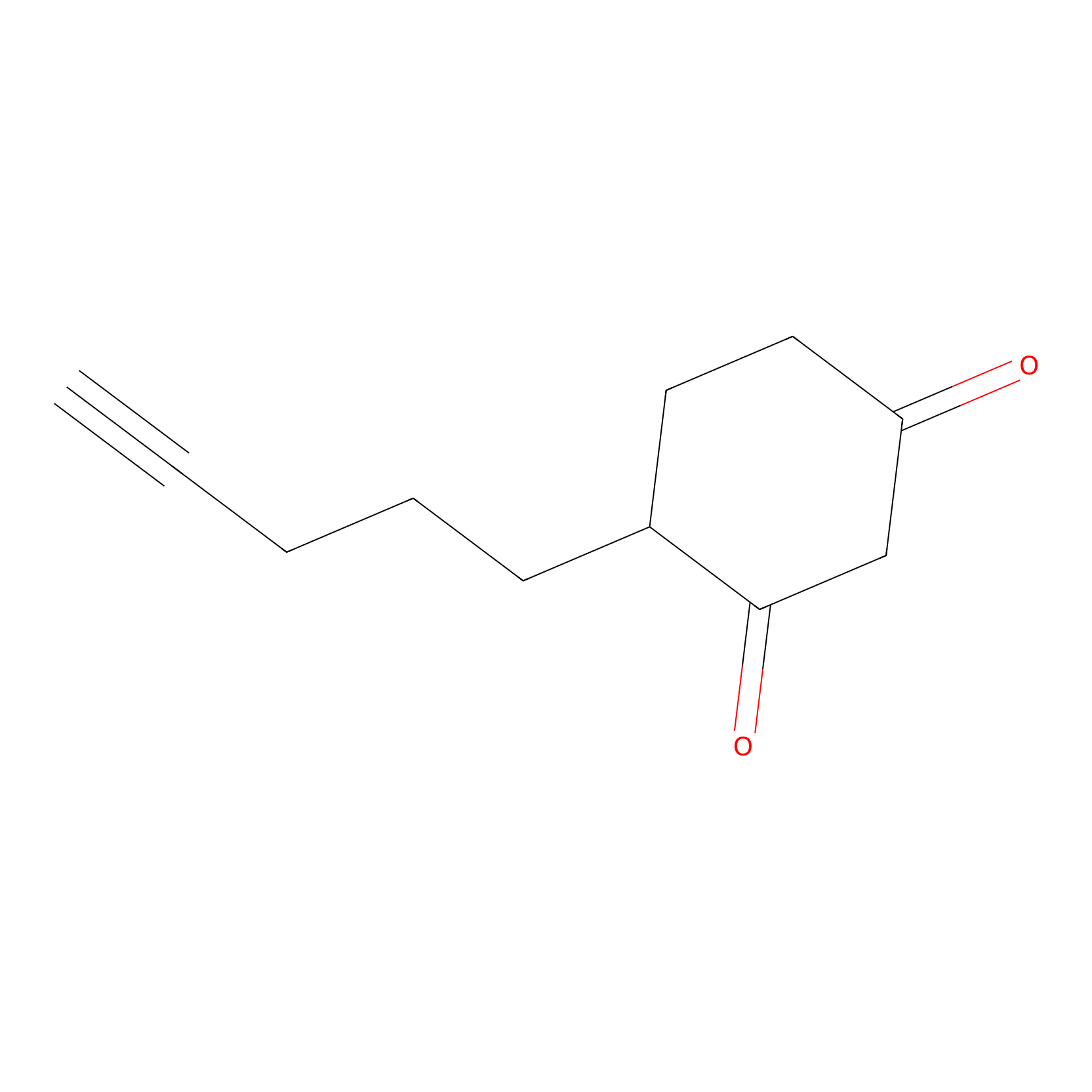 |
N.A. | LDD0013 | [18] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [19] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [20] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [19] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [18] | |
|
aHNE Probe Info |
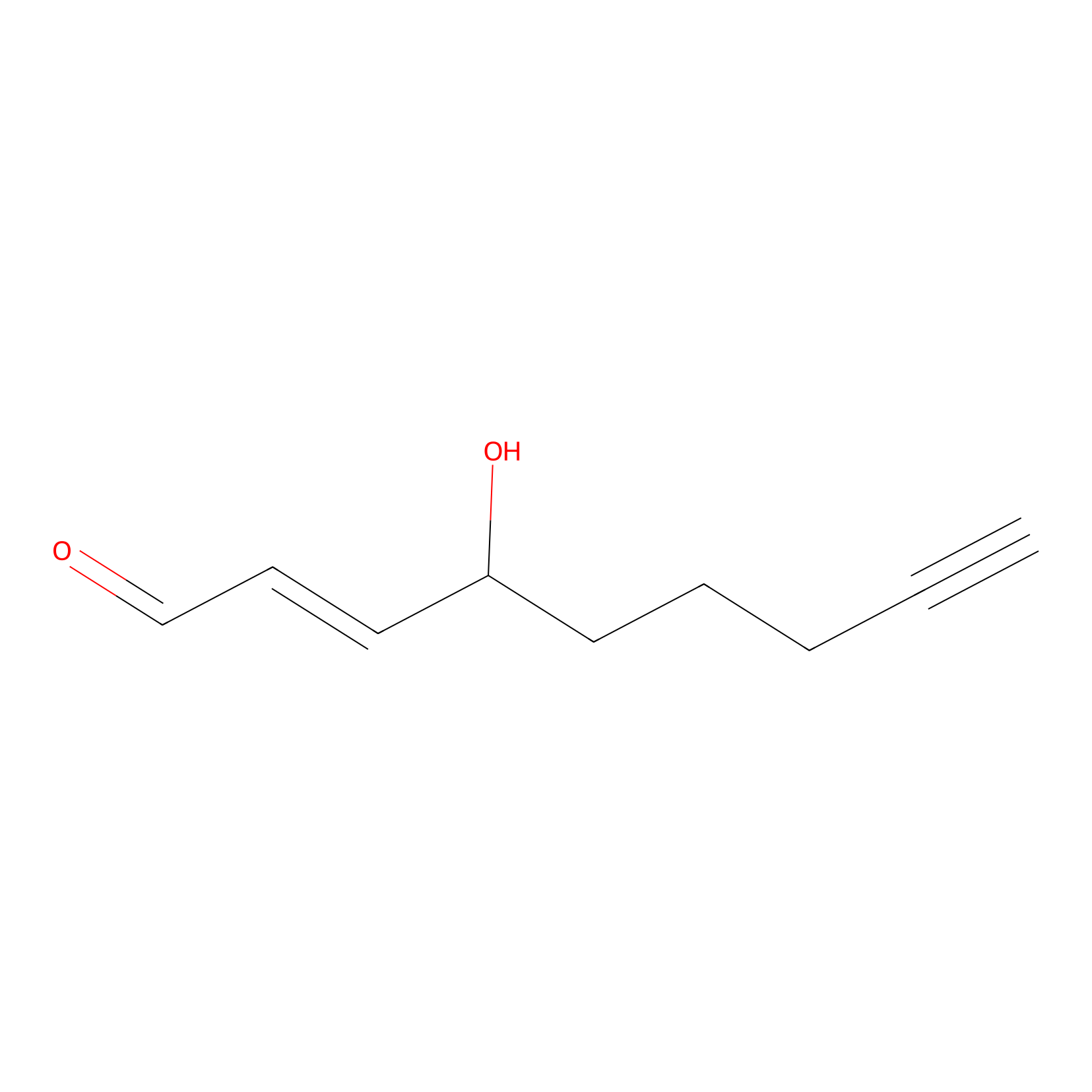 |
N.A. | LDD0001 | [18] | |
|
DBIA Probe Info |
 |
C3(12.13) | LDD2236 | [21] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [18] | |
|
NHS Probe Info |
 |
K78(0.00); K39(0.00) | LDD0010 | [18] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0017 | [22] | |
|
PPMS Probe Info |
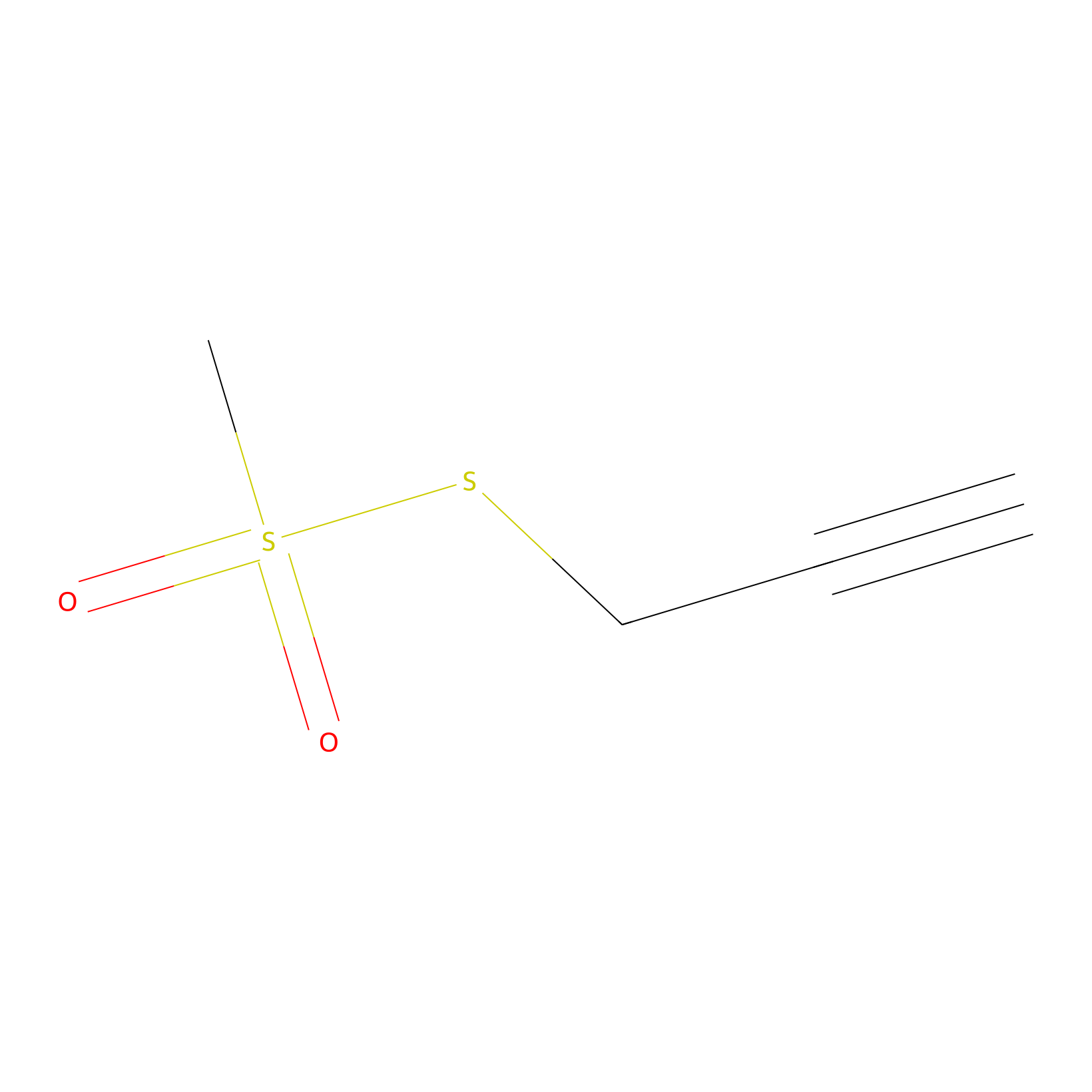 |
N.A. | LDD0008 | [18] | |
|
SF Probe Info |
 |
Y97(0.00); K39(0.00) | LDD0028 | [23] | |
|
STPyne Probe Info |
 |
K39(0.00); K78(0.00) | LDD0009 | [18] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [18] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [24] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [25] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [26] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [20] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C232 Probe Info |
 |
37.27 | LDD1905 | [27] | |
|
FFF probe4 Probe Info |
 |
6.63 | LDD0466 | [28] | |
|
Diazir Probe Info |
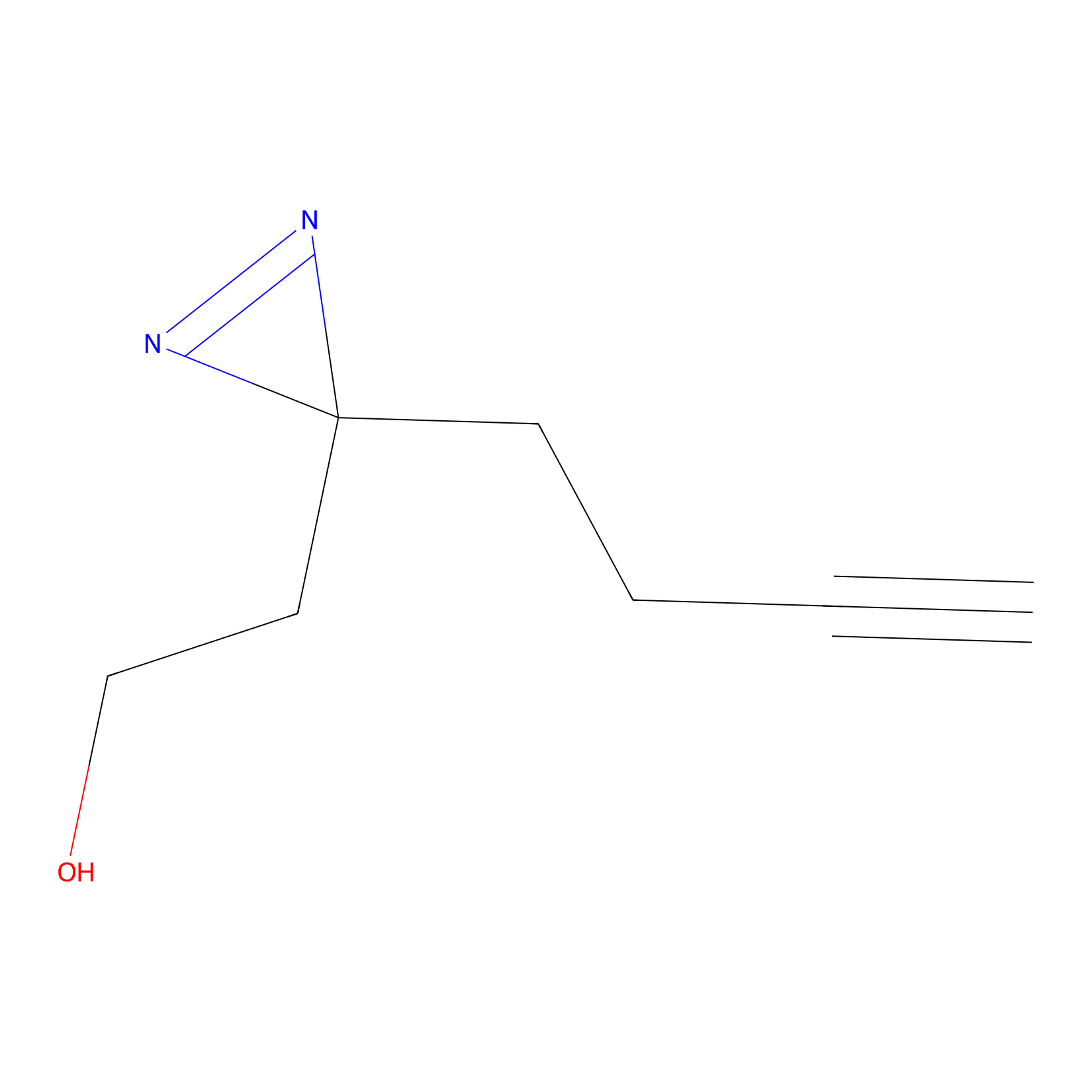 |
N.A. | LDD0011 | [18] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0156 | Aniline | NCI-H1299 | 13.06 | LDD0403 | [1] |
| LDCM0108 | Chloroacetamide | HeLa | H58(0.00); H92(0.00) | LDD0222 | [13] |
| LDCM0632 | CL-Sc | Hep-G2 | C3(1.08); C3(0.80) | LDD2227 | [20] |
| LDCM0634 | CY-0357 | Hep-G2 | C3(0.67) | LDD2228 | [20] |
| LDCM0027 | Dopamine | HEK-293T | 4.46 | LDD0179 | [9] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C3(1.81) | LDD1702 | [29] |
| LDCM0031 | Epigallocatechin gallate | HEK-293T | 14.03 | LDD0183 | [9] |
| LDCM0625 | F8 | Ramos | C3(0.83) | LDD2187 | [30] |
| LDCM0572 | Fragment10 | Ramos | C3(0.89) | LDD2189 | [30] |
| LDCM0574 | Fragment12 | Ramos | C3(0.66) | LDD2191 | [30] |
| LDCM0575 | Fragment13 | Ramos | C3(1.00) | LDD2192 | [30] |
| LDCM0576 | Fragment14 | Ramos | C3(0.58) | LDD2193 | [30] |
| LDCM0579 | Fragment20 | Ramos | C3(0.43) | LDD2194 | [30] |
| LDCM0580 | Fragment21 | Ramos | C3(0.60) | LDD2195 | [30] |
| LDCM0582 | Fragment23 | Ramos | C3(0.79) | LDD2196 | [30] |
| LDCM0578 | Fragment27 | Ramos | C3(0.98) | LDD2197 | [30] |
| LDCM0588 | Fragment30 | Ramos | C3(0.55) | LDD2199 | [30] |
| LDCM0589 | Fragment31 | Ramos | C3(0.83) | LDD2200 | [30] |
| LDCM0590 | Fragment32 | Ramos | C3(0.92) | LDD2201 | [30] |
| LDCM0596 | Fragment38 | Ramos | C3(0.99) | LDD2203 | [30] |
| LDCM0566 | Fragment4 | Ramos | C3(0.66) | LDD2184 | [30] |
| LDCM0614 | Fragment56 | Ramos | C3(0.79) | LDD2205 | [30] |
| LDCM0569 | Fragment7 | Ramos | C3(0.82) | LDD2186 | [30] |
| LDCM0571 | Fragment9 | Ramos | C3(0.57) | LDD2188 | [30] |
| LDCM0116 | HHS-0101 | DM93 | Y85(0.79); Y53(0.80); Y97(0.96) | LDD0264 | [11] |
| LDCM0117 | HHS-0201 | DM93 | Y53(0.74); Y85(0.77); Y97(0.91) | LDD0265 | [11] |
| LDCM0118 | HHS-0301 | DM93 | Y53(0.86); Y97(0.91); Y85(1.07) | LDD0266 | [11] |
| LDCM0119 | HHS-0401 | DM93 | Y53(0.83); Y97(0.97); Y85(1.59) | LDD0267 | [11] |
| LDCM0120 | HHS-0701 | DM93 | Y53(0.90); Y97(0.91); Y85(1.27) | LDD0268 | [11] |
| LDCM0107 | IAA | HeLa | H58(0.00); H92(0.00) | LDD0221 | [13] |
| LDCM0123 | JWB131 | DM93 | Y53(1.22); Y85(1.63); Y97(0.92) | LDD0285 | [10] |
| LDCM0124 | JWB142 | DM93 | Y53(0.79); Y85(0.42); Y97(0.85) | LDD0286 | [10] |
| LDCM0125 | JWB146 | DM93 | Y53(1.06); Y85(1.24); Y97(1.04) | LDD0287 | [10] |
| LDCM0126 | JWB150 | DM93 | Y53(3.92); Y85(2.54); Y97(2.73) | LDD0288 | [10] |
| LDCM0127 | JWB152 | DM93 | Y53(2.34); Y85(3.04); Y97(1.81) | LDD0289 | [10] |
| LDCM0128 | JWB198 | DM93 | Y53(1.18); Y85(1.18); Y97(1.31) | LDD0290 | [10] |
| LDCM0129 | JWB202 | DM93 | Y53(0.67); Y85(0.54); Y97(0.65) | LDD0291 | [10] |
| LDCM0130 | JWB211 | DM93 | Y53(0.89); Y85(0.81); Y97(1.07) | LDD0292 | [10] |
| LDCM0022 | KB02 | Ramos | C3(0.98) | LDD2182 | [30] |
| LDCM0023 | KB03 | MDA-MB-231 | C3(1.64) | LDD1701 | [29] |
| LDCM0024 | KB05 | Ramos | C3(0.84) | LDD2185 | [30] |
| LDCM0030 | Luteolin | HEK-293T | 9.68 | LDD0182 | [9] |
| LDCM0109 | NEM | HeLa | H58(0.00); H66(0.00) | LDD0223 | [13] |
| LDCM0032 | Oleacein | HEK-293T | 4.33 | LDD0184 | [9] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C3(0.48) | LDD2207 | [31] |
The Interaction Atlas With This Target
References
