Details of the Target
General Information of Target
| Target ID | LDTP01699 | |||||
|---|---|---|---|---|---|---|
| Target Name | Acyl-CoA 6-desaturase (FADS2) | |||||
| Gene Name | FADS2 | |||||
| Gene ID | 9415 | |||||
| Synonyms |
Acyl-CoA 6-desaturase; EC 1.14.19.3; Delta(6) fatty acid desaturase; D6D; Delta(6) desaturase; Delta-6 desaturase; Fatty acid desaturase 2 |
|||||
| 3D Structure | ||||||
| Sequence |
MGKGGNQGEGAAEREVSVPTFSWEEIQKHNLRTDRWLVIDRKVYNITKWSIQHPGGQRVI
GHYAGEDATDAFRAFHPDLEFVGKFLKPLLIGELAPEEPSQDHGKNSKITEDFRALRKTA EDMNLFKTNHVFFLLLLAHIIALESIAWFTVFYFGNGWIPTLITAFVLATSQAQAGWLQH DYGHLSVYRKPKWNHLVHKFVIGHLKGASANWWNHRHFQHHAKPNIFHKDPDVNMLHVFV LGEWQPIEYGKKKLKYLPYNHQHEYFFLIGPPLLIPMYFQYQIIMTMIVHKNWVDLAWAV SYYIRFFITYIPFYGILGALLFLNFIRFLESHWFVWVTQMNHIVMEIDQEAYRDWFSSQL TATCNVEQSFFNDWFSGHLNFQIEHHLFPTMPRHNLHKIAPLVKSLCAKHGIEYQEKPLL RALLDIIRSLKKSGKLWLDAYLHK |
|||||
| Target Type |
Literature-reported
|
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Fatty acid desaturase type 1 family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Involved in the biosynthesis of highly unsaturated fatty acids (HUFA) from the essential polyunsaturated fatty acids (PUFA) linoleic acid (LA) (18:2n-6) and alpha-linolenic acid (ALA) (18:3n-3) precursors, acting as a fatty acyl-coenzyme A (CoA) desaturase that introduces a cis double bond at carbon 6 of the fatty acyl chain. Catalyzes the first and rate limiting step in this pathway which is the desaturation of LA (18:2n-6) and ALA (18:3n-3) into gamma-linoleate (GLA) (18:3n-6) and stearidonate (18:4n-3), respectively. Subsequently, in the biosynthetic pathway of HUFA n-3 series, it desaturates tetracosapentaenoate (24:5n-3) to tetracosahexaenoate (24:6n-3), which is then converted to docosahexaenoate (DHA)(22:6n-3), an important lipid for nervous system function. Desaturates hexadecanate (palmitate) to produce 6Z-hexadecenoate (sapienate), a fatty acid unique to humans and major component of human sebum, that has been implicated in the development of acne and may have potent antibacterial activity. It can also desaturate (11E)-octadecenoate (trans-vaccenoate, the predominant trans fatty acid in human milk) at carbon 6 generating (6Z,11E)-octadecadienoate. In addition to Delta-6 activity, this enzyme exhibits Delta-8 activity with slight biases toward n-3 fatty acyl-CoA substrates.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Alkylaryl probe 2 Probe Info |
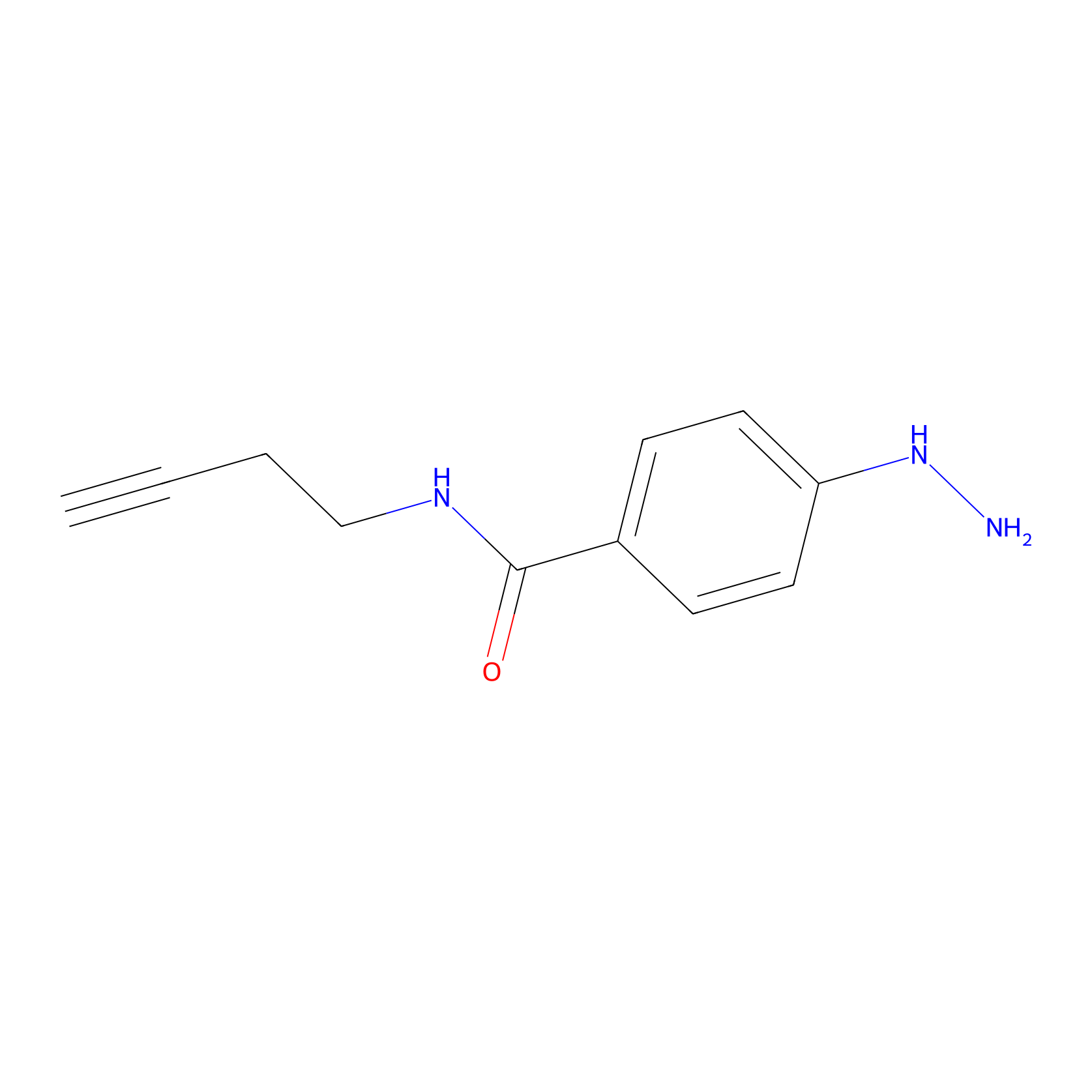 |
20.00 | LDD0389 | [1] | |
|
TH211 Probe Info |
 |
Y441(12.97) | LDD0260 | [2] | |
|
STPyne Probe Info |
 |
K28(6.67); K398(5.44); K404(10.00); K42(1.44) | LDD0277 | [3] | |
|
OPA-S-S-alkyne Probe Info |
 |
K108(1.76) | LDD3494 | [4] | |
|
P11 Probe Info |
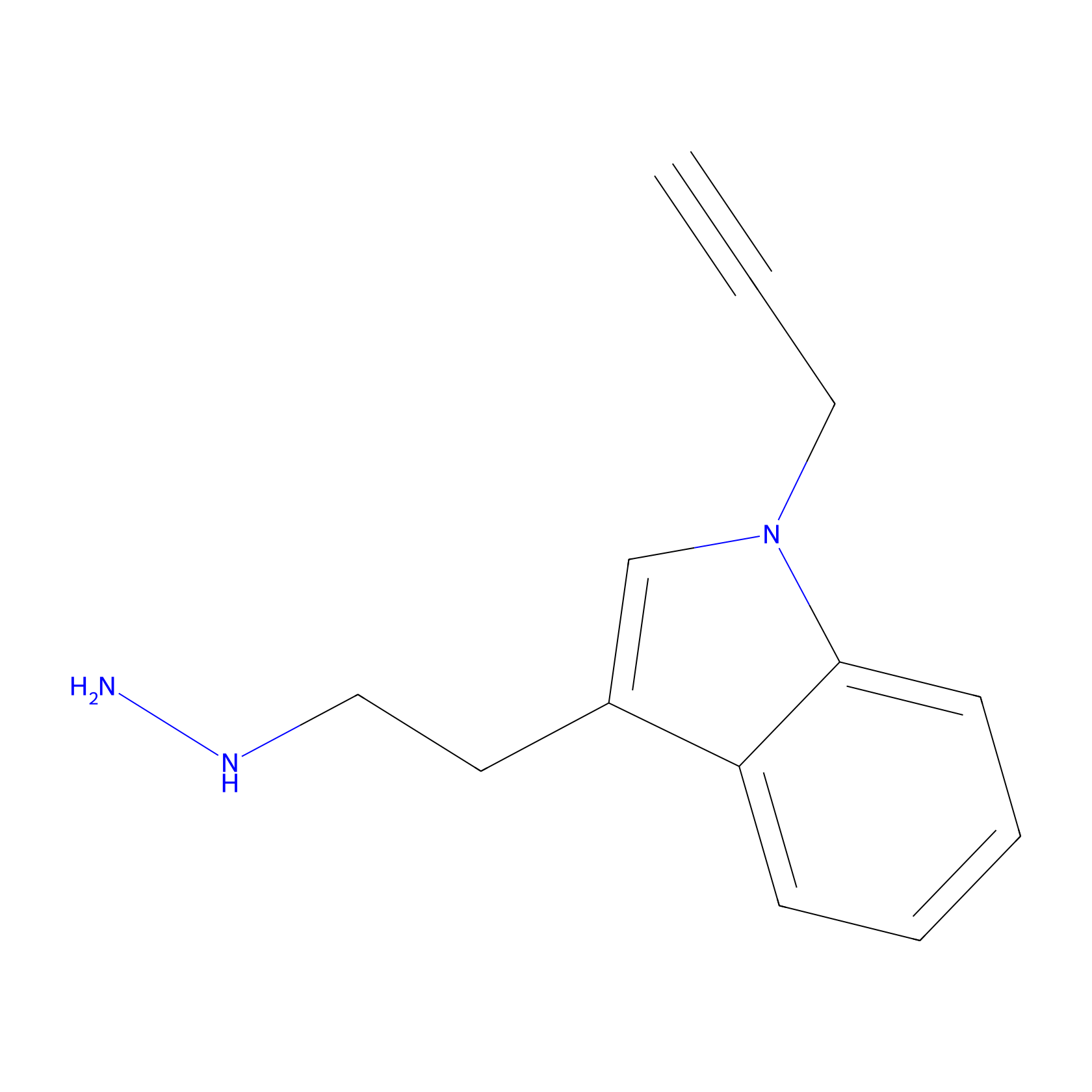 |
12.38 | LDD0201 | [5] | |
|
Alkylaryl probe 3 Probe Info |
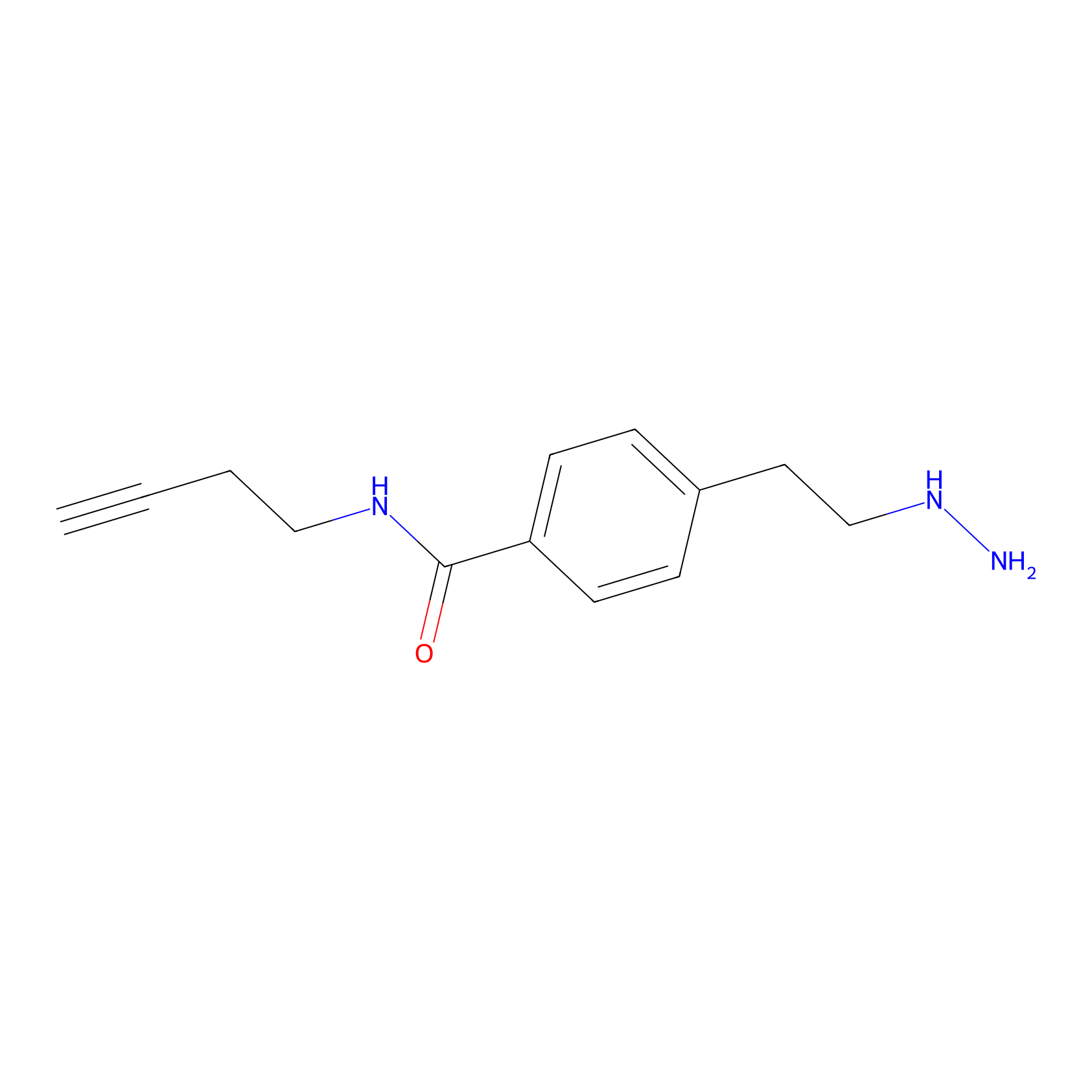 |
20.00 | LDD0282 | [6] | |
|
Jackson_1 Probe Info |
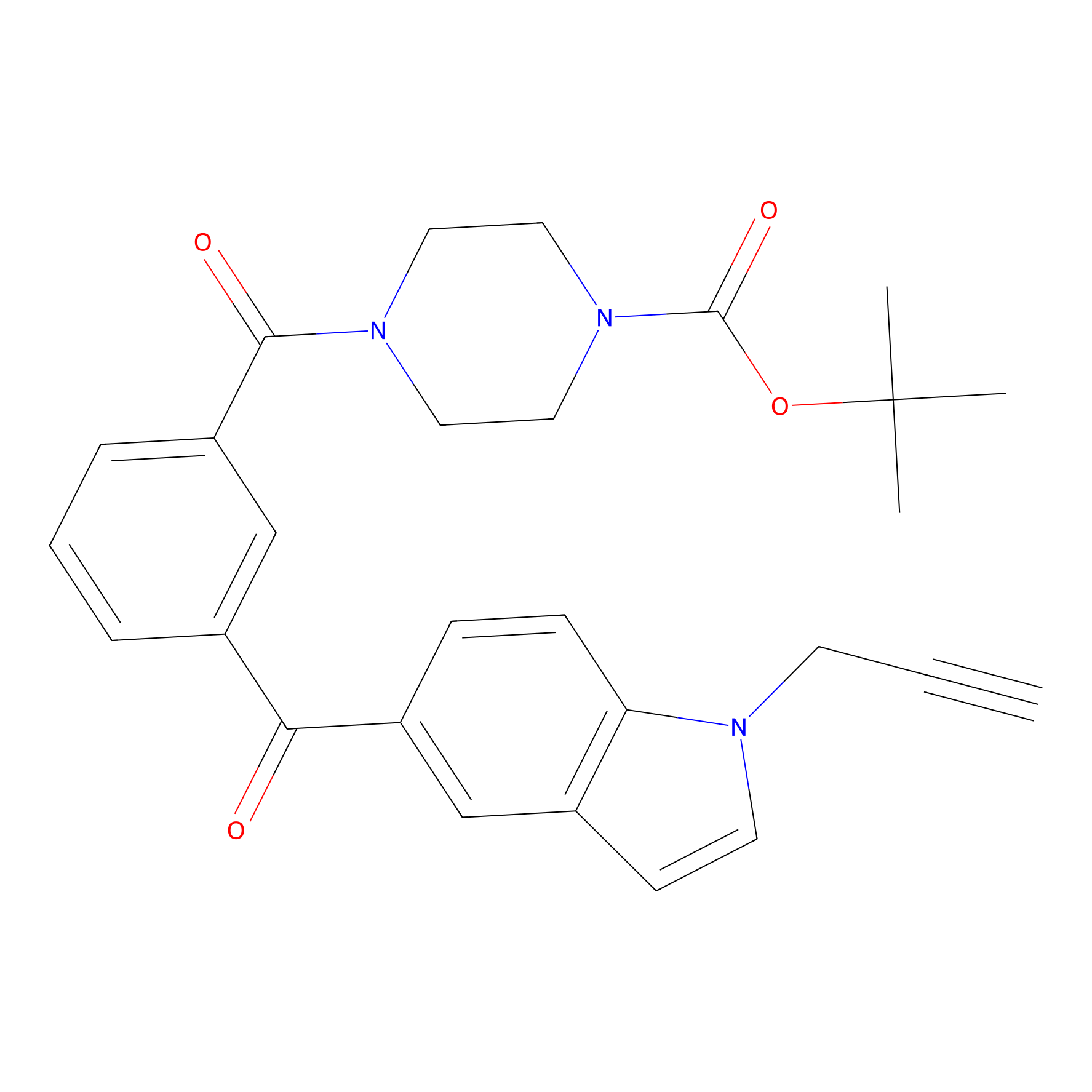 |
2.95 | LDD0121 | [7] | |
|
Acrolein Probe Info |
 |
H76(0.00); H62(0.00) | LDD0223 | [8] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [8] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C027 Probe Info |
 |
5.90 | LDD1733 | [9] | |
|
C040 Probe Info |
 |
6.68 | LDD1740 | [9] | |
|
C094 Probe Info |
 |
24.42 | LDD1785 | [9] | |
|
C161 Probe Info |
 |
10.48 | LDD1841 | [9] | |
|
C220 Probe Info |
 |
12.73 | LDD1894 | [9] | |
|
C235 Probe Info |
 |
17.03 | LDD1908 | [9] | |
|
C275 Probe Info |
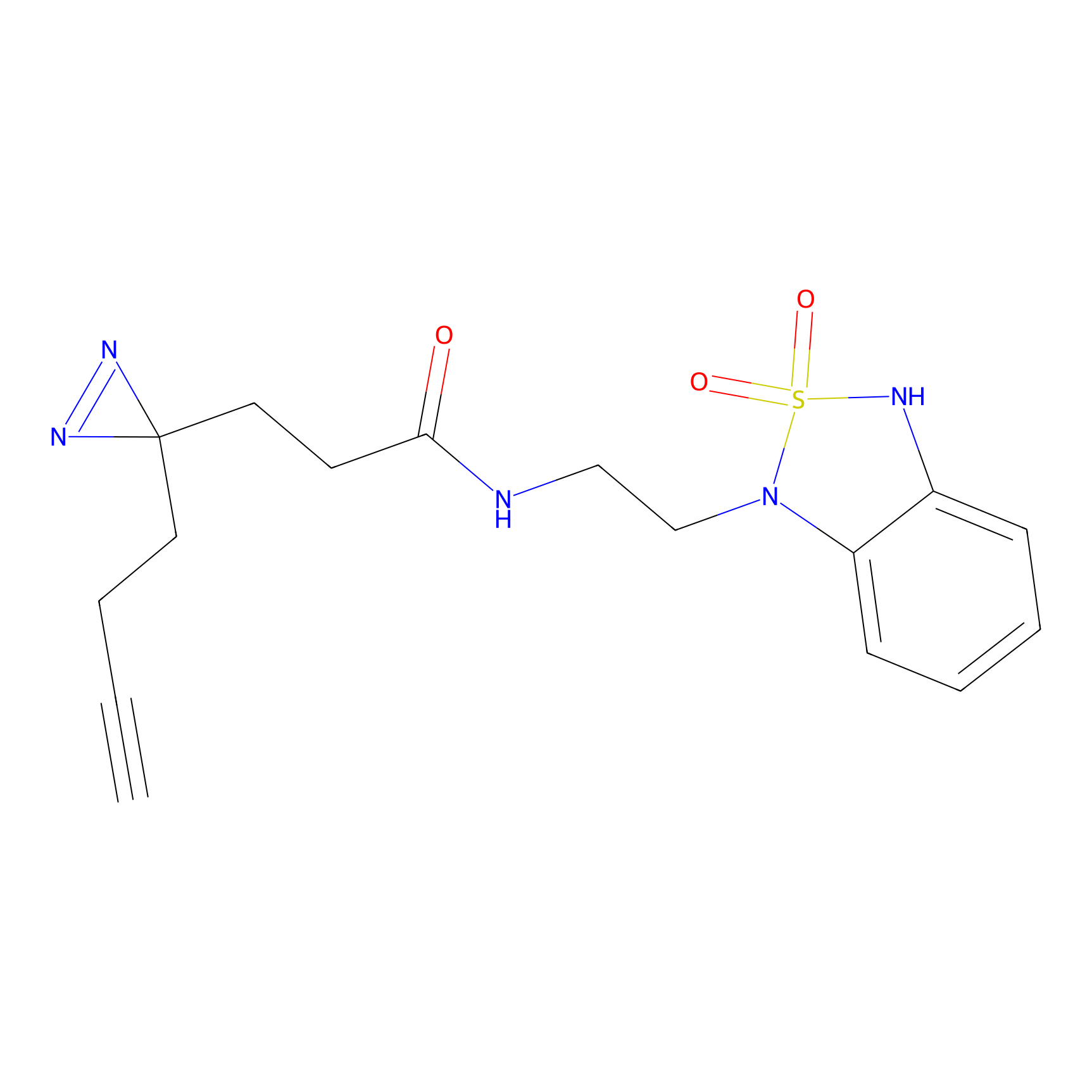 |
5.54 | LDD1945 | [9] | |
|
C349 Probe Info |
 |
9.13 | LDD2010 | [9] | |
|
C350 Probe Info |
 |
22.01 | LDD2011 | [9] | |
|
C353 Probe Info |
 |
17.39 | LDD2014 | [9] | |
|
C382 Probe Info |
 |
9.78 | LDD2041 | [9] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [10] | |
|
FFF probe15 Probe Info |
 |
20.00 | LDD0478 | [10] | |
|
FFF probe7 Probe Info |
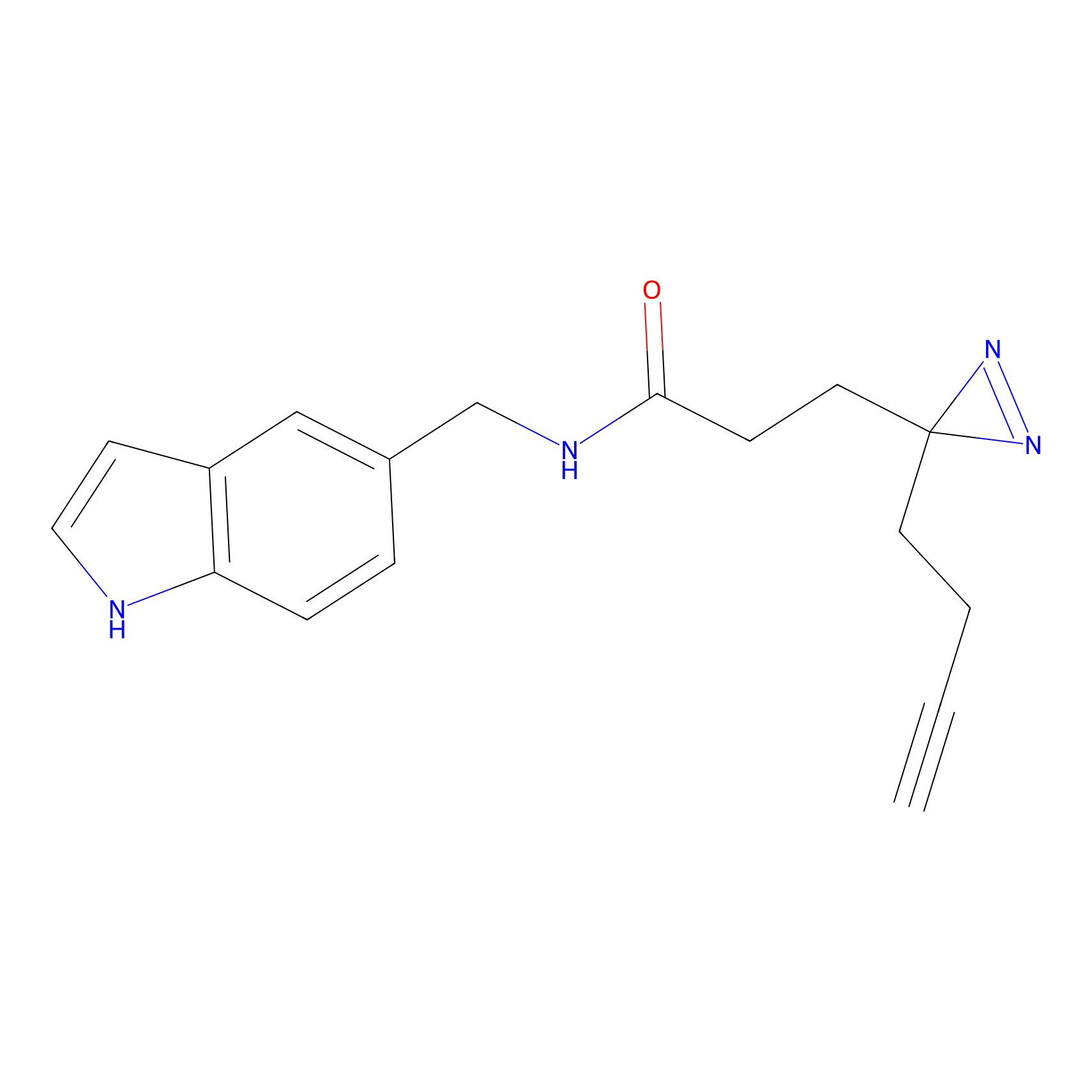 |
20.00 | LDD0483 | [10] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [11] | |
|
OEA-DA Probe Info |
 |
20.00 | LDD0046 | [12] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Zinc finger protein 655 (ZNF655) | Krueppel C2H2-type zinc-finger protein family | Q8N720 | |||
Other
References
