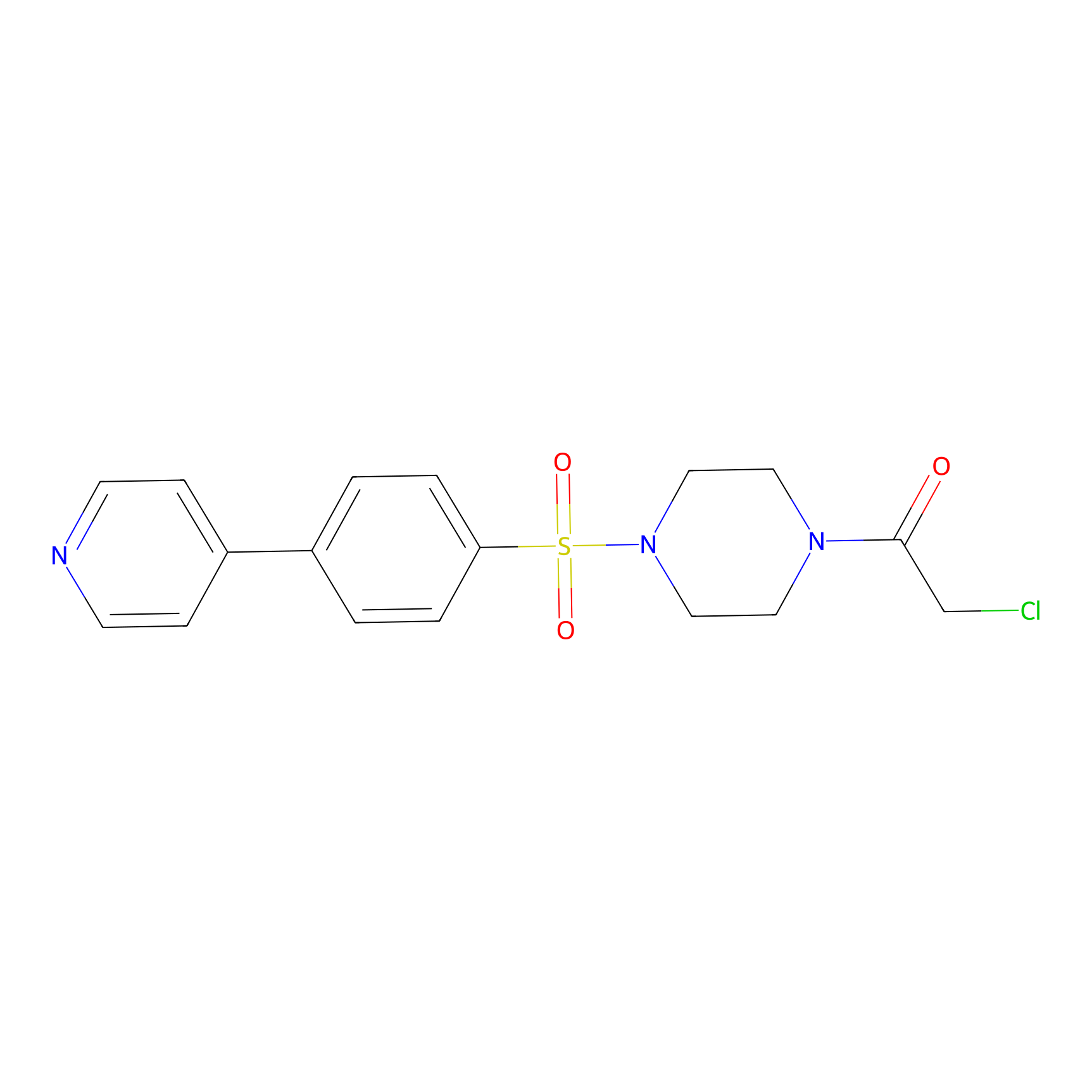
Details of the Competitor
General Information of Competitor
The Competitor Interaction Atlas
Probe(s) Related This Competitor
ABPP Probe
| Probe Name | Structure | Concentration | Cell-system | Ref | |
|---|---|---|---|---|---|
|
IA-alkyne Probe Info |
 |
100 uM | Human normal cell lysate (T cell) | [1] | |
Target(s) List of this Competitor
|
36 Enzyme Competed by This
Competitor
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
2 Transporter and channel Competed by This
Competitor
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
1 Immunoglobulin Competed by This
Competitor
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
5 Other Competed by This
Competitor
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Full Information of The Labelling Profiles of This Competitor

IA-alkyne
Quantification: Probe vs (Probe+Competitor)
|
Experiment 1 Reporting the Labelling Profiles of This Probe
|
||||
| Probe concentration | ||||
| Quantitative Method | ||||
| Competitor Name | ||||
| Competitor Concentration | ||||
| In Vitro Experiment Model |

|
|||

|
||||
| Normal | ||||
| Human normal cell lysate (T cell) | ||||
| Interaction Atlas ID |
 Download The Altas
Download The Altas
|
|||
|
Experiment 2 Reporting the Labelling Profiles of This Probe
|
||||
| Probe concentration | ||||
| Quantitative Method | ||||
| Competitor Name | ||||
| Competitor Concentration | ||||
| In Vitro Experiment Model |

|
|||

|
||||
| Normal | ||||
| Human normal cell lysate (T cell) | ||||
| Interaction Atlas ID |
 Download The Altas
Download The Altas
|
|||