Details of the Target
General Information of Target
| Target ID | LDTP10046 | |||||
|---|---|---|---|---|---|---|
| Target Name | Endoplasmic reticulum-Golgi intermediate compartment protein 1 (ERGIC1) | |||||
| Gene Name | ERGIC1 | |||||
| Gene ID | 57222 | |||||
| Synonyms |
ERGIC32; KIAA1181; Endoplasmic reticulum-Golgi intermediate compartment protein 1; ER-Golgi intermediate compartment 32 kDa protein; ERGIC-32 |
|||||
| 3D Structure | ||||||
| Sequence |
MACSRPPSQCEPTSLPPGPPAGRRHLPLSRRRREMSSNKEQRSAVFVILFALITILILYS
SNSANEVFHYGSLRGRSRRPVNLKKWSITDGYVPILGNKTLPSRCHQCVIVSSSSHLLGT KLGPEIERAECTIRMNDAPTTGYSADVGNKTTYRVVAHSSVFRVLRRPQEFVNRTPETVF IFWGPPSKMQKPQGSLVRVIQRAGLVFPNMEAYAVSPGRMRQFDDLFRGETGKDREKSHS WLSTGWFTMVIAVELCDHVHVYGMVPPNYCSQRPRLQRMPYHYYEPKGPDECVTYIQNEH SRKGNHHRFITEKRVFSSWAQLYGITFSHPSWT |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
ERGIC family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function | Possible role in transport between endoplasmic reticulum and Golgi. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| HCT15 | SNV: p.S272P | DBIA Probe Info | |||
| IGROV1 | Deletion: p.G173EfsTer17 | DBIA Probe Info | |||
| NCIH1155 | SNV: p.C82Y | . | |||
| SKMEL5 | SNV: p.Q285Ter | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FBPP2 Probe Info |
 |
2.27 | LDD0318 | [1] | |
|
TH211 Probe Info |
 |
Y219(20.00); Y196(16.04) | LDD0260 | [2] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [3] | |
|
STPyne Probe Info |
 |
K190(10.00); K288(6.14) | LDD0277 | [4] | |
|
DBIA Probe Info |
 |
C70(3.37) | LDD3310 | [5] | |
|
BTD Probe Info |
 |
C115(0.68) | LDD2117 | [6] | |
|
IPM Probe Info |
 |
C115(1.83) | LDD1702 | [6] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [7] | |
|
m-APA Probe Info |
 |
H221(0.00); H165(0.00) | LDD2231 | [7] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [8] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [8] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [8] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [9] | |
|
Crotonaldehyde Probe Info |
 |
H221(0.00); C115(0.00) | LDD0219 | [9] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [10] | |
|
MPP-AC Probe Info |
 |
N.A. | LDD0428 | [11] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [12] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [11] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [11] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C091 Probe Info |
 |
10.70 | LDD1782 | [13] | |
|
C092 Probe Info |
 |
22.94 | LDD1783 | [13] | |
|
C094 Probe Info |
 |
33.36 | LDD1785 | [13] | |
|
C141 Probe Info |
 |
9.13 | LDD1823 | [13] | |
|
C210 Probe Info |
 |
67.65 | LDD1884 | [13] | |
|
C213 Probe Info |
 |
17.27 | LDD1887 | [13] | |
|
C228 Probe Info |
 |
14.62 | LDD1901 | [13] | |
|
C246 Probe Info |
 |
12.30 | LDD1919 | [13] | |
|
C289 Probe Info |
 |
38.05 | LDD1959 | [13] | |
|
C362 Probe Info |
 |
52.71 | LDD2023 | [13] | |
|
C363 Probe Info |
 |
19.84 | LDD2024 | [13] | |
|
C364 Probe Info |
 |
19.16 | LDD2025 | [13] | |
|
C366 Probe Info |
 |
5.62 | LDD2027 | [13] | |
|
C388 Probe Info |
 |
38.32 | LDD2047 | [13] | |
|
C403 Probe Info |
 |
14.93 | LDD2061 | [13] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0136 | [14] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [14] | |
|
HM-2 Probe Info |
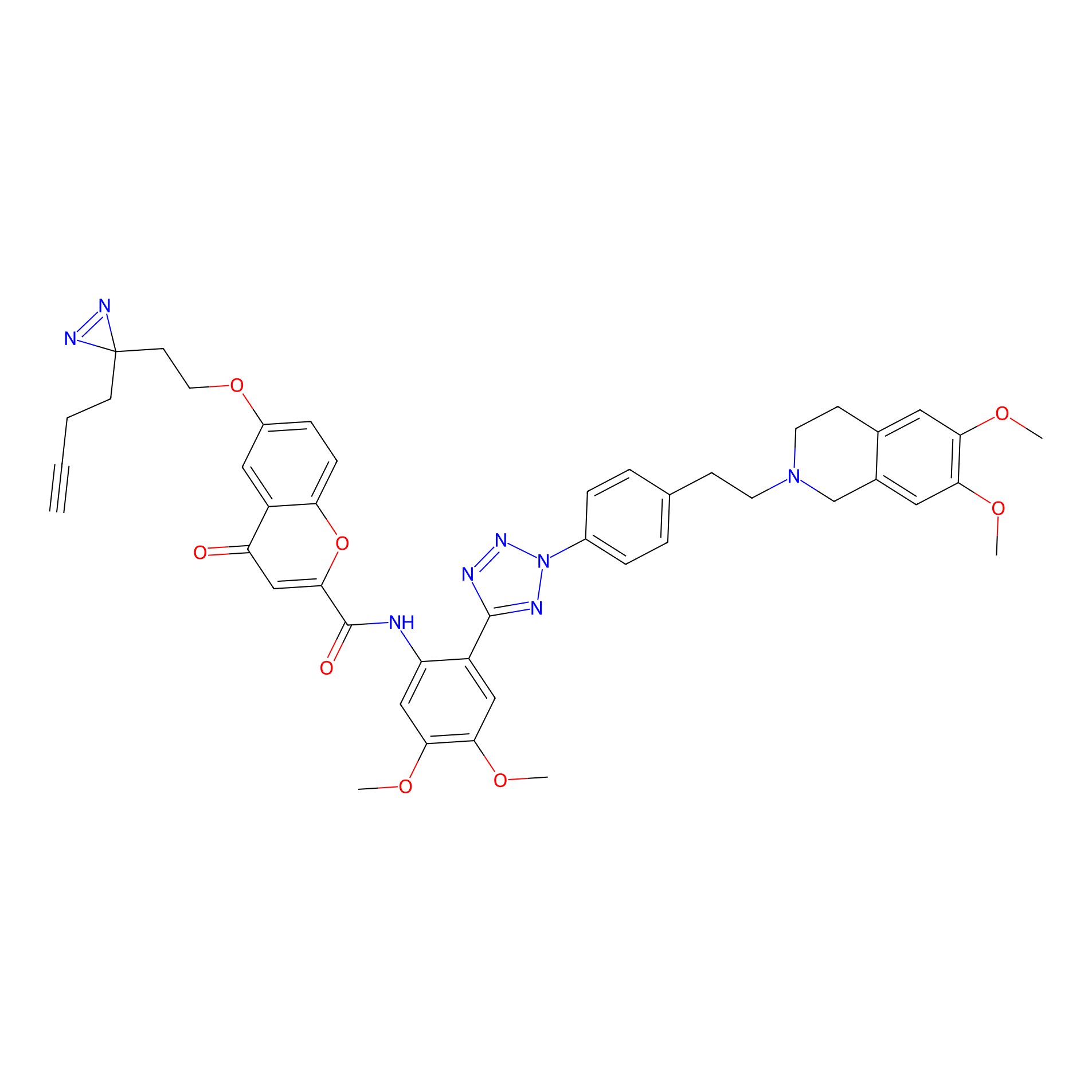 |
1.97 | LDD0436 | [15] | |
|
OEA-DA Probe Info |
 |
12.51 | LDD0046 | [16] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C115(0.68) | LDD2117 | [6] |
| LDCM0108 | Chloroacetamide | HeLa | H96(0.00); H165(0.00) | LDD0222 | [9] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C115(1.83) | LDD1702 | [6] |
| LDCM0173 | HM30181 | Hep-G2 | 1.97 | LDD0436 | [15] |
| LDCM0107 | IAA | HeLa | H165(0.00); H96(0.00) | LDD0221 | [9] |
| LDCM0022 | KB02 | 697 | C70(1.78) | LDD2245 | [5] |
| LDCM0023 | KB03 | 697 | C70(2.28) | LDD2662 | [5] |
| LDCM0024 | KB05 | COLO792 | C70(3.37) | LDD3310 | [5] |
| LDCM0109 | NEM | HeLa | H165(0.00); H96(0.00); H100(0.00); H185(0.00) | LDD0223 | [9] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C115(0.77) | LDD2123 | [6] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C115(0.49) | LDD2125 | [6] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C115(0.39) | LDD2127 | [6] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C115(1.08) | LDD2136 | [6] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C115(0.34) | LDD2140 | [6] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Peroxisomal bifunctional enzyme (EHHADH) | Enoyl-CoA hydratase/isomerase family; 3-hydroxyacyl-CoA dehydrogenase family | Q08426 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Nucleoporin p58/p45 (NUP58) | NUP58 family | Q9BVL2 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Krueppel-like factor 11 (KLF11) | Sp1 C2H2-type zinc-finger protein family | O14901 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Endoplasmic reticulum-Golgi intermediate compartment protein 3 (ERGIC3) | ERGIC family | Q9Y282 | |||
References
