Details of the Target
General Information of Target
| Target ID | LDTP08950 | |||||
|---|---|---|---|---|---|---|
| Target Name | ADP-ribosylation factor GTPase-activating protein 1 (ARFGAP1) | |||||
| Gene Name | ARFGAP1 | |||||
| Gene ID | 55738 | |||||
| Synonyms |
ARF1GAP; ADP-ribosylation factor GTPase-activating protein 1; ARF GAP 1; ADP-ribosylation factor 1 GTPase-activating protein; ARF1 GAP; ARF1-directed GTPase-activating protein |
|||||
| 3D Structure | ||||||
| Sequence |
MASPRTRKVLKEVRVQDENNVCFECGAFNPQWVSVTYGIWICLECSGRHRGLGVHLSFVR
SVTMDKWKDIELEKMKAGGNAKFREFLESQEDYDPCWSLQEKYNSRAAALFRDKVVALAE GREWSLESSPAQNWTPPQPRTLPSMVHRVSGQPQSVTASSDKAFEDWLNDDLGSYQGAQG NRYVGFGNTPPPQKKEDDFLNNAMSSLYSGWSSFTTGASRFASAAKEGATKFGSQASQKA SELGHSLNENVLKPAQEKVKEGKIFDDVSSGVSQLASKVQGVGSKGWRDVTTFFSGKAEG PLDSPSEGHSYQNSGLDHFQNSNIDQSFWETFGSAEPTKTRKSPSSDSWTCADTSTERRS SDSWEVWGSASTNRNSNSDGGEGGEGTKKAVPPAVPTDDGWDNQNW |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
GTPase-activating protein (GAP) for the ADP ribosylation factor 1 (ARF1). Involved in membrane trafficking and /or vesicle transport. Promotes hydrolysis of the ARF1-bound GTP and thus, is required for the dissociation of coat proteins from Golgi-derived membranes and vesicles, a prerequisite for vesicle's fusion with target compartment. Probably regulates ARF1-mediated transport via its interaction with the KDELR proteins and TMED2. Overexpression induces the redistribution of the entire Golgi complex to the endoplasmic reticulum, as when ARF1 is deactivated. Its activity is stimulated by phosphoinosides and inhibited by phosphatidylcholine.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| HCT116 | SNV: p.A276V | . | |||
| RKO | SNV: p.A27T | . | |||
| SKMEL2 | Deletion: p.S295RfsTer30 | DBIA Probe Info | |||
| SUPT1 | SNV: p.R182H; p.G386C | DBIA Probe Info | |||
| TGBC1TKB | SNV: p.L99S | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [2] | |
|
TH211 Probe Info |
 |
Y183(8.08) | LDD0260 | [3] | |
|
STPyne Probe Info |
 |
K11(5.38); K114(10.00); K226(5.84); K231(1.43) | LDD0277 | [4] | |
|
ONAyne Probe Info |
 |
K226(5.56) | LDD0275 | [4] | |
|
Probe 1 Probe Info |
 |
Y175(6.36); Y183(13.50) | LDD3495 | [5] | |
|
DBIA Probe Info |
 |
C96(0.72) | LDD3314 | [6] | |
|
BTD Probe Info |
 |
C351(0.89) | LDD2091 | [7] | |
|
Jackson_1 Probe Info |
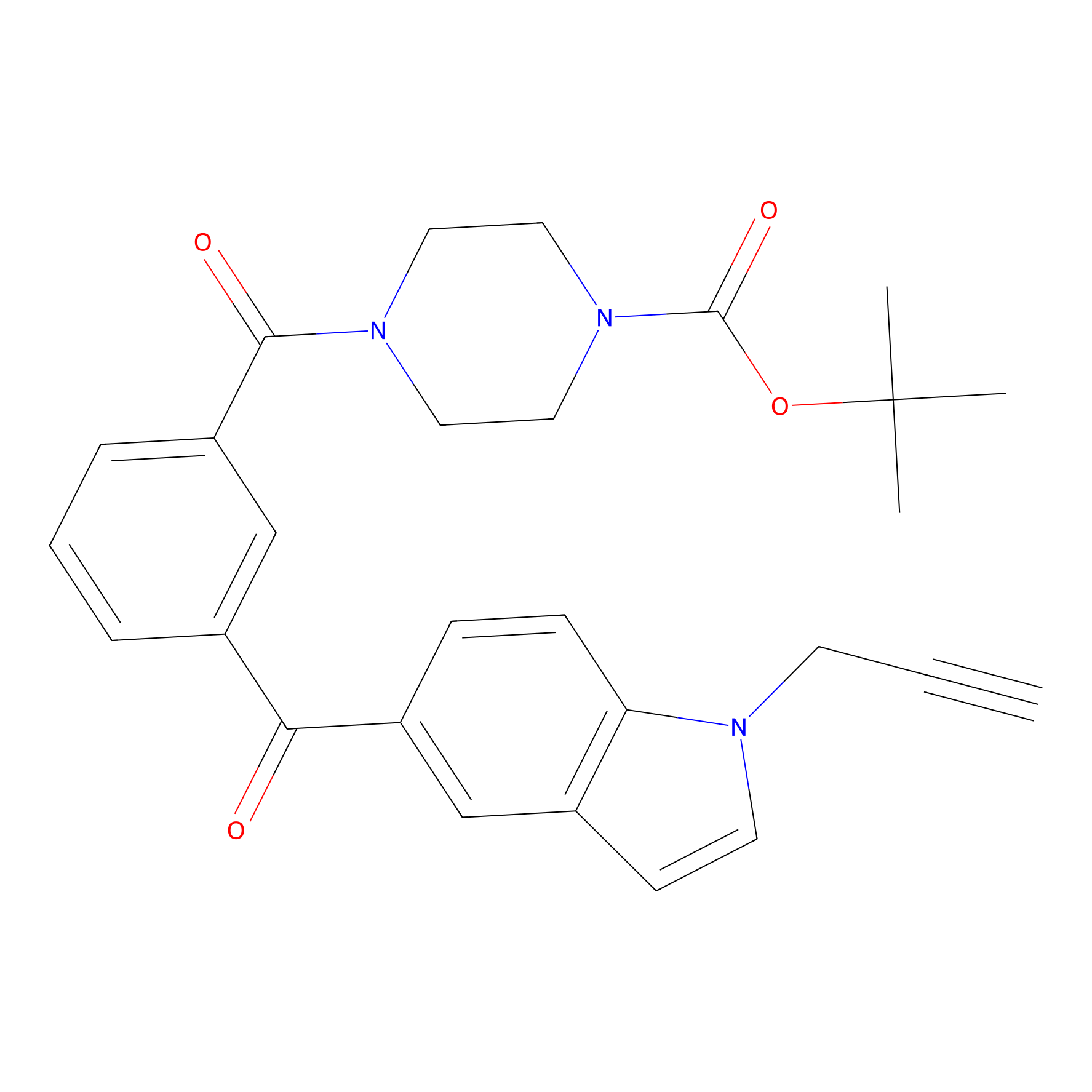 |
20.00 | LDD0121 | [8] | |
|
AHL-Pu-1 Probe Info |
 |
C96(2.60); C351(2.54) | LDD0168 | [9] | |
|
HHS-475 Probe Info |
 |
Y183(0.81); Y311(1.22); Y175(1.68) | LDD0264 | [10] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [11] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [11] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [11] | |
|
NAIA_4 Probe Info |
 |
C96(0.00); C351(0.00) | LDD2226 | [12] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [13] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [14] | |
|
DA-P3 Probe Info |
 |
N.A. | LDD0178 | [15] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [16] | |
|
Compound 11 Probe Info |
 |
N.A. | LDD2213 | [16] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [14] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [14] | |
|
Phosphinate-6 Probe Info |
 |
C96(0.00); C351(0.00) | LDD0018 | [17] | |
|
Acrolein Probe Info |
 |
C351(0.00); C96(0.00) | LDD0217 | [18] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [18] | |
|
W1 Probe Info |
 |
C96(0.00); C351(0.00) | LDD0236 | [19] | |
|
AOyne Probe Info |
 |
11.50 | LDD0443 | [20] | |
|
NAIA_5 Probe Info |
 |
C96(0.00); C351(0.00) | LDD2223 | [12] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C187 Probe Info |
 |
18.51 | LDD1865 | [21] | |
|
C191 Probe Info |
 |
14.62 | LDD1868 | [21] | |
|
C193 Probe Info |
 |
6.23 | LDD1869 | [21] | |
|
A-DA Probe Info |
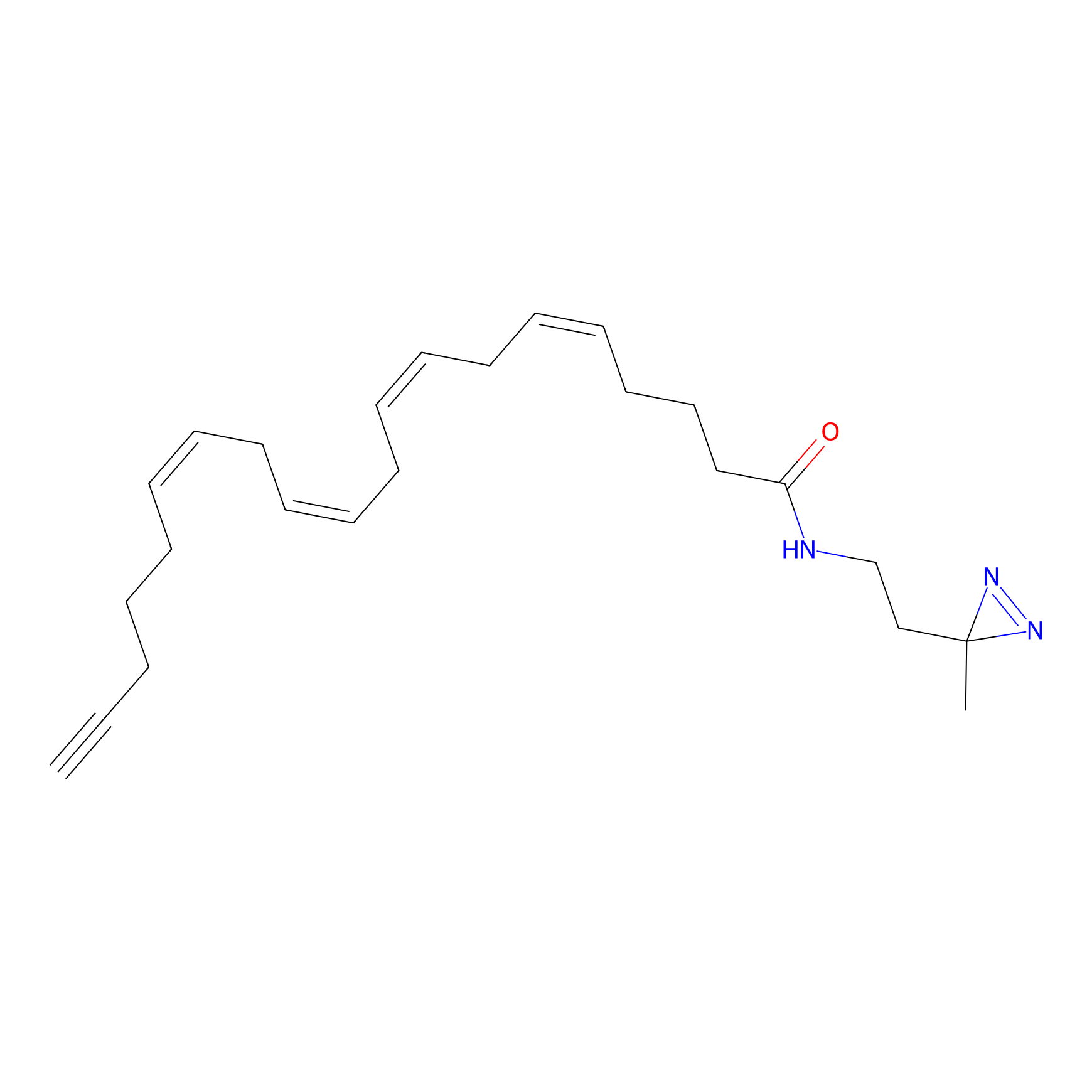 |
14.31 | LDD0143 | [22] | |
|
Alk-rapa Probe Info |
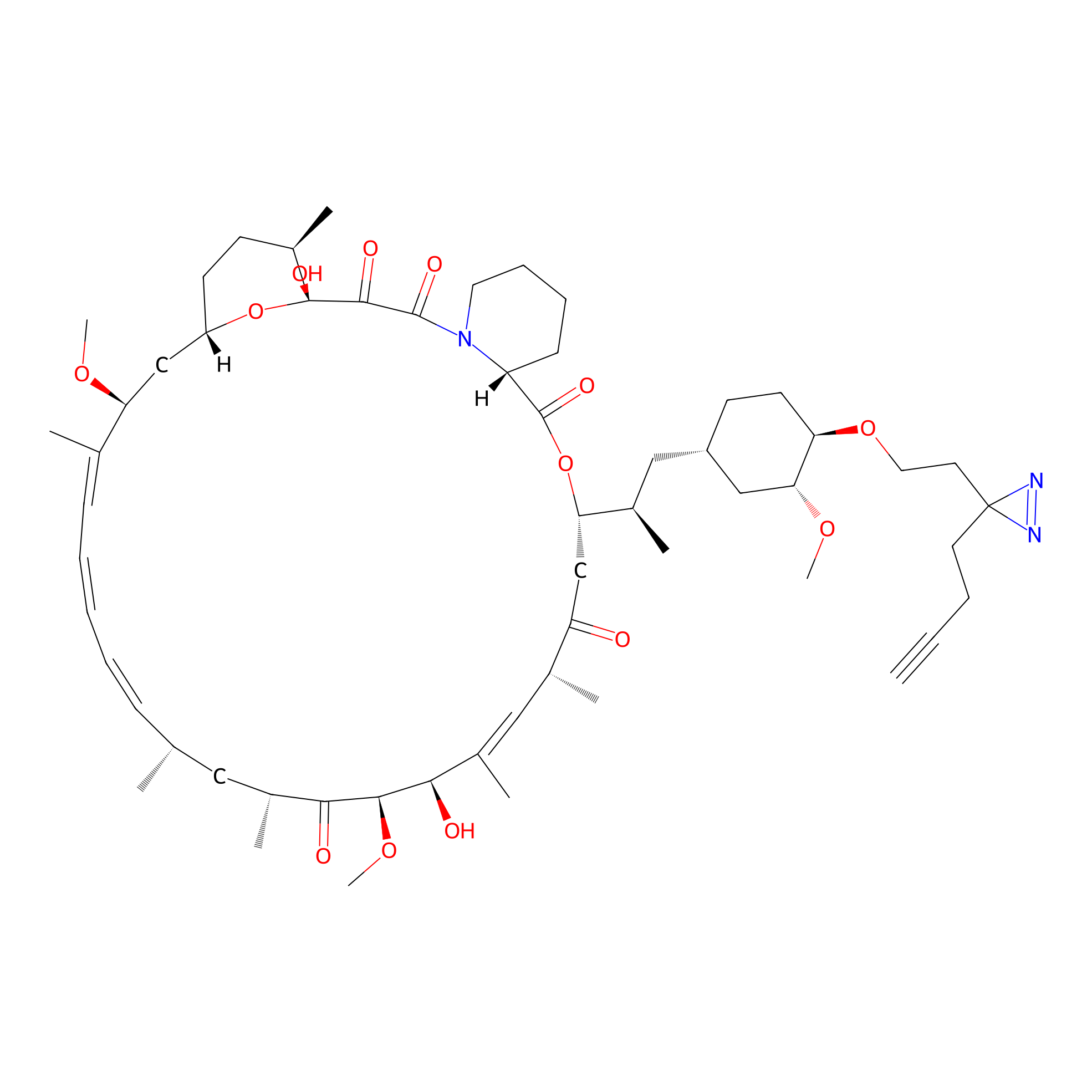 |
4.02 | LDD0213 | [23] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C351(1.19) | LDD2130 | [7] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C351(0.72) | LDD2132 | [7] |
| LDCM0025 | 4SU-RNA | HEK-293T | C96(2.60); C351(2.54) | LDD0168 | [9] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C351(5.56) | LDD0169 | [9] |
| LDCM0214 | AC1 | HEK-293T | C351(1.05); C96(1.09) | LDD1507 | [24] |
| LDCM0215 | AC10 | HEK-293T | C351(0.92); C96(0.88) | LDD1508 | [24] |
| LDCM0226 | AC11 | HEK-293T | C351(1.10); C96(0.99) | LDD1509 | [24] |
| LDCM0237 | AC12 | HEK-293T | C351(1.07); C96(0.99) | LDD1510 | [24] |
| LDCM0259 | AC14 | HEK-293T | C351(1.02); C96(0.92) | LDD1512 | [24] |
| LDCM0270 | AC15 | HEK-293T | C351(1.01); C96(0.92) | LDD1513 | [24] |
| LDCM0276 | AC17 | HEK-293T | C351(1.01); C96(0.98) | LDD1515 | [24] |
| LDCM0277 | AC18 | HEK-293T | C351(1.00); C96(0.92) | LDD1516 | [24] |
| LDCM0278 | AC19 | HEK-293T | C351(1.15); C96(1.03) | LDD1517 | [24] |
| LDCM0279 | AC2 | HEK-293T | C351(1.12); C96(0.98) | LDD1518 | [24] |
| LDCM0280 | AC20 | HEK-293T | C351(1.04); C96(0.99) | LDD1519 | [24] |
| LDCM0281 | AC21 | HEK-293T | C351(1.06); C96(1.00) | LDD1520 | [24] |
| LDCM0282 | AC22 | HEK-293T | C351(1.03); C96(0.91) | LDD1521 | [24] |
| LDCM0283 | AC23 | HEK-293T | C351(1.01); C96(1.09) | LDD1522 | [24] |
| LDCM0284 | AC24 | HEK-293T | C351(0.97); C96(0.94) | LDD1523 | [24] |
| LDCM0285 | AC25 | HEK-293T | C351(1.02); C96(1.00) | LDD1524 | [24] |
| LDCM0286 | AC26 | HEK-293T | C351(0.97); C96(0.87) | LDD1525 | [24] |
| LDCM0287 | AC27 | HEK-293T | C351(1.14); C96(1.05) | LDD1526 | [24] |
| LDCM0288 | AC28 | HEK-293T | C351(0.96); C96(0.93) | LDD1527 | [24] |
| LDCM0289 | AC29 | HEK-293T | C351(1.19); C96(0.96) | LDD1528 | [24] |
| LDCM0290 | AC3 | HEK-293T | C351(1.13); C96(1.11) | LDD1529 | [24] |
| LDCM0291 | AC30 | HEK-293T | C351(1.10); C96(1.04) | LDD1530 | [24] |
| LDCM0292 | AC31 | HEK-293T | C351(1.04); C96(0.98) | LDD1531 | [24] |
| LDCM0293 | AC32 | HEK-293T | C351(0.92); C96(0.95) | LDD1532 | [24] |
| LDCM0294 | AC33 | HEK-293T | C351(1.09); C96(0.95) | LDD1533 | [24] |
| LDCM0295 | AC34 | HEK-293T | C351(1.08); C96(0.92) | LDD1534 | [24] |
| LDCM0296 | AC35 | HEK-293T | C351(1.12); C96(1.12) | LDD1535 | [24] |
| LDCM0297 | AC36 | HEK-293T | C351(1.07); C96(0.99) | LDD1536 | [24] |
| LDCM0298 | AC37 | HEK-293T | C351(1.18); C96(1.00) | LDD1537 | [24] |
| LDCM0299 | AC38 | HEK-293T | C351(1.13); C96(0.86) | LDD1538 | [24] |
| LDCM0300 | AC39 | HEK-293T | C351(1.07); C96(0.98) | LDD1539 | [24] |
| LDCM0301 | AC4 | HEK-293T | C351(1.07); C96(1.01) | LDD1540 | [24] |
| LDCM0302 | AC40 | HEK-293T | C351(0.94); C96(0.84) | LDD1541 | [24] |
| LDCM0303 | AC41 | HEK-293T | C351(1.05); C96(0.99) | LDD1542 | [24] |
| LDCM0304 | AC42 | HEK-293T | C351(1.01); C96(0.88) | LDD1543 | [24] |
| LDCM0305 | AC43 | HEK-293T | C351(1.04); C96(1.05) | LDD1544 | [24] |
| LDCM0306 | AC44 | HEK-293T | C351(1.07); C96(0.95) | LDD1545 | [24] |
| LDCM0307 | AC45 | HEK-293T | C351(1.13); C96(0.95) | LDD1546 | [24] |
| LDCM0308 | AC46 | HEK-293T | C351(0.98); C96(0.92) | LDD1547 | [24] |
| LDCM0309 | AC47 | HEK-293T | C351(0.99); C96(0.96) | LDD1548 | [24] |
| LDCM0310 | AC48 | HEK-293T | C351(0.93); C96(0.94) | LDD1549 | [24] |
| LDCM0311 | AC49 | HEK-293T | C351(1.18); C96(0.99) | LDD1550 | [24] |
| LDCM0312 | AC5 | HEK-293T | C351(0.98); C96(0.97) | LDD1551 | [24] |
| LDCM0313 | AC50 | HEK-293T | C351(0.97); C96(0.89) | LDD1552 | [24] |
| LDCM0314 | AC51 | HEK-293T | C351(1.07); C96(1.04) | LDD1553 | [24] |
| LDCM0315 | AC52 | HEK-293T | C351(1.07); C96(0.91) | LDD1554 | [24] |
| LDCM0316 | AC53 | HEK-293T | C351(1.14); C96(0.99) | LDD1555 | [24] |
| LDCM0317 | AC54 | HEK-293T | C351(1.06); C96(0.95) | LDD1556 | [24] |
| LDCM0318 | AC55 | HEK-293T | C351(0.98); C96(0.97) | LDD1557 | [24] |
| LDCM0319 | AC56 | HEK-293T | C351(1.04); C96(0.93) | LDD1558 | [24] |
| LDCM0320 | AC57 | HEK-293T | C351(1.16); C96(0.99) | LDD1559 | [24] |
| LDCM0321 | AC58 | HEK-293T | C351(1.01); C96(0.95) | LDD1560 | [24] |
| LDCM0322 | AC59 | HEK-293T | C351(1.03); C96(1.09) | LDD1561 | [24] |
| LDCM0323 | AC6 | HEK-293T | C351(1.02); C96(0.95) | LDD1562 | [24] |
| LDCM0324 | AC60 | HEK-293T | C351(1.11); C96(1.12) | LDD1563 | [24] |
| LDCM0325 | AC61 | HEK-293T | C351(1.09); C96(0.94) | LDD1564 | [24] |
| LDCM0326 | AC62 | HEK-293T | C351(1.35); C96(0.92) | LDD1565 | [24] |
| LDCM0327 | AC63 | HEK-293T | C351(1.03); C96(0.95) | LDD1566 | [24] |
| LDCM0328 | AC64 | HEK-293T | C351(0.93); C96(1.06) | LDD1567 | [24] |
| LDCM0334 | AC7 | HEK-293T | C351(1.01); C96(1.01) | LDD1568 | [24] |
| LDCM0345 | AC8 | HEK-293T | C351(0.93); C96(0.91) | LDD1569 | [24] |
| LDCM0248 | AKOS034007472 | HEK-293T | C351(1.16); C96(0.98) | LDD1511 | [24] |
| LDCM0356 | AKOS034007680 | HEK-293T | C351(1.09); C96(0.97) | LDD1570 | [24] |
| LDCM0275 | AKOS034007705 | HEK-293T | C351(0.90); C96(0.94) | LDD1514 | [24] |
| LDCM0156 | Aniline | NCI-H1299 | 12.29 | LDD0403 | [1] |
| LDCM0083 | Avasimibe | A-549 | 14.31 | LDD0143 | [22] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C351(0.89) | LDD2091 | [7] |
| LDCM0630 | CCW28-3 | 231MFP | C96(1.64) | LDD2214 | [25] |
| LDCM0108 | Chloroacetamide | HeLa | C351(0.00); C96(0.00) | LDD0222 | [18] |
| LDCM0071 | Cisar_cp14 | MDA-MB-231 | 20.00 | LDD0121 | [8] |
| LDCM0632 | CL-Sc | Hep-G2 | C351(1.08); C351(0.74); C351(0.64) | LDD2227 | [12] |
| LDCM0367 | CL1 | HEK-293T | C351(1.05); C96(0.83) | LDD1571 | [24] |
| LDCM0368 | CL10 | HEK-293T | C351(0.88); C96(0.77) | LDD1572 | [24] |
| LDCM0369 | CL100 | HEK-293T | C351(0.98); C96(1.00) | LDD1573 | [24] |
| LDCM0370 | CL101 | HEK-293T | C351(1.05); C96(0.80) | LDD1574 | [24] |
| LDCM0371 | CL102 | HEK-293T | C351(0.99); C96(0.95) | LDD1575 | [24] |
| LDCM0372 | CL103 | HEK-293T | C351(1.08); C96(1.04) | LDD1576 | [24] |
| LDCM0373 | CL104 | HEK-293T | C351(0.93); C96(0.98) | LDD1577 | [24] |
| LDCM0374 | CL105 | HEK-293T | C351(1.01); C96(0.85) | LDD1578 | [24] |
| LDCM0375 | CL106 | HEK-293T | C351(1.05); C96(1.20) | LDD1579 | [24] |
| LDCM0376 | CL107 | HEK-293T | C351(0.96); C96(1.05) | LDD1580 | [24] |
| LDCM0377 | CL108 | HEK-293T | C351(0.98); C96(0.96) | LDD1581 | [24] |
| LDCM0378 | CL109 | HEK-293T | C351(0.90); C96(0.90) | LDD1582 | [24] |
| LDCM0379 | CL11 | HEK-293T | C351(1.13); C96(1.18) | LDD1583 | [24] |
| LDCM0380 | CL110 | HEK-293T | C351(0.98); C96(1.11) | LDD1584 | [24] |
| LDCM0381 | CL111 | HEK-293T | C351(0.96); C96(0.91) | LDD1585 | [24] |
| LDCM0382 | CL112 | HEK-293T | C351(1.00); C96(0.93) | LDD1586 | [24] |
| LDCM0383 | CL113 | HEK-293T | C351(1.04); C96(0.79) | LDD1587 | [24] |
| LDCM0384 | CL114 | HEK-293T | C351(1.00); C96(0.88) | LDD1588 | [24] |
| LDCM0385 | CL115 | HEK-293T | C351(0.99); C96(0.88) | LDD1589 | [24] |
| LDCM0386 | CL116 | HEK-293T | C351(0.98); C96(0.97) | LDD1590 | [24] |
| LDCM0387 | CL117 | HEK-293T | C351(0.96); C96(0.74) | LDD1591 | [24] |
| LDCM0388 | CL118 | HEK-293T | C351(1.10); C96(0.98) | LDD1592 | [24] |
| LDCM0389 | CL119 | HEK-293T | C351(0.98); C96(0.95) | LDD1593 | [24] |
| LDCM0390 | CL12 | HEK-293T | C351(0.79); C96(1.00) | LDD1594 | [24] |
| LDCM0391 | CL120 | HEK-293T | C351(0.91); C96(1.04) | LDD1595 | [24] |
| LDCM0392 | CL121 | HEK-293T | C351(0.96); C96(0.96) | LDD1596 | [24] |
| LDCM0393 | CL122 | HEK-293T | C351(0.98); C96(1.16) | LDD1597 | [24] |
| LDCM0394 | CL123 | HEK-293T | C351(1.11); C96(0.84) | LDD1598 | [24] |
| LDCM0395 | CL124 | HEK-293T | C351(0.99); C96(0.90) | LDD1599 | [24] |
| LDCM0396 | CL125 | HEK-293T | C351(1.02); C96(0.77) | LDD1600 | [24] |
| LDCM0397 | CL126 | HEK-293T | C351(1.06); C96(0.94) | LDD1601 | [24] |
| LDCM0398 | CL127 | HEK-293T | C351(0.98); C96(0.99) | LDD1602 | [24] |
| LDCM0399 | CL128 | HEK-293T | C351(0.96); C96(0.90) | LDD1603 | [24] |
| LDCM0400 | CL13 | HEK-293T | C351(1.02); C96(0.83) | LDD1604 | [24] |
| LDCM0401 | CL14 | HEK-293T | C351(1.15); C96(1.14) | LDD1605 | [24] |
| LDCM0402 | CL15 | HEK-293T | C351(1.11); C96(0.90) | LDD1606 | [24] |
| LDCM0403 | CL16 | HEK-293T | C351(0.96); C96(0.90) | LDD1607 | [24] |
| LDCM0404 | CL17 | HEK-293T | C351(1.18); C96(0.90) | LDD1608 | [24] |
| LDCM0405 | CL18 | HEK-293T | C351(1.07); C96(0.82) | LDD1609 | [24] |
| LDCM0406 | CL19 | HEK-293T | C351(1.11); C96(0.93) | LDD1610 | [24] |
| LDCM0407 | CL2 | HEK-293T | C351(1.07); C96(1.24) | LDD1611 | [24] |
| LDCM0408 | CL20 | HEK-293T | C351(1.14); C96(0.63) | LDD1612 | [24] |
| LDCM0409 | CL21 | HEK-293T | C351(0.92); C96(0.99) | LDD1613 | [24] |
| LDCM0410 | CL22 | HEK-293T | C351(0.84); C96(0.92) | LDD1614 | [24] |
| LDCM0411 | CL23 | HEK-293T | C351(1.20); C96(1.21) | LDD1615 | [24] |
| LDCM0412 | CL24 | HEK-293T | C351(0.83); C96(0.93) | LDD1616 | [24] |
| LDCM0413 | CL25 | HEK-293T | C351(0.84); C96(0.88) | LDD1617 | [24] |
| LDCM0414 | CL26 | HEK-293T | C351(1.21); C96(1.02) | LDD1618 | [24] |
| LDCM0415 | CL27 | HEK-293T | C351(1.08); C96(0.96) | LDD1619 | [24] |
| LDCM0416 | CL28 | HEK-293T | C351(0.90); C96(0.92) | LDD1620 | [24] |
| LDCM0417 | CL29 | HEK-293T | C351(1.09); C96(1.07) | LDD1621 | [24] |
| LDCM0418 | CL3 | HEK-293T | C351(1.10); C96(0.94) | LDD1622 | [24] |
| LDCM0419 | CL30 | HEK-293T | C351(1.00); C96(0.89) | LDD1623 | [24] |
| LDCM0420 | CL31 | HEK-293T | C351(1.16); C96(0.87) | LDD1624 | [24] |
| LDCM0421 | CL32 | HEK-293T | C351(1.15); C96(0.68) | LDD1625 | [24] |
| LDCM0422 | CL33 | HEK-293T | C351(0.89); C96(0.90) | LDD1626 | [24] |
| LDCM0423 | CL34 | HEK-293T | C351(0.82); C96(0.86) | LDD1627 | [24] |
| LDCM0424 | CL35 | HEK-293T | C351(1.13); C96(1.22) | LDD1628 | [24] |
| LDCM0425 | CL36 | HEK-293T | C351(0.74); C96(0.87) | LDD1629 | [24] |
| LDCM0426 | CL37 | HEK-293T | C351(1.11); C96(0.79) | LDD1630 | [24] |
| LDCM0428 | CL39 | HEK-293T | C351(1.11); C96(0.96) | LDD1632 | [24] |
| LDCM0429 | CL4 | HEK-293T | C351(1.03); C96(0.94) | LDD1633 | [24] |
| LDCM0430 | CL40 | HEK-293T | C351(0.91); C96(1.03) | LDD1634 | [24] |
| LDCM0431 | CL41 | HEK-293T | C351(1.14); C96(0.90) | LDD1635 | [24] |
| LDCM0432 | CL42 | HEK-293T | C351(1.04); C96(0.87) | LDD1636 | [24] |
| LDCM0433 | CL43 | HEK-293T | C351(1.05); C96(0.92) | LDD1637 | [24] |
| LDCM0434 | CL44 | HEK-293T | C351(1.21); C96(0.76) | LDD1638 | [24] |
| LDCM0435 | CL45 | HEK-293T | C351(1.12); C96(1.07) | LDD1639 | [24] |
| LDCM0436 | CL46 | HEK-293T | C351(0.93); C96(0.88) | LDD1640 | [24] |
| LDCM0437 | CL47 | HEK-293T | C351(1.02); C96(1.12) | LDD1641 | [24] |
| LDCM0438 | CL48 | HEK-293T | C351(0.68); C96(0.89) | LDD1642 | [24] |
| LDCM0439 | CL49 | HEK-293T | C351(0.95); C96(0.71) | LDD1643 | [24] |
| LDCM0440 | CL5 | HEK-293T | C351(1.09); C96(1.01) | LDD1644 | [24] |
| LDCM0441 | CL50 | HEK-293T | C351(1.08); C96(0.86) | LDD1645 | [24] |
| LDCM0443 | CL52 | HEK-293T | C351(0.95); C96(1.01) | LDD1646 | [24] |
| LDCM0444 | CL53 | HEK-293T | C351(1.26); C96(0.96) | LDD1647 | [24] |
| LDCM0445 | CL54 | HEK-293T | C351(1.01); C96(0.75) | LDD1648 | [24] |
| LDCM0446 | CL55 | HEK-293T | C351(1.05); C96(1.05) | LDD1649 | [24] |
| LDCM0447 | CL56 | HEK-293T | C351(1.13); C96(0.69) | LDD1650 | [24] |
| LDCM0448 | CL57 | HEK-293T | C351(1.04); C96(1.05) | LDD1651 | [24] |
| LDCM0449 | CL58 | HEK-293T | C351(0.81); C96(0.89) | LDD1652 | [24] |
| LDCM0450 | CL59 | HEK-293T | C351(1.12); C96(1.20) | LDD1653 | [24] |
| LDCM0451 | CL6 | HEK-293T | C351(0.97); C96(0.81) | LDD1654 | [24] |
| LDCM0452 | CL60 | HEK-293T | C351(0.75); C96(0.80) | LDD1655 | [24] |
| LDCM0453 | CL61 | HEK-293T | C351(1.00); C96(0.84) | LDD1656 | [24] |
| LDCM0454 | CL62 | HEK-293T | C351(1.00); C96(0.94) | LDD1657 | [24] |
| LDCM0455 | CL63 | HEK-293T | C351(1.12); C96(0.88) | LDD1658 | [24] |
| LDCM0456 | CL64 | HEK-293T | C351(1.07); C96(0.87) | LDD1659 | [24] |
| LDCM0457 | CL65 | HEK-293T | C351(1.08); C96(1.06) | LDD1660 | [24] |
| LDCM0458 | CL66 | HEK-293T | C351(1.03); C96(0.86) | LDD1661 | [24] |
| LDCM0459 | CL67 | HEK-293T | C351(1.11); C96(0.94) | LDD1662 | [24] |
| LDCM0460 | CL68 | HEK-293T | C351(1.09); C96(0.69) | LDD1663 | [24] |
| LDCM0461 | CL69 | HEK-293T | C351(1.04); C96(1.08) | LDD1664 | [24] |
| LDCM0462 | CL7 | HEK-293T | C351(1.16); C96(0.97) | LDD1665 | [24] |
| LDCM0463 | CL70 | HEK-293T | C351(0.80); C96(0.95) | LDD1666 | [24] |
| LDCM0464 | CL71 | HEK-293T | C351(1.03); C96(1.16) | LDD1667 | [24] |
| LDCM0465 | CL72 | HEK-293T | C351(0.85); C96(0.89) | LDD1668 | [24] |
| LDCM0466 | CL73 | HEK-293T | C351(0.89); C96(0.72) | LDD1669 | [24] |
| LDCM0467 | CL74 | HEK-293T | C351(1.16); C96(1.06) | LDD1670 | [24] |
| LDCM0469 | CL76 | HEK-293T | C351(0.96); C96(0.93) | LDD1672 | [24] |
| LDCM0470 | CL77 | HEK-293T | C351(1.18); C96(0.87) | LDD1673 | [24] |
| LDCM0471 | CL78 | HEK-293T | C351(1.02); C96(0.86) | LDD1674 | [24] |
| LDCM0472 | CL79 | HEK-293T | C351(1.02); C96(1.03) | LDD1675 | [24] |
| LDCM0473 | CL8 | HEK-293T | C351(0.95); C96(0.62) | LDD1676 | [24] |
| LDCM0474 | CL80 | HEK-293T | C351(1.10); C96(0.83) | LDD1677 | [24] |
| LDCM0475 | CL81 | HEK-293T | C351(1.12); C96(1.05) | LDD1678 | [24] |
| LDCM0476 | CL82 | HEK-293T | C351(0.95); C96(0.97) | LDD1679 | [24] |
| LDCM0477 | CL83 | HEK-293T | C351(1.01); C96(1.12) | LDD1680 | [24] |
| LDCM0478 | CL84 | HEK-293T | C351(0.76); C96(0.92) | LDD1681 | [24] |
| LDCM0479 | CL85 | HEK-293T | C351(0.96); C96(0.69) | LDD1682 | [24] |
| LDCM0480 | CL86 | HEK-293T | C351(1.11); C96(0.88) | LDD1683 | [24] |
| LDCM0481 | CL87 | HEK-293T | C351(1.09); C96(0.94) | LDD1684 | [24] |
| LDCM0482 | CL88 | HEK-293T | C351(0.90); C96(1.02) | LDD1685 | [24] |
| LDCM0483 | CL89 | HEK-293T | C351(1.06); C96(0.98) | LDD1686 | [24] |
| LDCM0484 | CL9 | HEK-293T | C351(0.96); C96(1.06) | LDD1687 | [24] |
| LDCM0485 | CL90 | HEK-293T | C351(0.99); C96(0.72) | LDD1688 | [24] |
| LDCM0486 | CL91 | HEK-293T | C351(1.23); C96(1.05) | LDD1689 | [24] |
| LDCM0487 | CL92 | HEK-293T | C351(0.97); C96(0.83) | LDD1690 | [24] |
| LDCM0488 | CL93 | HEK-293T | C351(0.97); C96(1.03) | LDD1691 | [24] |
| LDCM0489 | CL94 | HEK-293T | C351(0.92); C96(0.94) | LDD1692 | [24] |
| LDCM0490 | CL95 | HEK-293T | C351(1.16); C96(0.90) | LDD1693 | [24] |
| LDCM0491 | CL96 | HEK-293T | C351(0.79); C96(1.09) | LDD1694 | [24] |
| LDCM0492 | CL97 | HEK-293T | C351(0.91); C96(0.84) | LDD1695 | [24] |
| LDCM0493 | CL98 | HEK-293T | C351(1.12); C96(1.09) | LDD1696 | [24] |
| LDCM0494 | CL99 | HEK-293T | C351(1.09); C96(1.00) | LDD1697 | [24] |
| LDCM0634 | CY-0357 | Hep-G2 | C351(1.50); C351(0.65) | LDD2228 | [12] |
| LDCM0028 | Dobutamine | HEK-293T | 9.24 | LDD0180 | [15] |
| LDCM0027 | Dopamine | HEK-293T | 25.77 | LDD0179 | [15] |
| LDCM0495 | E2913 | HEK-293T | C351(1.05); C96(1.05) | LDD1698 | [24] |
| LDCM0031 | Epigallocatechin gallate | HEK-293T | 20.17 | LDD0183 | [15] |
| LDCM0625 | F8 | Ramos | C351(1.66) | LDD2187 | [26] |
| LDCM0572 | Fragment10 | Ramos | C351(2.27) | LDD2189 | [26] |
| LDCM0573 | Fragment11 | Ramos | C351(0.60); C96(1.06) | LDD2190 | [26] |
| LDCM0574 | Fragment12 | Ramos | C351(0.85) | LDD2191 | [26] |
| LDCM0575 | Fragment13 | Ramos | C351(1.17) | LDD2192 | [26] |
| LDCM0576 | Fragment14 | Ramos | C351(0.93); C96(0.86) | LDD2193 | [26] |
| LDCM0579 | Fragment20 | Ramos | C351(1.17) | LDD2194 | [26] |
| LDCM0580 | Fragment21 | Ramos | C351(0.81) | LDD2195 | [26] |
| LDCM0582 | Fragment23 | Ramos | C351(1.73) | LDD2196 | [26] |
| LDCM0578 | Fragment27 | Ramos | C351(1.29) | LDD2197 | [26] |
| LDCM0586 | Fragment28 | Ramos | C351(0.97); C96(1.24) | LDD2198 | [26] |
| LDCM0588 | Fragment30 | Ramos | C351(1.51) | LDD2199 | [26] |
| LDCM0589 | Fragment31 | Ramos | C351(1.22) | LDD2200 | [26] |
| LDCM0590 | Fragment32 | Ramos | C351(2.00) | LDD2201 | [26] |
| LDCM0468 | Fragment33 | HEK-293T | C351(1.07); C96(0.88) | LDD1671 | [24] |
| LDCM0596 | Fragment38 | Ramos | C351(1.61) | LDD2203 | [26] |
| LDCM0566 | Fragment4 | Ramos | C351(0.79); C96(0.60) | LDD2184 | [26] |
| LDCM0427 | Fragment51 | HEK-293T | C351(1.02); C96(1.13) | LDD1631 | [24] |
| LDCM0610 | Fragment52 | Ramos | C351(1.18) | LDD2204 | [26] |
| LDCM0614 | Fragment56 | Ramos | C351(1.26) | LDD2205 | [26] |
| LDCM0569 | Fragment7 | Ramos | C351(0.90); C96(0.62) | LDD2186 | [26] |
| LDCM0571 | Fragment9 | Ramos | C351(1.27) | LDD2188 | [26] |
| LDCM0116 | HHS-0101 | DM93 | Y183(0.81); Y311(1.22); Y175(1.68) | LDD0264 | [10] |
| LDCM0117 | HHS-0201 | DM93 | Y183(0.49); Y311(1.51); Y175(3.66) | LDD0265 | [10] |
| LDCM0118 | HHS-0301 | DM93 | Y183(0.39) | LDD0266 | [10] |
| LDCM0119 | HHS-0401 | DM93 | Y183(0.81); Y175(0.89) | LDD0267 | [10] |
| LDCM0120 | HHS-0701 | DM93 | Y183(0.56); Y175(1.44) | LDD0268 | [10] |
| LDCM0022 | KB02 | HEK-293T | C96(1.12) | LDD1492 | [24] |
| LDCM0023 | KB03 | HEK-293T | C96(1.05) | LDD1497 | [24] |
| LDCM0024 | KB05 | IGR37 | C96(0.72) | LDD3314 | [6] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C351(0.94) | LDD2121 | [7] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C351(0.68) | LDD2150 | [7] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C351(0.88); C351(0.40) | LDD2206 | [27] |
| LDCM0032 | Oleacein | HEK-293T | 11.17 | LDD0184 | [15] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C351(0.90); C351(0.72) | LDD2207 | [27] |
| LDCM0131 | RA190 | MM1.R | C96(1.33); C351(1.20) | LDD0304 | [28] |
| LDCM0090 | Rapamycin | JHH-7 | 4.02 | LDD0213 | [23] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Alpha-synuclein (SNCA) | Synuclein family | P37840 | |||
Other
References
