Details of the Target
General Information of Target
| Target ID | LDTP06902 | |||||
|---|---|---|---|---|---|---|
| Target Name | mRNA export factor GLE1 (GLE1) | |||||
| Gene Name | GLE1 | |||||
| Gene ID | 2733 | |||||
| Synonyms |
GLE1L; mRNA export factor GLE1; hGLE1; GLE1 RNA export mediator; GLE1-like protein; Nucleoporin GLE1 |
|||||
| 3D Structure | ||||||
| Sequence |
MPSEGRCWETLKALRSSDKGRLCYYRDWLLRREDVLEECMSLPKLSSYSGWVVEHVLPHM
QENQPLSETSPSSTSASALDQPSFVPKSPDASSAFSPASPATPNGTKGKDESQHTESMVL QSSRGIKVEGCVRMYELVHRMKGTEGLRLWQEEQERKVQALSEMASEQLKRFDEWKELKQ HKEFQDLREVMEKSSREALGHQEKLKAEHRHRAKILNLKLREAEQQRVKQAEQERLRKEE GQIRLRALYALQEEMLQLSQQLDASEQHKALLKVDLAAFQTRGNQLCSLISGIIRASSES SYPTAESQAEAERALREMRDLLMNLGQEITRACEDKRRQDEEEAQVKLQEAQMQQGPEAH KEPPAPSQGPGGKQNEDLQVKVQDITMQWYQQLQDASMQCVLTFEGLTNSKDSQAKKIKM DLQKAATIPVSQISTIAGSKLKEIFDKIHSLLSGKPVQSGGRSVSVTLNPQGLDFVQYKL AEKFVKQGEEEVASHHEAAFPIAVVASGIWELHPRVGDLILAHLHKKCPYSVPFYPTFKE GMALEDYQRMLGYQVKDSKVEQQDNFLKRMSGMIRLYAAIIQLRWPYGNRQEIHPHGLNH GWRWLAQILNMEPLSDVTATLLFDFLEVCGNALMKQYQVQFWKMLILIKEDYFPRIEAIT SSGQMGSFIRLKQFLEKCLQHKDIPVPKGFLTSSFWRS |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
GLE1 family
|
|||||
| Subcellular location |
Cytoplasm; Nucleus
|
|||||
| Function | Required for the export of mRNAs containing poly(A) tails from the nucleus into the cytoplasm. May be involved in the terminal step of the mRNA transport through the nuclear pore complex (NPC). | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| A549 | SNV: p.A231V | DBIA Probe Info | |||
| COLO792 | SNV: p.S83L | DBIA Probe Info | |||
| COLO800 | SNV: p.R196K | DBIA Probe Info | |||
| MFE319 | SNV: p.A632V | DBIA Probe Info | |||
| MOLT4 | SNV: p.P536R | IA-alkyne Probe Info | |||
| NALM6 | SNV: p.T386A | DBIA Probe Info | |||
| REH | SNV: p.V492M | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
13.15 | LDD0402 | [1] | |
|
DA-P2 Probe Info |
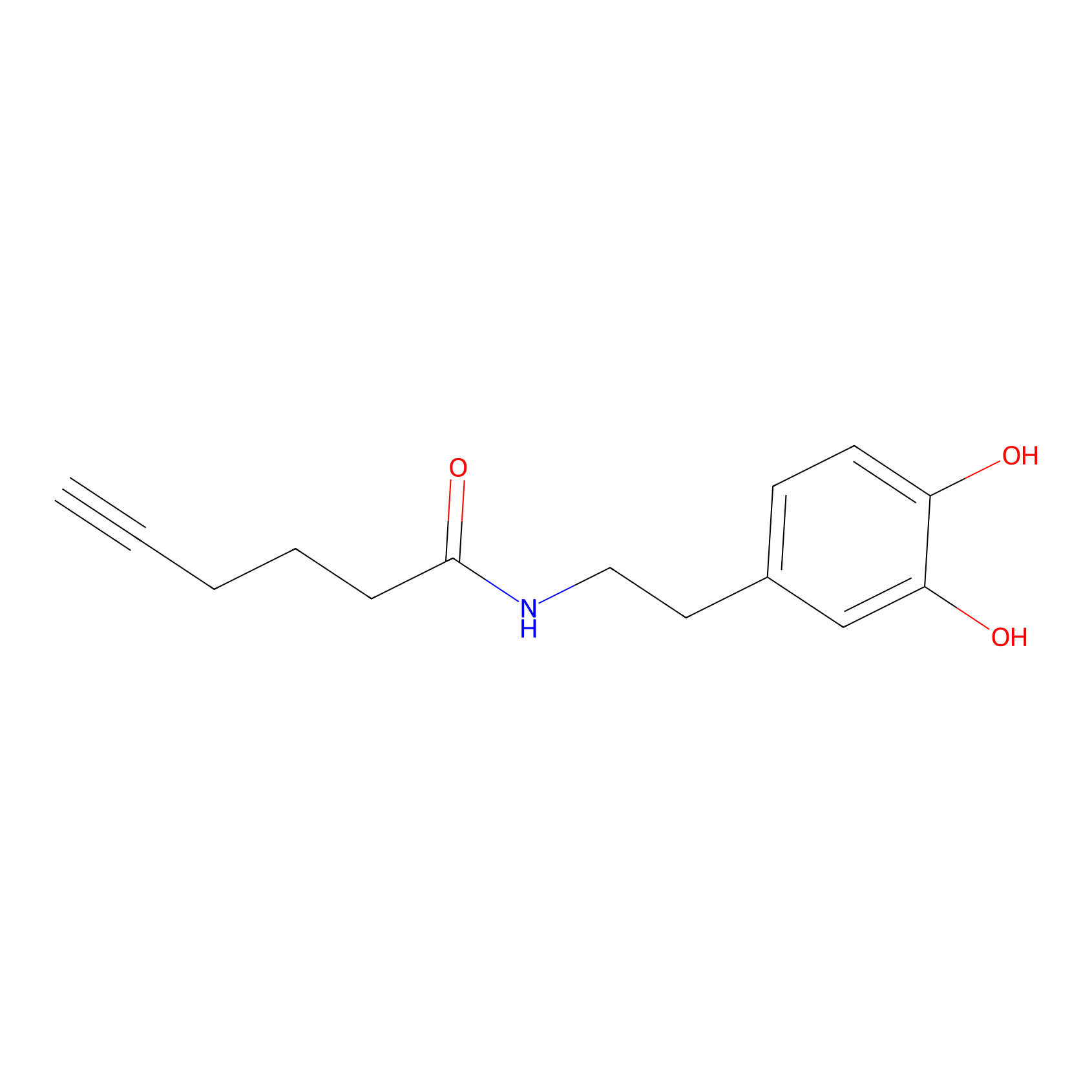 |
8.00 | LDD0348 | [2] | |
|
HRP Probe Info |
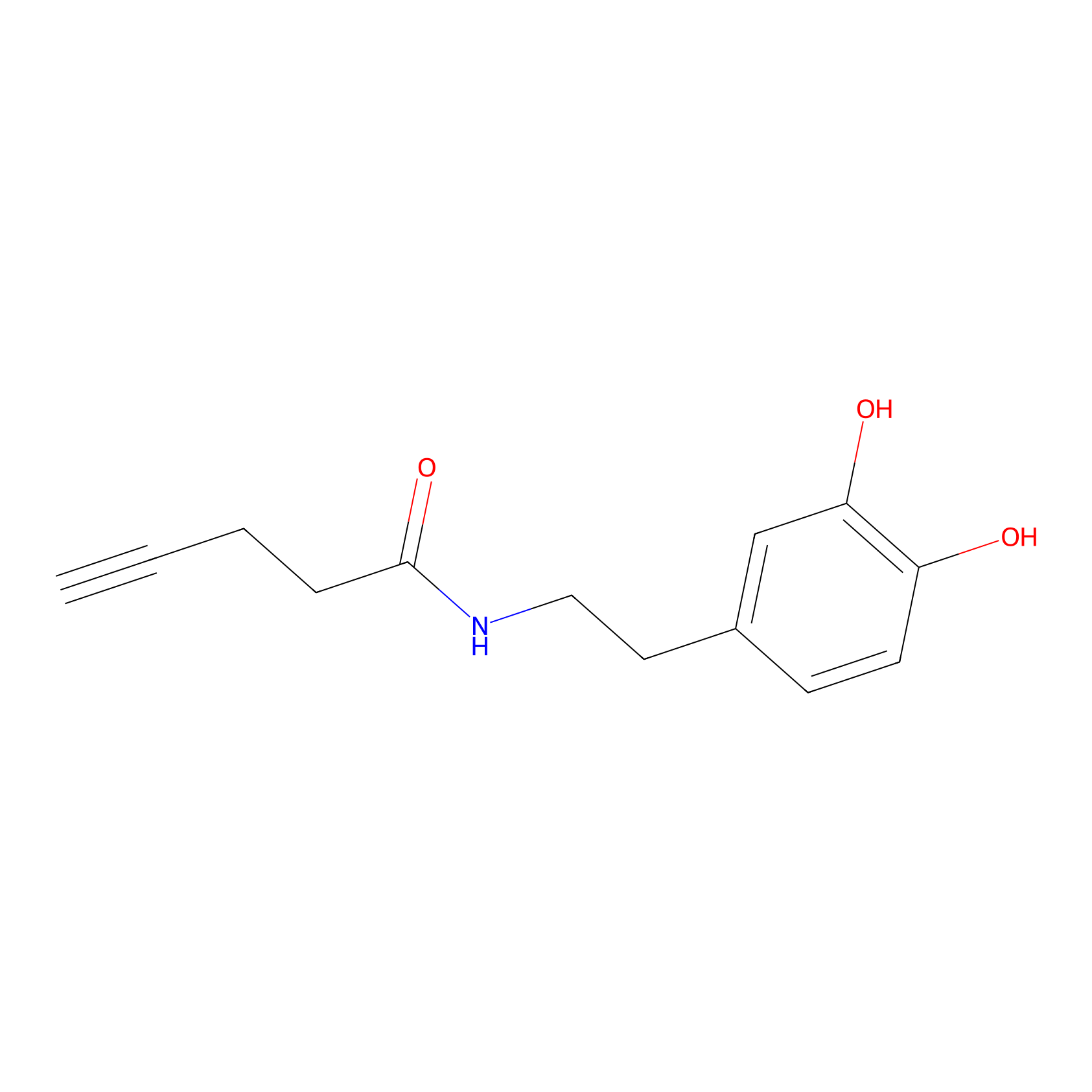 |
7.18 | LDD0347 | [2] | |
|
STPyne Probe Info |
 |
K204(5.76); K672(0.63) | LDD0277 | [3] | |
|
Probe 1 Probe Info |
 |
Y302(8.88) | LDD3495 | [4] | |
|
BTD Probe Info |
 |
C333(2.43) | LDD2100 | [5] | |
|
DA-P3 Probe Info |
 |
11.19 | LDD0179 | [2] | |
|
AHL-Pu-1 Probe Info |
 |
C39(2.28) | LDD0169 | [6] | |
|
DBIA Probe Info |
 |
C287(1.03) | LDD0078 | [7] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [8] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [8] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [8] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [9] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2224 | [9] | |
|
TFBX Probe Info |
 |
C528(0.00); C39(0.00) | LDD0027 | [10] | |
|
WYneN Probe Info |
 |
C528(0.00); C7(0.00) | LDD0021 | [11] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [11] | |
|
Phosphinate-6 Probe Info |
 |
C528(0.00); C7(0.00); C131(0.00) | LDD0018 | [12] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [13] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [13] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [14] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C333(1.20) | LDD2117 | [5] |
| LDCM0025 | 4SU-RNA | HEK-293T | C39(5.52) | LDD0371 | [6] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C39(2.28) | LDD0169 | [6] |
| LDCM0214 | AC1 | HCT 116 | C287(1.18); C528(1.02) | LDD0531 | [7] |
| LDCM0215 | AC10 | HCT 116 | C287(1.16); C528(0.95) | LDD0532 | [7] |
| LDCM0216 | AC100 | HCT 116 | C528(0.73) | LDD0533 | [7] |
| LDCM0217 | AC101 | HCT 116 | C528(0.74) | LDD0534 | [7] |
| LDCM0218 | AC102 | HCT 116 | C528(0.77) | LDD0535 | [7] |
| LDCM0219 | AC103 | HCT 116 | C528(0.79) | LDD0536 | [7] |
| LDCM0220 | AC104 | HCT 116 | C528(0.81) | LDD0537 | [7] |
| LDCM0221 | AC105 | HCT 116 | C528(0.84) | LDD0538 | [7] |
| LDCM0222 | AC106 | HCT 116 | C528(0.79) | LDD0539 | [7] |
| LDCM0223 | AC107 | HCT 116 | C528(0.86) | LDD0540 | [7] |
| LDCM0224 | AC108 | HCT 116 | C528(0.75) | LDD0541 | [7] |
| LDCM0225 | AC109 | HCT 116 | C528(0.84) | LDD0542 | [7] |
| LDCM0226 | AC11 | HCT 116 | C287(1.22); C528(0.93) | LDD0543 | [7] |
| LDCM0227 | AC110 | HCT 116 | C528(0.78) | LDD0544 | [7] |
| LDCM0228 | AC111 | HCT 116 | C528(0.75) | LDD0545 | [7] |
| LDCM0229 | AC112 | HCT 116 | C528(0.77) | LDD0546 | [7] |
| LDCM0230 | AC113 | HCT 116 | C287(0.94); C528(1.03) | LDD0547 | [7] |
| LDCM0231 | AC114 | HCT 116 | C287(1.13); C528(0.97) | LDD0548 | [7] |
| LDCM0232 | AC115 | HCT 116 | C287(1.18); C528(1.03) | LDD0549 | [7] |
| LDCM0233 | AC116 | HCT 116 | C287(1.36); C528(0.88) | LDD0550 | [7] |
| LDCM0234 | AC117 | HCT 116 | C287(1.19); C528(1.01) | LDD0551 | [7] |
| LDCM0235 | AC118 | HCT 116 | C287(1.10); C528(1.02) | LDD0552 | [7] |
| LDCM0236 | AC119 | HCT 116 | C287(1.13); C528(0.90) | LDD0553 | [7] |
| LDCM0237 | AC12 | HCT 116 | C287(1.06); C528(0.98) | LDD0554 | [7] |
| LDCM0238 | AC120 | HCT 116 | C287(1.15); C528(1.08) | LDD0555 | [7] |
| LDCM0239 | AC121 | HCT 116 | C287(1.14); C528(1.01) | LDD0556 | [7] |
| LDCM0240 | AC122 | HCT 116 | C287(1.13); C528(1.01) | LDD0557 | [7] |
| LDCM0241 | AC123 | HCT 116 | C287(1.32); C528(1.10) | LDD0558 | [7] |
| LDCM0242 | AC124 | HCT 116 | C287(1.04); C528(1.01) | LDD0559 | [7] |
| LDCM0243 | AC125 | HCT 116 | C287(1.04); C528(1.09) | LDD0560 | [7] |
| LDCM0244 | AC126 | HCT 116 | C287(1.35); C528(1.05) | LDD0561 | [7] |
| LDCM0245 | AC127 | HCT 116 | C287(1.21); C528(1.08) | LDD0562 | [7] |
| LDCM0246 | AC128 | HCT 116 | C287(1.57); C528(1.71) | LDD0563 | [7] |
| LDCM0247 | AC129 | HCT 116 | C287(0.99); C528(1.57) | LDD0564 | [7] |
| LDCM0249 | AC130 | HCT 116 | C287(1.20); C528(1.30) | LDD0566 | [7] |
| LDCM0250 | AC131 | HCT 116 | C287(1.12); C528(1.28) | LDD0567 | [7] |
| LDCM0251 | AC132 | HCT 116 | C287(1.51); C528(1.09) | LDD0568 | [7] |
| LDCM0252 | AC133 | HCT 116 | C287(1.15); C528(1.19) | LDD0569 | [7] |
| LDCM0253 | AC134 | HCT 116 | C287(1.27); C528(1.26) | LDD0570 | [7] |
| LDCM0254 | AC135 | HCT 116 | C287(1.41); C528(1.26) | LDD0571 | [7] |
| LDCM0255 | AC136 | HCT 116 | C287(1.18); C528(1.20) | LDD0572 | [7] |
| LDCM0256 | AC137 | HCT 116 | C287(1.14); C528(1.28) | LDD0573 | [7] |
| LDCM0257 | AC138 | HCT 116 | C287(1.31); C528(1.63) | LDD0574 | [7] |
| LDCM0258 | AC139 | HCT 116 | C287(1.26); C528(1.11) | LDD0575 | [7] |
| LDCM0259 | AC14 | HCT 116 | C287(1.29); C528(1.01) | LDD0576 | [7] |
| LDCM0260 | AC140 | HCT 116 | C287(1.28); C528(1.35) | LDD0577 | [7] |
| LDCM0261 | AC141 | HCT 116 | C287(1.39); C528(1.29) | LDD0578 | [7] |
| LDCM0262 | AC142 | HCT 116 | C287(1.22); C528(1.22) | LDD0579 | [7] |
| LDCM0263 | AC143 | HCT 116 | C287(1.03); C528(1.15) | LDD0580 | [7] |
| LDCM0264 | AC144 | HCT 116 | C528(1.01); C287(1.01) | LDD0581 | [7] |
| LDCM0265 | AC145 | HCT 116 | C528(1.04); C287(1.14) | LDD0582 | [7] |
| LDCM0266 | AC146 | HCT 116 | C528(1.12); C287(1.19) | LDD0583 | [7] |
| LDCM0267 | AC147 | HCT 116 | C528(1.05); C287(1.16) | LDD0584 | [7] |
| LDCM0268 | AC148 | HCT 116 | C528(1.21); C287(1.51) | LDD0585 | [7] |
| LDCM0269 | AC149 | HCT 116 | C528(1.09); C287(1.15) | LDD0586 | [7] |
| LDCM0270 | AC15 | HCT 116 | C528(0.99); C287(1.21) | LDD0587 | [7] |
| LDCM0271 | AC150 | HCT 116 | C528(1.03); C287(1.09) | LDD0588 | [7] |
| LDCM0272 | AC151 | HCT 116 | C528(0.94); C287(1.04) | LDD0589 | [7] |
| LDCM0273 | AC152 | HCT 116 | C287(1.09); C528(1.22) | LDD0590 | [7] |
| LDCM0274 | AC153 | HCT 116 | C528(1.36); C287(1.63) | LDD0591 | [7] |
| LDCM0621 | AC154 | HCT 116 | C287(1.15); C528(1.07) | LDD2158 | [7] |
| LDCM0622 | AC155 | HCT 116 | C287(1.10); C528(1.11) | LDD2159 | [7] |
| LDCM0623 | AC156 | HCT 116 | C287(0.88); C528(1.04) | LDD2160 | [7] |
| LDCM0624 | AC157 | HCT 116 | C287(0.98); C528(1.07) | LDD2161 | [7] |
| LDCM0276 | AC17 | HCT 116 | C287(0.93); C528(1.06) | LDD0593 | [7] |
| LDCM0277 | AC18 | HCT 116 | C287(1.02); C528(1.10) | LDD0594 | [7] |
| LDCM0278 | AC19 | HCT 116 | C287(0.88); C528(0.91) | LDD0595 | [7] |
| LDCM0279 | AC2 | HCT 116 | C528(0.96); C287(1.20) | LDD0596 | [7] |
| LDCM0280 | AC20 | HCT 116 | C287(0.89); C528(1.09) | LDD0597 | [7] |
| LDCM0281 | AC21 | HCT 116 | C287(0.92); C528(1.07) | LDD0598 | [7] |
| LDCM0282 | AC22 | HCT 116 | C287(0.84); C528(1.03) | LDD0599 | [7] |
| LDCM0283 | AC23 | HCT 116 | C287(0.82); C528(1.00) | LDD0600 | [7] |
| LDCM0284 | AC24 | HCT 116 | C287(0.85); C528(1.03) | LDD0601 | [7] |
| LDCM0285 | AC25 | HCT 116 | C528(0.71); C287(0.97) | LDD0602 | [7] |
| LDCM0286 | AC26 | HCT 116 | C528(0.82); C287(1.10) | LDD0603 | [7] |
| LDCM0287 | AC27 | HCT 116 | C528(0.88); C287(1.05) | LDD0604 | [7] |
| LDCM0288 | AC28 | HCT 116 | C528(0.93); C287(1.00) | LDD0605 | [7] |
| LDCM0289 | AC29 | HCT 116 | C528(0.79); C287(1.05) | LDD0606 | [7] |
| LDCM0290 | AC3 | HCT 116 | C528(0.97); C287(1.18) | LDD0607 | [7] |
| LDCM0291 | AC30 | HCT 116 | C528(0.77); C287(1.06) | LDD0608 | [7] |
| LDCM0292 | AC31 | HCT 116 | C528(0.78); C287(1.05) | LDD0609 | [7] |
| LDCM0293 | AC32 | HCT 116 | C528(0.84); C287(1.25) | LDD0610 | [7] |
| LDCM0294 | AC33 | HCT 116 | C528(0.83); C287(1.15) | LDD0611 | [7] |
| LDCM0295 | AC34 | HCT 116 | C528(1.05); C287(1.25) | LDD0612 | [7] |
| LDCM0296 | AC35 | HCT 116 | C287(0.91); C528(1.16) | LDD0613 | [7] |
| LDCM0297 | AC36 | HCT 116 | C287(0.95); C528(1.12) | LDD0614 | [7] |
| LDCM0298 | AC37 | HCT 116 | C287(1.00); C528(1.04) | LDD0615 | [7] |
| LDCM0299 | AC38 | HCT 116 | C287(0.99); C528(1.18) | LDD0616 | [7] |
| LDCM0300 | AC39 | HCT 116 | C287(1.05); C528(1.11) | LDD0617 | [7] |
| LDCM0301 | AC4 | HCT 116 | C528(1.11); C287(1.27) | LDD0618 | [7] |
| LDCM0302 | AC40 | HCT 116 | C528(1.10); C287(1.10) | LDD0619 | [7] |
| LDCM0303 | AC41 | HCT 116 | C528(1.06); C287(1.11) | LDD0620 | [7] |
| LDCM0304 | AC42 | HCT 116 | C528(1.07); C287(1.12) | LDD0621 | [7] |
| LDCM0305 | AC43 | HCT 116 | C528(0.96); C287(1.16) | LDD0622 | [7] |
| LDCM0306 | AC44 | HCT 116 | C528(0.94); C287(1.12) | LDD0623 | [7] |
| LDCM0307 | AC45 | HCT 116 | C528(0.97); C287(1.24) | LDD0624 | [7] |
| LDCM0308 | AC46 | HCT 116 | C528(0.96); C287(1.05) | LDD0625 | [7] |
| LDCM0309 | AC47 | HCT 116 | C528(1.02); C287(1.04) | LDD0626 | [7] |
| LDCM0310 | AC48 | HCT 116 | C528(0.92); C287(1.19) | LDD0627 | [7] |
| LDCM0311 | AC49 | HCT 116 | C528(0.95); C287(1.34) | LDD0628 | [7] |
| LDCM0312 | AC5 | HCT 116 | C528(1.14); C287(1.27) | LDD0629 | [7] |
| LDCM0313 | AC50 | HCT 116 | C528(0.97); C287(1.21) | LDD0630 | [7] |
| LDCM0314 | AC51 | HCT 116 | C528(0.91); C287(0.97) | LDD0631 | [7] |
| LDCM0315 | AC52 | HCT 116 | C528(0.91); C287(1.04) | LDD0632 | [7] |
| LDCM0316 | AC53 | HCT 116 | C528(1.01); C287(1.16) | LDD0633 | [7] |
| LDCM0317 | AC54 | HCT 116 | C528(1.03); C287(1.13) | LDD0634 | [7] |
| LDCM0318 | AC55 | HCT 116 | C528(0.94); C287(1.23) | LDD0635 | [7] |
| LDCM0319 | AC56 | HCT 116 | C528(1.00); C287(1.50) | LDD0636 | [7] |
| LDCM0320 | AC57 | HCT 116 | C528(0.94) | LDD0637 | [7] |
| LDCM0321 | AC58 | HCT 116 | C528(1.03) | LDD0638 | [7] |
| LDCM0322 | AC59 | HCT 116 | C528(0.91) | LDD0639 | [7] |
| LDCM0323 | AC6 | HCT 116 | C528(0.98); C287(1.20) | LDD0640 | [7] |
| LDCM0324 | AC60 | HCT 116 | C528(0.91) | LDD0641 | [7] |
| LDCM0325 | AC61 | HCT 116 | C528(0.76) | LDD0642 | [7] |
| LDCM0326 | AC62 | HCT 116 | C528(0.91) | LDD0643 | [7] |
| LDCM0327 | AC63 | HCT 116 | C528(0.89) | LDD0644 | [7] |
| LDCM0328 | AC64 | HCT 116 | C528(0.97) | LDD0645 | [7] |
| LDCM0329 | AC65 | HCT 116 | C528(0.84) | LDD0646 | [7] |
| LDCM0330 | AC66 | HCT 116 | C528(0.86) | LDD0647 | [7] |
| LDCM0331 | AC67 | HCT 116 | C528(0.91) | LDD0648 | [7] |
| LDCM0332 | AC68 | HCT 116 | C287(0.98); C528(1.16) | LDD0649 | [7] |
| LDCM0333 | AC69 | HCT 116 | C287(1.00); C528(1.27) | LDD0650 | [7] |
| LDCM0334 | AC7 | HCT 116 | C528(0.99); C287(1.07) | LDD0651 | [7] |
| LDCM0335 | AC70 | HCT 116 | C287(0.94); C528(1.14) | LDD0652 | [7] |
| LDCM0336 | AC71 | HCT 116 | C287(0.97); C528(1.24) | LDD0653 | [7] |
| LDCM0337 | AC72 | HCT 116 | C287(1.03); C528(1.18) | LDD0654 | [7] |
| LDCM0338 | AC73 | HCT 116 | C287(1.19); C528(1.42) | LDD0655 | [7] |
| LDCM0339 | AC74 | HCT 116 | C287(1.15); C528(1.51) | LDD0656 | [7] |
| LDCM0340 | AC75 | HCT 116 | C287(1.33); C528(1.45) | LDD0657 | [7] |
| LDCM0341 | AC76 | HCT 116 | C287(1.04); C528(1.21) | LDD0658 | [7] |
| LDCM0342 | AC77 | HCT 116 | C287(0.99); C528(1.18) | LDD0659 | [7] |
| LDCM0343 | AC78 | HCT 116 | C287(0.98); C528(1.08) | LDD0660 | [7] |
| LDCM0344 | AC79 | HCT 116 | C287(1.08); C528(1.18) | LDD0661 | [7] |
| LDCM0345 | AC8 | HCT 116 | C528(1.06); C287(1.12) | LDD0662 | [7] |
| LDCM0346 | AC80 | HCT 116 | C287(1.00); C528(1.14) | LDD0663 | [7] |
| LDCM0347 | AC81 | HCT 116 | C287(0.82); C528(1.15) | LDD0664 | [7] |
| LDCM0348 | AC82 | HCT 116 | C287(1.22); C528(1.60) | LDD0665 | [7] |
| LDCM0349 | AC83 | HCT 116 | C528(0.94); C287(1.27) | LDD0666 | [7] |
| LDCM0350 | AC84 | HCT 116 | C528(0.99); C287(1.18) | LDD0667 | [7] |
| LDCM0351 | AC85 | HCT 116 | C528(0.83); C287(0.93) | LDD0668 | [7] |
| LDCM0352 | AC86 | HCT 116 | C528(0.73); C287(1.11) | LDD0669 | [7] |
| LDCM0353 | AC87 | HCT 116 | C528(0.80); C287(0.96) | LDD0670 | [7] |
| LDCM0354 | AC88 | HCT 116 | C528(0.77); C287(0.99) | LDD0671 | [7] |
| LDCM0355 | AC89 | HCT 116 | C528(0.77); C287(1.12) | LDD0672 | [7] |
| LDCM0357 | AC90 | HCT 116 | C528(0.69); C287(1.00) | LDD0674 | [7] |
| LDCM0358 | AC91 | HCT 116 | C528(0.92); C287(1.22) | LDD0675 | [7] |
| LDCM0359 | AC92 | HCT 116 | C528(0.87); C287(1.23) | LDD0676 | [7] |
| LDCM0360 | AC93 | HCT 116 | C528(0.86); C287(0.93) | LDD0677 | [7] |
| LDCM0361 | AC94 | HCT 116 | C287(0.98); C528(1.06) | LDD0678 | [7] |
| LDCM0362 | AC95 | HCT 116 | C528(0.79); C287(1.02) | LDD0679 | [7] |
| LDCM0363 | AC96 | HCT 116 | C528(0.98); C287(1.01) | LDD0680 | [7] |
| LDCM0364 | AC97 | HCT 116 | C528(0.77); C287(1.06) | LDD0681 | [7] |
| LDCM0365 | AC98 | HCT 116 | C528(0.97) | LDD0682 | [7] |
| LDCM0366 | AC99 | HCT 116 | C528(0.78) | LDD0683 | [7] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C333(0.82) | LDD2113 | [5] |
| LDCM0248 | AKOS034007472 | HCT 116 | C287(1.02); C528(0.88) | LDD0565 | [7] |
| LDCM0356 | AKOS034007680 | HCT 116 | C528(0.90); C287(1.31) | LDD0673 | [7] |
| LDCM0275 | AKOS034007705 | HCT 116 | C528(1.09); C287(1.46) | LDD0592 | [7] |
| LDCM0156 | Aniline | NCI-H1299 | 11.47 | LDD0403 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C287(1.03) | LDD0078 | [7] |
| LDCM0087 | Capsaicin | HEK-293T | 6.80 | LDD0185 | [2] |
| LDCM0632 | CL-Sc | Hep-G2 | C39(20.00); C287(0.74) | LDD2227 | [9] |
| LDCM0367 | CL1 | HCT 116 | C287(0.91); C528(0.96) | LDD0684 | [7] |
| LDCM0368 | CL10 | HCT 116 | C528(0.91); C287(1.09) | LDD0685 | [7] |
| LDCM0369 | CL100 | HCT 116 | C528(1.02); C287(1.19) | LDD0686 | [7] |
| LDCM0370 | CL101 | HCT 116 | C528(0.88); C287(1.22) | LDD0687 | [7] |
| LDCM0371 | CL102 | HCT 116 | C528(1.04); C287(1.21) | LDD0688 | [7] |
| LDCM0372 | CL103 | HCT 116 | C528(0.89); C287(1.05) | LDD0689 | [7] |
| LDCM0373 | CL104 | HCT 116 | C287(0.93); C528(0.98) | LDD0690 | [7] |
| LDCM0374 | CL105 | HCT 116 | C528(1.01); C287(1.01) | LDD0691 | [7] |
| LDCM0375 | CL106 | HCT 116 | C287(1.10); C528(1.23) | LDD0692 | [7] |
| LDCM0376 | CL107 | HCT 116 | C287(1.01); C528(1.14) | LDD0693 | [7] |
| LDCM0377 | CL108 | HCT 116 | C287(1.15); C528(1.15) | LDD0694 | [7] |
| LDCM0378 | CL109 | HCT 116 | C287(0.94); C528(0.96) | LDD0695 | [7] |
| LDCM0379 | CL11 | HCT 116 | C528(0.98); C287(1.27) | LDD0696 | [7] |
| LDCM0380 | CL110 | HCT 116 | C528(0.94); C287(0.95) | LDD0697 | [7] |
| LDCM0381 | CL111 | HCT 116 | C287(0.85); C528(1.01) | LDD0698 | [7] |
| LDCM0382 | CL112 | HCT 116 | C528(0.90); C287(0.95) | LDD0699 | [7] |
| LDCM0383 | CL113 | HCT 116 | C528(0.85); C287(1.12) | LDD0700 | [7] |
| LDCM0384 | CL114 | HCT 116 | C528(0.69); C287(1.00) | LDD0701 | [7] |
| LDCM0385 | CL115 | HCT 116 | C528(0.80); C287(0.96) | LDD0702 | [7] |
| LDCM0386 | CL116 | HCT 116 | C528(0.91); C287(1.08) | LDD0703 | [7] |
| LDCM0387 | CL117 | HCT 116 | C528(1.53); C287(1.54) | LDD0704 | [7] |
| LDCM0388 | CL118 | HCT 116 | C528(1.04); C287(1.10) | LDD0705 | [7] |
| LDCM0389 | CL119 | HCT 116 | C528(0.99); C287(1.10) | LDD0706 | [7] |
| LDCM0390 | CL12 | HCT 116 | C528(0.99); C287(1.24) | LDD0707 | [7] |
| LDCM0391 | CL120 | HCT 116 | C528(0.93); C287(1.05) | LDD0708 | [7] |
| LDCM0392 | CL121 | HCT 116 | C528(0.57); C287(0.91) | LDD0709 | [7] |
| LDCM0393 | CL122 | HCT 116 | C528(1.07); C287(1.14) | LDD0710 | [7] |
| LDCM0394 | CL123 | HCT 116 | C528(0.91); C287(1.17) | LDD0711 | [7] |
| LDCM0395 | CL124 | HCT 116 | C528(0.90); C287(1.23) | LDD0712 | [7] |
| LDCM0396 | CL125 | HCT 116 | C528(0.94) | LDD0713 | [7] |
| LDCM0397 | CL126 | HCT 116 | C528(1.05) | LDD0714 | [7] |
| LDCM0398 | CL127 | HCT 116 | C528(0.82) | LDD0715 | [7] |
| LDCM0399 | CL128 | HCT 116 | C528(1.05) | LDD0716 | [7] |
| LDCM0400 | CL13 | HCT 116 | C528(1.00); C287(1.07) | LDD0717 | [7] |
| LDCM0401 | CL14 | HCT 116 | C528(0.92); C287(1.00) | LDD0718 | [7] |
| LDCM0402 | CL15 | HCT 116 | C528(1.02); C287(1.22) | LDD0719 | [7] |
| LDCM0403 | CL16 | HCT 116 | C528(0.77); C287(1.12) | LDD0720 | [7] |
| LDCM0404 | CL17 | HCT 116 | C528(1.08); C287(1.12) | LDD0721 | [7] |
| LDCM0405 | CL18 | HCT 116 | C528(0.89); C287(1.25) | LDD0722 | [7] |
| LDCM0406 | CL19 | HCT 116 | C528(0.91); C287(1.18) | LDD0723 | [7] |
| LDCM0407 | CL2 | HCT 116 | C528(0.90); C287(1.01) | LDD0724 | [7] |
| LDCM0408 | CL20 | HCT 116 | C528(0.88); C287(1.16) | LDD0725 | [7] |
| LDCM0409 | CL21 | HCT 116 | C528(1.04); C287(1.43) | LDD0726 | [7] |
| LDCM0410 | CL22 | HCT 116 | C528(1.15); C287(1.79) | LDD0727 | [7] |
| LDCM0411 | CL23 | HCT 116 | C528(0.92); C287(1.12) | LDD0728 | [7] |
| LDCM0412 | CL24 | HCT 116 | C528(0.82); C287(1.41) | LDD0729 | [7] |
| LDCM0413 | CL25 | HCT 116 | C287(1.32); C528(0.78) | LDD0730 | [7] |
| LDCM0414 | CL26 | HCT 116 | C287(1.07); C528(0.87) | LDD0731 | [7] |
| LDCM0415 | CL27 | HCT 116 | C287(1.13); C528(0.78) | LDD0732 | [7] |
| LDCM0416 | CL28 | HCT 116 | C287(1.17); C528(0.76) | LDD0733 | [7] |
| LDCM0417 | CL29 | HCT 116 | C287(1.22); C528(0.77) | LDD0734 | [7] |
| LDCM0418 | CL3 | HCT 116 | C287(1.02); C528(0.96) | LDD0735 | [7] |
| LDCM0419 | CL30 | HCT 116 | C287(1.00); C528(0.73) | LDD0736 | [7] |
| LDCM0420 | CL31 | HCT 116 | C287(0.97); C528(1.01) | LDD0737 | [7] |
| LDCM0421 | CL32 | HCT 116 | C287(1.05); C528(1.01) | LDD0738 | [7] |
| LDCM0422 | CL33 | HCT 116 | C287(0.99); C528(1.21) | LDD0739 | [7] |
| LDCM0423 | CL34 | HCT 116 | C287(1.08); C528(1.04) | LDD0740 | [7] |
| LDCM0424 | CL35 | HCT 116 | C287(0.99); C528(1.04) | LDD0741 | [7] |
| LDCM0425 | CL36 | HCT 116 | C287(0.97); C528(1.08) | LDD0742 | [7] |
| LDCM0426 | CL37 | HCT 116 | C287(1.11); C528(1.21) | LDD0743 | [7] |
| LDCM0428 | CL39 | HCT 116 | C287(0.97); C528(1.05) | LDD0745 | [7] |
| LDCM0429 | CL4 | HCT 116 | C287(0.98); C528(0.93) | LDD0746 | [7] |
| LDCM0430 | CL40 | HCT 116 | C287(0.94); C528(1.05) | LDD0747 | [7] |
| LDCM0431 | CL41 | HCT 116 | C287(1.11); C528(1.01) | LDD0748 | [7] |
| LDCM0432 | CL42 | HCT 116 | C287(1.35); C528(1.52) | LDD0749 | [7] |
| LDCM0433 | CL43 | HCT 116 | C287(1.12); C528(1.19) | LDD0750 | [7] |
| LDCM0434 | CL44 | HCT 116 | C287(0.99); C528(1.02) | LDD0751 | [7] |
| LDCM0435 | CL45 | HCT 116 | C287(1.13); C528(1.11) | LDD0752 | [7] |
| LDCM0436 | CL46 | HCT 116 | C287(0.79); C528(0.80) | LDD0753 | [7] |
| LDCM0437 | CL47 | HCT 116 | C287(0.87); C528(0.74) | LDD0754 | [7] |
| LDCM0438 | CL48 | HCT 116 | C287(0.85); C528(0.91) | LDD0755 | [7] |
| LDCM0439 | CL49 | HCT 116 | C287(0.83); C528(1.03) | LDD0756 | [7] |
| LDCM0440 | CL5 | HCT 116 | C287(1.07); C528(0.97) | LDD0757 | [7] |
| LDCM0441 | CL50 | HCT 116 | C287(0.78); C528(0.84) | LDD0758 | [7] |
| LDCM0442 | CL51 | HCT 116 | C287(0.84); C528(0.84) | LDD0759 | [7] |
| LDCM0443 | CL52 | HCT 116 | C287(0.83); C528(0.79) | LDD0760 | [7] |
| LDCM0444 | CL53 | HCT 116 | C287(0.77); C528(0.96) | LDD0761 | [7] |
| LDCM0445 | CL54 | HCT 116 | C287(0.78); C528(0.88) | LDD0762 | [7] |
| LDCM0446 | CL55 | HCT 116 | C287(0.80); C528(0.84) | LDD0763 | [7] |
| LDCM0447 | CL56 | HCT 116 | C287(0.92); C528(0.87) | LDD0764 | [7] |
| LDCM0448 | CL57 | HCT 116 | C287(0.79); C528(0.98) | LDD0765 | [7] |
| LDCM0449 | CL58 | HCT 116 | C287(0.85); C528(0.80) | LDD0766 | [7] |
| LDCM0450 | CL59 | HCT 116 | C287(0.93); C528(0.89) | LDD0767 | [7] |
| LDCM0451 | CL6 | HCT 116 | C287(1.10); C528(0.86) | LDD0768 | [7] |
| LDCM0452 | CL60 | HCT 116 | C287(0.94); C528(0.87) | LDD0769 | [7] |
| LDCM0453 | CL61 | HCT 116 | C287(1.02); C528(1.04) | LDD0770 | [7] |
| LDCM0454 | CL62 | HCT 116 | C287(1.26); C528(0.99) | LDD0771 | [7] |
| LDCM0455 | CL63 | HCT 116 | C287(1.20); C528(1.01) | LDD0772 | [7] |
| LDCM0456 | CL64 | HCT 116 | C287(1.14); C528(1.03) | LDD0773 | [7] |
| LDCM0457 | CL65 | HCT 116 | C287(1.19); C528(1.04) | LDD0774 | [7] |
| LDCM0458 | CL66 | HCT 116 | C287(1.53); C528(1.06) | LDD0775 | [7] |
| LDCM0459 | CL67 | HCT 116 | C287(1.27); C528(1.11) | LDD0776 | [7] |
| LDCM0460 | CL68 | HCT 116 | C287(1.12); C528(1.04) | LDD0777 | [7] |
| LDCM0461 | CL69 | HCT 116 | C287(1.03); C528(0.97) | LDD0778 | [7] |
| LDCM0462 | CL7 | HCT 116 | C287(1.13); C528(1.01) | LDD0779 | [7] |
| LDCM0463 | CL70 | HCT 116 | C287(1.21); C528(0.98) | LDD0780 | [7] |
| LDCM0464 | CL71 | HCT 116 | C287(1.13); C528(0.95) | LDD0781 | [7] |
| LDCM0465 | CL72 | HCT 116 | C287(1.06); C528(0.95) | LDD0782 | [7] |
| LDCM0466 | CL73 | HCT 116 | C287(1.10); C528(1.06) | LDD0783 | [7] |
| LDCM0467 | CL74 | HCT 116 | C287(1.10); C528(0.98) | LDD0784 | [7] |
| LDCM0469 | CL76 | HCT 116 | C287(1.18); C528(1.11) | LDD0786 | [7] |
| LDCM0470 | CL77 | HCT 116 | C287(1.05); C528(1.18) | LDD0787 | [7] |
| LDCM0471 | CL78 | HCT 116 | C287(1.03); C528(0.96) | LDD0788 | [7] |
| LDCM0472 | CL79 | HCT 116 | C287(1.16); C528(1.16) | LDD0789 | [7] |
| LDCM0473 | CL8 | HCT 116 | C287(1.53); C528(1.19) | LDD0790 | [7] |
| LDCM0474 | CL80 | HCT 116 | C287(1.12); C528(1.15) | LDD0791 | [7] |
| LDCM0475 | CL81 | HCT 116 | C287(1.06); C528(1.09) | LDD0792 | [7] |
| LDCM0476 | CL82 | HCT 116 | C287(1.45); C528(1.18) | LDD0793 | [7] |
| LDCM0477 | CL83 | HCT 116 | C287(1.31); C528(1.13) | LDD0794 | [7] |
| LDCM0478 | CL84 | HCT 116 | C287(1.66); C528(1.21) | LDD0795 | [7] |
| LDCM0479 | CL85 | HCT 116 | C287(1.14); C528(1.08) | LDD0796 | [7] |
| LDCM0480 | CL86 | HCT 116 | C287(1.12); C528(1.12) | LDD0797 | [7] |
| LDCM0481 | CL87 | HCT 116 | C287(1.18); C528(0.98) | LDD0798 | [7] |
| LDCM0482 | CL88 | HCT 116 | C287(1.34); C528(1.13) | LDD0799 | [7] |
| LDCM0483 | CL89 | HCT 116 | C287(1.95); C528(1.24) | LDD0800 | [7] |
| LDCM0484 | CL9 | HCT 116 | C287(1.01); C528(0.93) | LDD0801 | [7] |
| LDCM0485 | CL90 | HCT 116 | C287(1.14); C528(1.17) | LDD0802 | [7] |
| LDCM0486 | CL91 | HCT 116 | C287(1.10); C528(0.98) | LDD0803 | [7] |
| LDCM0487 | CL92 | HCT 116 | C287(1.08); C528(1.07) | LDD0804 | [7] |
| LDCM0488 | CL93 | HCT 116 | C287(1.09); C528(1.04) | LDD0805 | [7] |
| LDCM0489 | CL94 | HCT 116 | C287(1.20); C528(0.99) | LDD0806 | [7] |
| LDCM0490 | CL95 | HCT 116 | C287(1.21); C528(1.06) | LDD0807 | [7] |
| LDCM0491 | CL96 | HCT 116 | C287(1.10); C528(0.95) | LDD0808 | [7] |
| LDCM0492 | CL97 | HCT 116 | C287(1.10); C528(0.95) | LDD0809 | [7] |
| LDCM0493 | CL98 | HCT 116 | C287(1.21); C528(0.99) | LDD0810 | [7] |
| LDCM0494 | CL99 | HCT 116 | C287(1.25); C528(0.93) | LDD0811 | [7] |
| LDCM0027 | Dopamine | HEK-293T | 11.19 | LDD0179 | [2] |
| LDCM0495 | E2913 | HEK-293T | C528(1.69); C287(0.87); C400(0.84); C131(1.05) | LDD1698 | [15] |
| LDCM0625 | F8 | Ramos | C131(1.04); C528(1.01); C287(0.21) | LDD2187 | [16] |
| LDCM0572 | Fragment10 | Ramos | C528(1.03); C287(0.41) | LDD2189 | [16] |
| LDCM0573 | Fragment11 | Ramos | C131(8.69); C528(1.26); C287(0.82) | LDD2190 | [16] |
| LDCM0575 | Fragment13 | Ramos | C287(0.28) | LDD2192 | [16] |
| LDCM0576 | Fragment14 | Ramos | C528(1.48) | LDD2193 | [16] |
| LDCM0580 | Fragment21 | Ramos | C528(0.72); C287(0.62) | LDD2195 | [16] |
| LDCM0582 | Fragment23 | Ramos | C131(1.02); C528(0.71); C287(0.49) | LDD2196 | [16] |
| LDCM0578 | Fragment27 | Ramos | C131(2.10); C528(1.04); C287(0.30) | LDD2197 | [16] |
| LDCM0586 | Fragment28 | Ramos | C131(0.73); C528(1.86); C287(0.64) | LDD2198 | [16] |
| LDCM0588 | Fragment30 | Ramos | C131(1.69); C287(0.59) | LDD2199 | [16] |
| LDCM0589 | Fragment31 | Ramos | C131(1.50); C287(0.49) | LDD2200 | [16] |
| LDCM0590 | Fragment32 | Ramos | C528(1.20) | LDD2201 | [16] |
| LDCM0468 | Fragment33 | HCT 116 | C287(1.21); C528(0.93) | LDD0785 | [7] |
| LDCM0596 | Fragment38 | Ramos | C528(1.81); C287(0.80) | LDD2203 | [16] |
| LDCM0427 | Fragment51 | HCT 116 | C287(1.25); C528(1.28) | LDD0744 | [7] |
| LDCM0610 | Fragment52 | Ramos | C287(0.18) | LDD2204 | [16] |
| LDCM0614 | Fragment56 | Ramos | C528(0.49); C287(0.21) | LDD2205 | [16] |
| LDCM0569 | Fragment7 | Ramos | C528(2.59); C287(1.80) | LDD2186 | [16] |
| LDCM0571 | Fragment9 | Ramos | C528(1.28) | LDD2188 | [16] |
| LDCM0022 | KB02 | HEK-293T | C287(1.00); C400(1.17); C39(0.92) | LDD1492 | [15] |
| LDCM0023 | KB03 | Jurkat | C528(4.81); C7(53.72) | LDD0209 | [17] |
| LDCM0024 | KB05 | COLO792 | C131(2.57) | LDD3310 | [18] |
| LDCM0030 | Luteolin | HEK-293T | 4.63 | LDD0182 | [2] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C333(2.43) | LDD2100 | [5] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C333(0.76) | LDD2101 | [5] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C333(0.60) | LDD2125 | [5] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C333(0.44) | LDD2133 | [5] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C333(0.54) | LDD2134 | [5] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C333(0.81) | LDD2135 | [5] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C333(1.09) | LDD2137 | [5] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C333(0.80) | LDD2146 | [5] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C23(2.40) | LDD2206 | [19] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
Other
References
