Details of the Target
General Information of Target
| Target ID | LDTP06868 | |||||
|---|---|---|---|---|---|---|
| Target Name | Angiomotin (AMOT) | |||||
| Gene Name | AMOT | |||||
| Gene ID | 154796 | |||||
| Synonyms |
KIAA1071; Angiomotin |
|||||
| 3D Structure | ||||||
| Sequence |
MRNSEEQPSGGTTVLQRLLQEQLRYGNPSENRSLLAIHQQATGNGPPFPSGSGNPGPQSD
VLSPQDHHQQLVAHAARQEPQGQEIQSENLIMEKQLSPRMQNNEELPTYEEAKVQSQYFR GQQHASVGAAFYVTGVTNQKMRTEGRPSVQRLNPGKMHQDEGLRDLKQGHVRSLSERLMQ MSLATSGVKAHPPVTSAPLSPPQPNDLYKNPTSSSEFYKAQGPLPNQHSLKGMEHRGPPP EYPFKGMPPQSVVCKPQEPGHFYSEHRLNQPGRTEGQLMRYQHPPEYGAARPAQDISLPL SARNSQPHSPTSSLTSGGSLPLLQSPPSTRLSPARHPLVPNQGDHSAHLPRPQQHFLPNQ AHQGDHYRLSQPGLSQQQQQQQQQHHHHHHHQQQQQQQPQQQPGEAYSAMPRAQPSSASY QPVPADPFAIVSRAQQMVEILSDENRNLRQELEGCYEKVARLQKVETEIQRVSEAYENLV KSSSKREALEKAMRNKLEGEIRRMHDFNRDLRERLETANKQLAEKEYEGSEDTRKTISQL FAKNKESQREKEKLEAELATARSTNEDQRRHIEIRDQALSNAQAKVVKLEEELKKKQVYV DKVEKMQQALVQLQAACEKREQLEHRLRTRLERELESLRIQQRQGNCQPTNVSEYNAAAL MELLREKEERILALEADMTKWEQKYLEENVMRHFALDAAATVAAQRDTTVISHSPNTSYD TALEARIQKEEEEILMANKRCLDMEGRIKTLHAQIIEKDAMIKVLQQRSRKEPSKTEQLS CMRPAKSLMSISNAGSGLLSHSSTLTGSPIMEEKRDDKSWKGSLGILLGGDYRAEYVPST PSPVPPSTPLLSAHSKTGSRDCSTQTERGTESNKTAAVAPISVPAPVAAAATAAAITATA ATITTTMVAAAPVAVAAAAAPAAAAAPSPATAAATAAAVSPAAAGQIPAAASVASAAAVA PSAAAAAAVQVAPAAPAPVPAPALVPVPAPAAAQASAPAQTQAPTSAPAVAPTPAPTPTP AVAQAEVPASPATGPGPHRLSIPSLTCNPDKTDGPVFHSNTLERKTPIQILGQEPDAEMV EYLI |
|||||
| Target Type |
Literature-reported
|
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Angiomotin family
|
|||||
| Subcellular location |
Cell junction, tight junction
|
|||||
| Function |
Plays a central role in tight junction maintenance via the complex formed with ARHGAP17, which acts by regulating the uptake of polarity proteins at tight junctions. Appears to regulate endothelial cell migration and tube formation. May also play a role in the assembly of endothelial cell-cell junctions.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
TH211 Probe Info |
 |
Y281(20.00); Y832(14.99); Y719(14.64); Y599(12.30) | LDD0257 | [1] | |
|
DA-P3 Probe Info |
 |
11.83 | LDD0185 | [2] | |
|
AHL-Pu-1 Probe Info |
 |
C647(2.23) | LDD0168 | [3] | |
|
HHS-465 Probe Info |
 |
Y25(10.00); Y281(8.40); Y420(5.01); Y599(10.00) | LDD2237 | [4] | |
|
DBIA Probe Info |
 |
C455(1.70); C741(1.40); C1047(1.75) | LDD0080 | [5] | |
|
ATP probe Probe Info |
 |
K458(0.00); K156(0.00); K1051(0.00); K543(0.00) | LDD0199 | [6] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [7] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0356 | [8] | |
|
ENE Probe Info |
 |
C862(0.00); C1047(0.00); C781(0.00); C455(0.00) | LDD0006 | [9] | |
|
IPM Probe Info |
 |
C741(0.00); C1047(0.00); C617(0.00); C254(0.00) | LDD0005 | [9] | |
|
NHS Probe Info |
 |
K585(0.00); K543(0.00) | LDD0010 | [9] | |
|
NPM Probe Info |
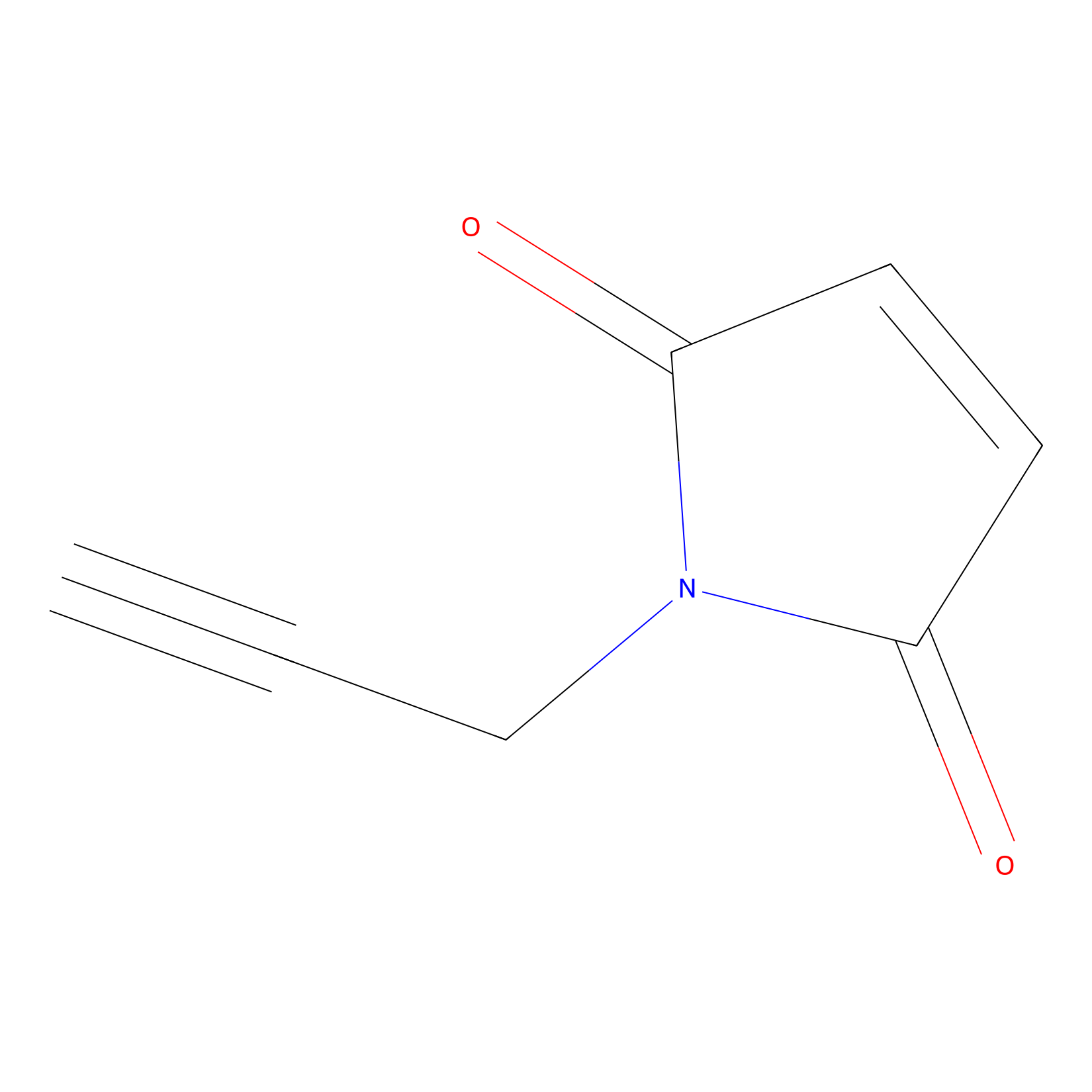 |
N.A. | LDD0016 | [9] | |
|
PPMS Probe Info |
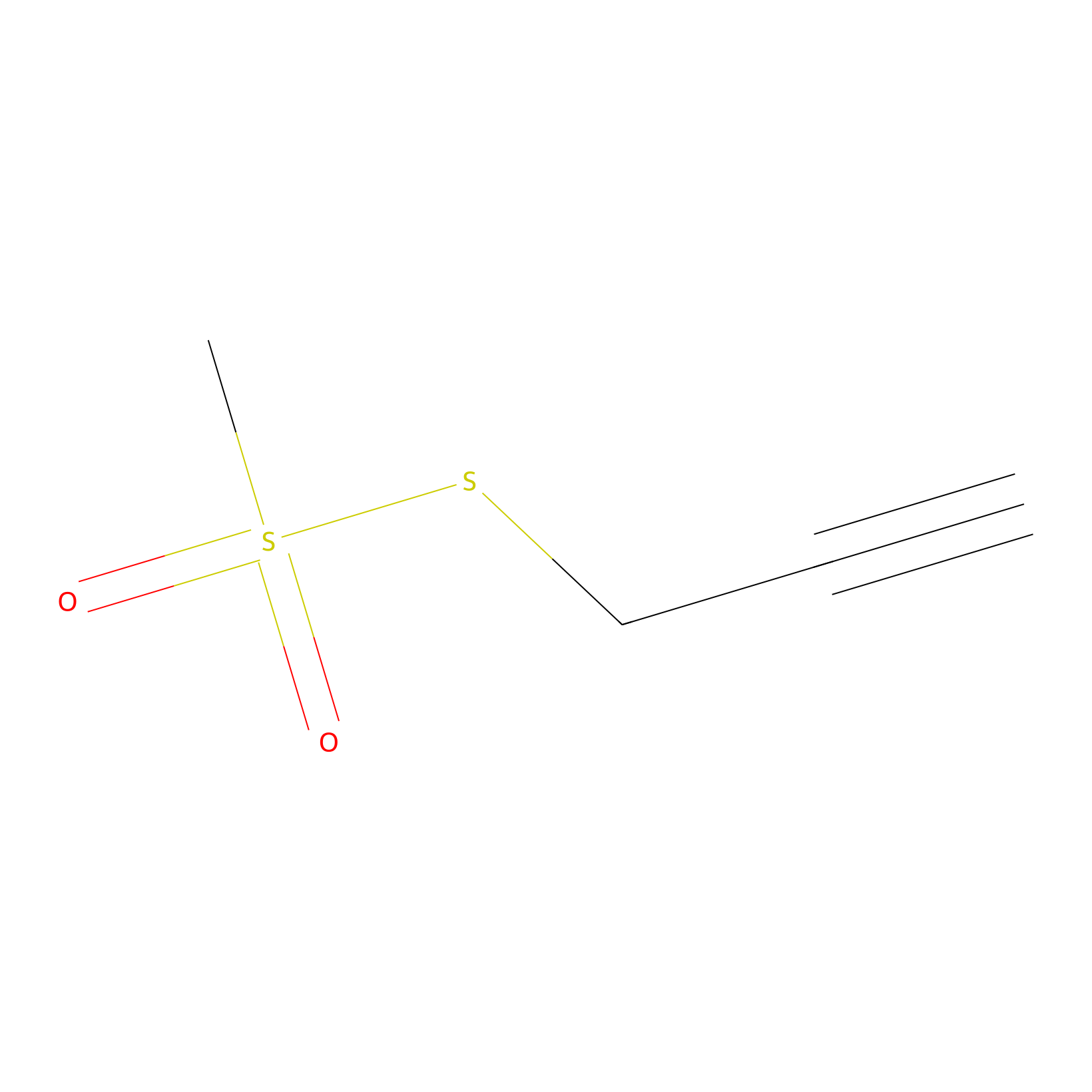 |
N.A. | LDD0008 | [9] | |
|
SF Probe Info |
 |
K596(0.00); Y719(0.00); Y599(0.00); Y287(0.00) | LDD0028 | [10] | |
|
STPyne Probe Info |
 |
K585(0.00); K520(0.00); K464(0.00); K481(0.00) | LDD0009 | [9] | |
|
VSF Probe Info |
 |
C455(0.00); C1047(0.00); C781(0.00); C862(0.00) | LDD0007 | [9] | |
|
Phosphinate-6 Probe Info |
 |
C617(0.00); C781(0.00); C647(0.00); C254(0.00) | LDD0018 | [11] | |
|
1c-yne Probe Info |
 |
K464(0.00); K856(0.00); K1051(0.00); K684(0.00) | LDD0228 | [8] | |
|
AOyne Probe Info |
 |
11.50 | LDD0443 | [12] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C112 Probe Info |
 |
18.00 | LDD1799 | [13] | |
|
C163 Probe Info |
 |
5.82 | LDD1843 | [13] | |
|
C177 Probe Info |
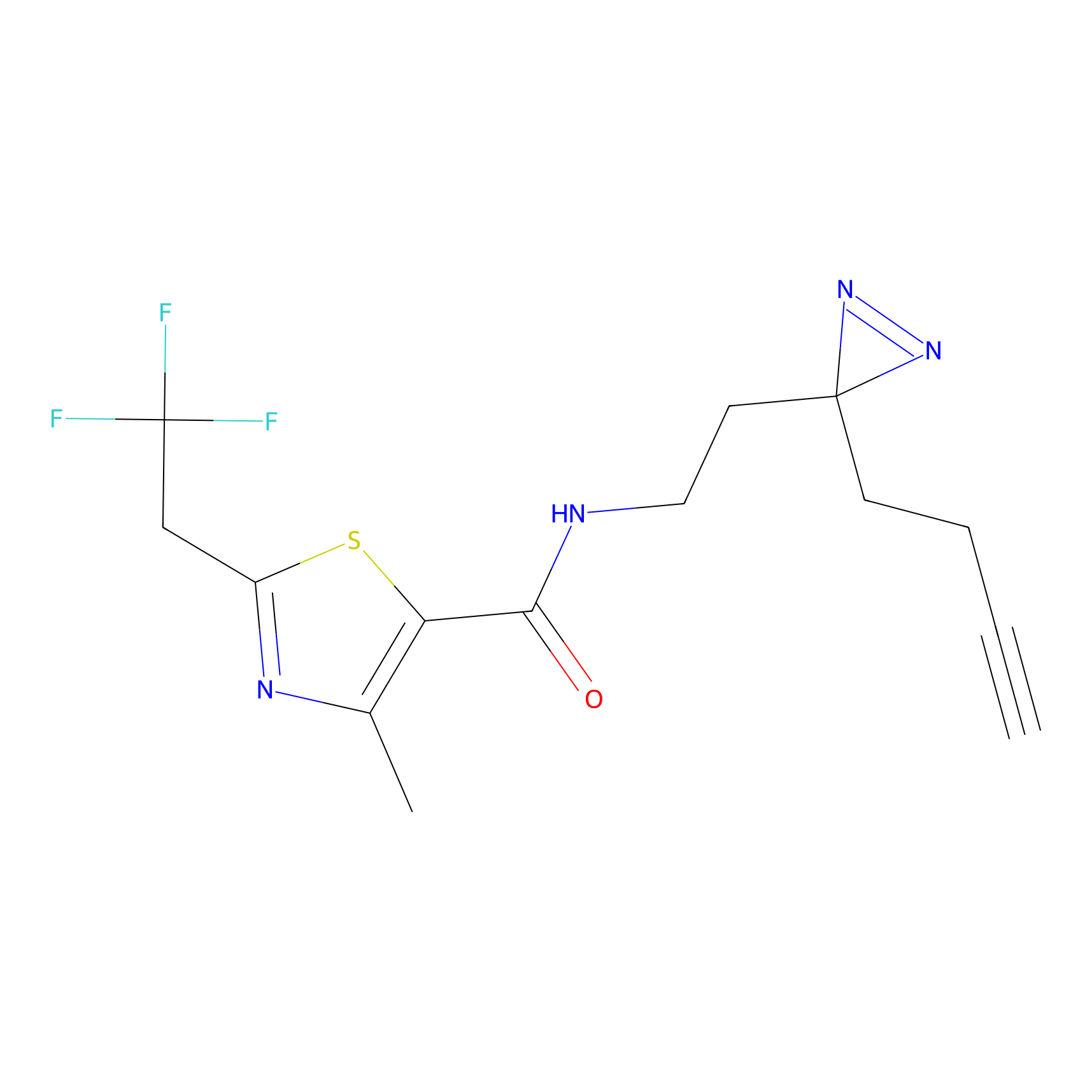 |
9.06 | LDD1856 | [13] | |
|
C245 Probe Info |
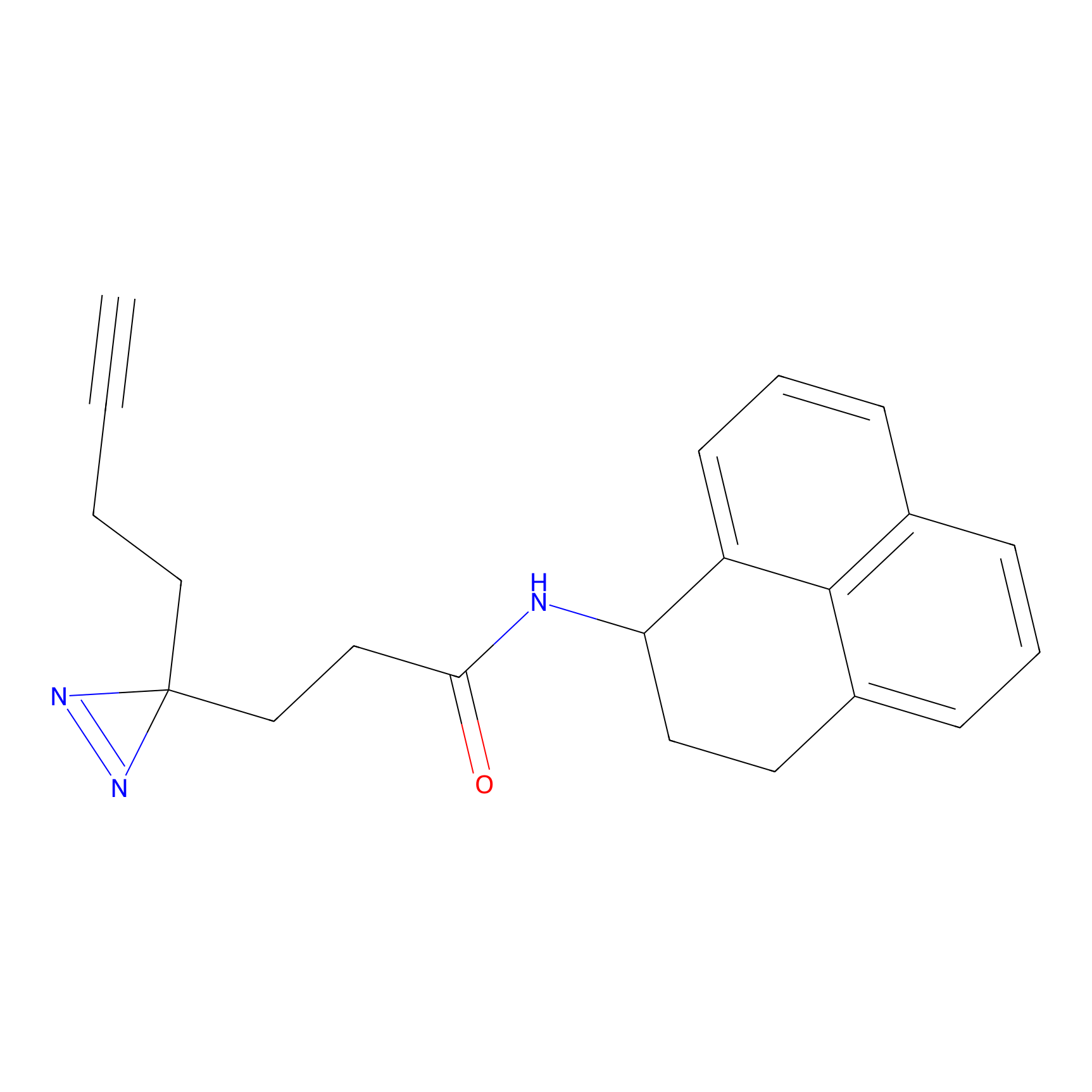 |
10.20 | LDD1918 | [13] | |
|
C246 Probe Info |
 |
13.45 | LDD1919 | [13] | |
|
C266 Probe Info |
 |
8.22 | LDD1937 | [13] | |
|
C285 Probe Info |
 |
24.08 | LDD1955 | [13] | |
|
C287 Probe Info |
 |
8.57 | LDD1957 | [13] | |
|
C338 Probe Info |
 |
11.55 | LDD2001 | [13] | |
|
C355 Probe Info |
 |
31.34 | LDD2016 | [13] | |
|
C391 Probe Info |
 |
13.18 | LDD2050 | [13] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | HEK-293T | C647(2.23) | LDD0168 | [3] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C647(2.40) | LDD0372 | [3] |
| LDCM0214 | AC1 | HEK-293T | C862(1.00); C741(0.96); C781(1.24); C455(1.07) | LDD1507 | [14] |
| LDCM0215 | AC10 | HEK-293T | C862(0.94); C741(1.04); C781(0.99); C455(1.02) | LDD1508 | [14] |
| LDCM0226 | AC11 | HEK-293T | C862(1.05); C741(0.98); C781(0.96); C455(0.89) | LDD1509 | [14] |
| LDCM0237 | AC12 | HEK-293T | C862(1.04); C741(0.99); C781(1.01); C455(1.02) | LDD1510 | [14] |
| LDCM0259 | AC14 | HEK-293T | C862(1.02); C741(1.04); C781(0.87); C455(1.08) | LDD1512 | [14] |
| LDCM0263 | AC143 | HEK-293T | C862(0.82) | LDD0861 | [5] |
| LDCM0264 | AC144 | HEK-293T | C862(1.09) | LDD0862 | [5] |
| LDCM0265 | AC145 | HEK-293T | C862(1.33) | LDD0863 | [5] |
| LDCM0266 | AC146 | HEK-293T | C862(1.16) | LDD0864 | [5] |
| LDCM0267 | AC147 | HEK-293T | C862(1.02) | LDD0865 | [5] |
| LDCM0268 | AC148 | HEK-293T | C862(1.08) | LDD0866 | [5] |
| LDCM0269 | AC149 | HEK-293T | C862(0.66) | LDD0867 | [5] |
| LDCM0270 | AC15 | HEK-293T | C862(1.01); C741(0.98); C781(0.98); C455(1.04) | LDD1513 | [14] |
| LDCM0271 | AC150 | HEK-293T | C862(0.99) | LDD0869 | [5] |
| LDCM0272 | AC151 | HEK-293T | C862(0.91) | LDD0870 | [5] |
| LDCM0273 | AC152 | HEK-293T | C862(0.80) | LDD0871 | [5] |
| LDCM0274 | AC153 | HEK-293T | C862(0.88) | LDD0872 | [5] |
| LDCM0621 | AC154 | HEK-293T | C862(0.88) | LDD2162 | [5] |
| LDCM0622 | AC155 | HEK-293T | C862(1.21) | LDD2163 | [5] |
| LDCM0623 | AC156 | HEK-293T | C862(0.83) | LDD2164 | [5] |
| LDCM0624 | AC157 | HEK-293T | C862(0.78) | LDD2165 | [5] |
| LDCM0276 | AC17 | HEK-293T | C862(2.22) | LDD0874 | [5] |
| LDCM0277 | AC18 | HEK-293T | C862(0.76) | LDD0875 | [5] |
| LDCM0278 | AC19 | HEK-293T | C862(0.74) | LDD0876 | [5] |
| LDCM0279 | AC2 | HEK-293T | C862(0.96); C741(0.97); C781(1.00); C455(1.03) | LDD1518 | [14] |
| LDCM0280 | AC20 | HEK-293T | C862(0.92) | LDD0878 | [5] |
| LDCM0281 | AC21 | HEK-293T | C862(0.75) | LDD0879 | [5] |
| LDCM0282 | AC22 | HEK-293T | C862(0.86) | LDD0880 | [5] |
| LDCM0283 | AC23 | HEK-293T | C862(0.79) | LDD0881 | [5] |
| LDCM0284 | AC24 | HEK-293T | C862(0.85) | LDD0882 | [5] |
| LDCM0285 | AC25 | HEK-293T | C862(0.59) | LDD0883 | [5] |
| LDCM0286 | AC26 | HEK-293T | C862(0.87) | LDD0884 | [5] |
| LDCM0287 | AC27 | HEK-293T | C862(0.96) | LDD0885 | [5] |
| LDCM0288 | AC28 | HEK-293T | C862(1.12) | LDD0886 | [5] |
| LDCM0289 | AC29 | HEK-293T | C862(0.77) | LDD0887 | [5] |
| LDCM0290 | AC3 | HEK-293T | C862(0.99); C741(0.93); C781(1.05); C455(1.00) | LDD1529 | [14] |
| LDCM0291 | AC30 | HEK-293T | C862(0.72) | LDD0889 | [5] |
| LDCM0292 | AC31 | HEK-293T | C862(1.01) | LDD0890 | [5] |
| LDCM0293 | AC32 | HEK-293T | C862(0.82) | LDD0891 | [5] |
| LDCM0294 | AC33 | HEK-293T | C862(0.72) | LDD0892 | [5] |
| LDCM0295 | AC34 | HEK-293T | C862(0.87) | LDD0893 | [5] |
| LDCM0296 | AC35 | HEK-293T | C862(1.03); C741(0.96); C781(0.88); C455(0.92) | LDD1535 | [14] |
| LDCM0297 | AC36 | HEK-293T | C862(0.91); C741(0.94); C781(1.01); C455(1.00) | LDD1536 | [14] |
| LDCM0298 | AC37 | HEK-293T | C862(0.98); C741(1.03); C781(0.94); C455(1.06) | LDD1537 | [14] |
| LDCM0299 | AC38 | HEK-293T | C862(0.93); C741(0.99); C781(0.91); C455(1.03) | LDD1538 | [14] |
| LDCM0300 | AC39 | HEK-293T | C862(1.01); C741(1.02); C781(0.99); C455(1.09) | LDD1539 | [14] |
| LDCM0301 | AC4 | HEK-293T | C862(0.98); C741(0.92); C781(1.09); C455(1.04) | LDD1540 | [14] |
| LDCM0302 | AC40 | HEK-293T | C862(0.99); C741(0.99); C781(0.86); C455(1.00) | LDD1541 | [14] |
| LDCM0303 | AC41 | HEK-293T | C862(1.08); C741(1.03); C781(1.07); C455(1.08) | LDD1542 | [14] |
| LDCM0304 | AC42 | HEK-293T | C862(1.03); C741(1.00); C781(1.04); C455(1.01) | LDD1543 | [14] |
| LDCM0305 | AC43 | HEK-293T | C862(0.98); C741(0.94); C781(0.91); C455(1.00) | LDD1544 | [14] |
| LDCM0306 | AC44 | HEK-293T | C862(1.02); C741(0.94); C781(1.01); C455(1.11) | LDD1545 | [14] |
| LDCM0307 | AC45 | HEK-293T | C862(1.04); C741(1.02); C781(0.99); C455(1.10) | LDD1546 | [14] |
| LDCM0308 | AC46 | HEK-293T | C862(1.01); C741(0.96); C781(1.00); C455(1.01) | LDD1547 | [14] |
| LDCM0309 | AC47 | HEK-293T | C862(0.99); C741(0.94); C781(1.00); C455(1.07) | LDD1548 | [14] |
| LDCM0310 | AC48 | HEK-293T | C862(1.01); C741(1.02); C781(0.97); C455(1.03) | LDD1549 | [14] |
| LDCM0311 | AC49 | HEK-293T | C862(1.02); C741(1.01); C781(1.14); C455(1.07) | LDD1550 | [14] |
| LDCM0312 | AC5 | HEK-293T | C862(0.95); C741(1.03); C781(1.07); C455(0.98) | LDD1551 | [14] |
| LDCM0313 | AC50 | HEK-293T | C862(0.96); C741(0.98); C781(1.03); C455(0.98) | LDD1552 | [14] |
| LDCM0314 | AC51 | HEK-293T | C862(1.01); C741(0.97); C781(1.03); C455(1.01) | LDD1553 | [14] |
| LDCM0315 | AC52 | HEK-293T | C862(1.02); C741(0.95); C781(1.02); C455(1.07) | LDD1554 | [14] |
| LDCM0316 | AC53 | HEK-293T | C862(0.95); C741(1.04); C781(1.01); C455(1.09) | LDD1555 | [14] |
| LDCM0317 | AC54 | HEK-293T | C862(0.98); C741(1.06); C781(0.98); C455(1.03) | LDD1556 | [14] |
| LDCM0318 | AC55 | HEK-293T | C862(1.00); C741(1.02); C781(0.98); C455(1.07) | LDD1557 | [14] |
| LDCM0319 | AC56 | HEK-293T | C862(1.03); C741(0.97); C781(0.98); C455(1.00) | LDD1558 | [14] |
| LDCM0320 | AC57 | HEK-293T | C862(1.26) | LDD0918 | [5] |
| LDCM0321 | AC58 | HEK-293T | C862(1.53) | LDD0919 | [5] |
| LDCM0322 | AC59 | HEK-293T | C862(0.99) | LDD0920 | [5] |
| LDCM0323 | AC6 | HEK-293T | C862(1.02); C741(0.95); C781(1.07); C455(1.01) | LDD1562 | [14] |
| LDCM0324 | AC60 | HEK-293T | C862(0.80) | LDD0922 | [5] |
| LDCM0325 | AC61 | HEK-293T | C862(0.98) | LDD0923 | [5] |
| LDCM0326 | AC62 | HEK-293T | C862(0.70) | LDD0924 | [5] |
| LDCM0327 | AC63 | HEK-293T | C862(1.15) | LDD0925 | [5] |
| LDCM0328 | AC64 | HEK-293T | C862(0.82) | LDD0926 | [5] |
| LDCM0329 | AC65 | HEK-293T | C862(1.02) | LDD0927 | [5] |
| LDCM0330 | AC66 | HEK-293T | C862(0.75) | LDD0928 | [5] |
| LDCM0331 | AC67 | HEK-293T | C862(0.99) | LDD0929 | [5] |
| LDCM0332 | AC68 | HEK-293T | C862(0.69) | LDD0930 | [5] |
| LDCM0333 | AC69 | HEK-293T | C862(0.82) | LDD0931 | [5] |
| LDCM0334 | AC7 | HEK-293T | C862(1.01); C741(0.99); C781(1.02); C455(1.12) | LDD1568 | [14] |
| LDCM0335 | AC70 | HEK-293T | C862(0.56) | LDD0933 | [5] |
| LDCM0336 | AC71 | HEK-293T | C862(0.77) | LDD0934 | [5] |
| LDCM0337 | AC72 | HEK-293T | C862(0.88) | LDD0935 | [5] |
| LDCM0338 | AC73 | HEK-293T | C862(0.65) | LDD0936 | [5] |
| LDCM0339 | AC74 | HEK-293T | C862(0.71) | LDD0937 | [5] |
| LDCM0340 | AC75 | HEK-293T | C862(0.69) | LDD0938 | [5] |
| LDCM0341 | AC76 | HEK-293T | C862(0.83) | LDD0939 | [5] |
| LDCM0342 | AC77 | HEK-293T | C862(0.80) | LDD0940 | [5] |
| LDCM0343 | AC78 | HEK-293T | C862(1.10) | LDD0941 | [5] |
| LDCM0344 | AC79 | HEK-293T | C862(0.60) | LDD0942 | [5] |
| LDCM0345 | AC8 | HEK-293T | C862(0.96); C741(0.97); C781(0.96); C455(1.09) | LDD1569 | [14] |
| LDCM0346 | AC80 | HEK-293T | C862(0.75) | LDD0944 | [5] |
| LDCM0347 | AC81 | HEK-293T | C862(0.72) | LDD0945 | [5] |
| LDCM0348 | AC82 | HEK-293T | C862(0.80) | LDD0946 | [5] |
| LDCM0349 | AC83 | HEK-293T | C862(1.06) | LDD0947 | [5] |
| LDCM0350 | AC84 | HEK-293T | C862(1.38) | LDD0948 | [5] |
| LDCM0351 | AC85 | HEK-293T | C862(1.27) | LDD0949 | [5] |
| LDCM0352 | AC86 | HEK-293T | C862(1.21) | LDD0950 | [5] |
| LDCM0353 | AC87 | HEK-293T | C862(1.00) | LDD0951 | [5] |
| LDCM0354 | AC88 | HEK-293T | C862(1.28) | LDD0952 | [5] |
| LDCM0355 | AC89 | HEK-293T | C862(1.35) | LDD0953 | [5] |
| LDCM0357 | AC90 | HEK-293T | C862(1.65) | LDD0955 | [5] |
| LDCM0358 | AC91 | HEK-293T | C862(1.19) | LDD0956 | [5] |
| LDCM0359 | AC92 | HEK-293T | C862(1.29) | LDD0957 | [5] |
| LDCM0360 | AC93 | HEK-293T | C862(1.05) | LDD0958 | [5] |
| LDCM0361 | AC94 | HEK-293T | C862(1.04) | LDD0959 | [5] |
| LDCM0362 | AC95 | HEK-293T | C862(1.04) | LDD0960 | [5] |
| LDCM0363 | AC96 | HEK-293T | C862(1.26) | LDD0961 | [5] |
| LDCM0364 | AC97 | HEK-293T | C862(1.16) | LDD0962 | [5] |
| LDCM0248 | AKOS034007472 | HEK-293T | C862(0.99); C741(1.09); C781(0.99); C455(1.01) | LDD1511 | [14] |
| LDCM0356 | AKOS034007680 | HEK-293T | C862(1.05); C741(1.00); C781(1.08); C455(1.06) | LDD1570 | [14] |
| LDCM0275 | AKOS034007705 | HEK-293T | C862(1.01); C741(0.96); C781(0.98); C455(0.96) | LDD1514 | [14] |
| LDCM0087 | Capsaicin | HEK-293T | 11.83 | LDD0185 | [2] |
| LDCM0367 | CL1 | HEK-293T | C862(0.91); C741(0.95); C781(1.02); C455(1.00) | LDD1571 | [14] |
| LDCM0368 | CL10 | HEK-293T | C862(0.77); C741(0.86); C781(0.67); C455(1.10) | LDD1572 | [14] |
| LDCM0369 | CL100 | HEK-293T | C862(0.94); C741(0.93); C781(1.10); C455(1.06) | LDD1573 | [14] |
| LDCM0370 | CL101 | HEK-293T | C862(1.01); C741(0.96); C781(1.10); C455(1.08) | LDD1574 | [14] |
| LDCM0371 | CL102 | HEK-293T | C862(1.02); C741(0.96); C781(1.02); C455(1.01) | LDD1575 | [14] |
| LDCM0372 | CL103 | HEK-293T | C862(0.97); C741(1.04); C781(0.94); C455(1.04) | LDD1576 | [14] |
| LDCM0373 | CL104 | HEK-293T | C862(0.97); C741(0.97); C781(1.05); C455(0.92) | LDD1577 | [14] |
| LDCM0374 | CL105 | HEK-293T | C862(1.19) | LDD0972 | [5] |
| LDCM0375 | CL106 | HEK-293T | C862(1.14) | LDD0973 | [5] |
| LDCM0376 | CL107 | HEK-293T | C862(1.15) | LDD0974 | [5] |
| LDCM0377 | CL108 | HEK-293T | C862(0.77) | LDD0975 | [5] |
| LDCM0378 | CL109 | HEK-293T | C862(0.90) | LDD0976 | [5] |
| LDCM0379 | CL11 | HEK-293T | C862(0.97); C741(0.92); C781(0.82); C455(1.23) | LDD1583 | [14] |
| LDCM0380 | CL110 | HEK-293T | C862(0.62) | LDD0978 | [5] |
| LDCM0381 | CL111 | HEK-293T | C862(0.95) | LDD0979 | [5] |
| LDCM0382 | CL112 | HEK-293T | C862(0.60) | LDD0980 | [5] |
| LDCM0383 | CL113 | HEK-293T | C862(0.67) | LDD0981 | [5] |
| LDCM0384 | CL114 | HEK-293T | C862(0.65) | LDD0982 | [5] |
| LDCM0385 | CL115 | HEK-293T | C862(0.96) | LDD0983 | [5] |
| LDCM0386 | CL116 | HEK-293T | C862(0.85) | LDD0984 | [5] |
| LDCM0387 | CL117 | HEK-293T | C862(0.99); C741(0.99); C781(1.16); C455(1.16) | LDD1591 | [14] |
| LDCM0388 | CL118 | HEK-293T | C862(1.03); C741(1.06); C781(1.12); C455(1.03) | LDD1592 | [14] |
| LDCM0389 | CL119 | HEK-293T | C862(1.02); C741(1.02); C781(0.99); C455(1.10) | LDD1593 | [14] |
| LDCM0390 | CL12 | HEK-293T | C862(0.98); C741(0.94); C781(0.67); C455(1.00) | LDD1594 | [14] |
| LDCM0391 | CL120 | HEK-293T | C862(1.01); C741(0.99); C781(1.05); C455(0.98) | LDD1595 | [14] |
| LDCM0392 | CL121 | HEK-293T | C862(0.97); C741(1.02); C781(1.10); C455(1.10) | LDD1596 | [14] |
| LDCM0393 | CL122 | HEK-293T | C862(0.95); C741(0.98); C781(1.07); C455(1.02) | LDD1597 | [14] |
| LDCM0394 | CL123 | HEK-293T | C862(0.85); C741(0.93); C781(1.04); C455(0.97) | LDD1598 | [14] |
| LDCM0395 | CL124 | HEK-293T | C862(0.96); C741(0.94); C781(1.18); C455(1.03) | LDD1599 | [14] |
| LDCM0396 | CL125 | HEK-293T | C862(1.25) | LDD0994 | [5] |
| LDCM0397 | CL126 | HEK-293T | C862(0.91) | LDD0995 | [5] |
| LDCM0398 | CL127 | HEK-293T | C862(1.20) | LDD0996 | [5] |
| LDCM0399 | CL128 | HEK-293T | C862(0.70) | LDD0997 | [5] |
| LDCM0400 | CL13 | HEK-293T | C862(0.97); C741(1.06); C781(1.23); C455(1.06) | LDD1604 | [14] |
| LDCM0401 | CL14 | HEK-293T | C862(1.01); C741(0.94); C781(1.09); C455(0.96) | LDD1605 | [14] |
| LDCM0402 | CL15 | HEK-293T | C862(0.89); C741(1.23); C781(0.92); C455(1.01) | LDD1606 | [14] |
| LDCM0403 | CL16 | HEK-293T | C862(0.96); C741(1.02); C781(0.88); C455(1.10) | LDD1607 | [14] |
| LDCM0404 | CL17 | HEK-293T | C862(0.94); C741(1.05); C781(0.95); C455(1.03) | LDD1608 | [14] |
| LDCM0405 | CL18 | HEK-293T | C862(0.95); C741(1.04); C781(0.81); C455(1.03) | LDD1609 | [14] |
| LDCM0406 | CL19 | HEK-293T | C862(1.02); C741(1.08); C781(0.62); C455(0.97) | LDD1610 | [14] |
| LDCM0407 | CL2 | HEK-293T | C862(0.99); C741(0.87); C781(1.10); C455(0.99) | LDD1611 | [14] |
| LDCM0408 | CL20 | HEK-293T | C862(1.07); C741(1.08); C781(0.91); C455(1.19) | LDD1612 | [14] |
| LDCM0409 | CL21 | HEK-293T | C862(0.94); C741(1.09); C781(0.52); C455(1.05) | LDD1613 | [14] |
| LDCM0410 | CL22 | HEK-293T | C862(1.00); C741(0.95); C781(0.62); C455(1.10) | LDD1614 | [14] |
| LDCM0411 | CL23 | HEK-293T | C862(1.05); C741(0.97); C781(0.80); C455(1.23) | LDD1615 | [14] |
| LDCM0412 | CL24 | HEK-293T | C862(0.98); C741(1.02); C781(0.68); C455(1.07) | LDD1616 | [14] |
| LDCM0413 | CL25 | HEK-293T | C862(0.88); C741(0.87); C781(1.15); C455(1.08) | LDD1617 | [14] |
| LDCM0414 | CL26 | HEK-293T | C862(0.98); C741(0.94); C781(0.92); C455(0.96) | LDD1618 | [14] |
| LDCM0415 | CL27 | HEK-293T | C862(0.95); C741(1.07); C781(0.93); C455(1.08) | LDD1619 | [14] |
| LDCM0416 | CL28 | HEK-293T | C862(1.01); C741(0.98); C781(0.97); C455(1.05) | LDD1620 | [14] |
| LDCM0417 | CL29 | HEK-293T | C862(1.10); C741(1.03); C781(0.89); C455(1.13) | LDD1621 | [14] |
| LDCM0418 | CL3 | HEK-293T | C862(0.93); C741(1.07); C781(0.90); C455(1.07) | LDD1622 | [14] |
| LDCM0419 | CL30 | HEK-293T | C862(1.00); C741(1.03); C781(0.77); C455(1.03) | LDD1623 | [14] |
| LDCM0420 | CL31 | HEK-293T | C862(1.04); C741(1.06); C781(0.71); C455(0.96) | LDD1624 | [14] |
| LDCM0421 | CL32 | HEK-293T | C862(1.06); C741(1.04); C781(0.95); C455(1.13) | LDD1625 | [14] |
| LDCM0422 | CL33 | HEK-293T | C862(0.88); C741(1.08); C781(0.53); C455(0.93) | LDD1626 | [14] |
| LDCM0423 | CL34 | HEK-293T | C862(1.00); C741(1.03); C781(0.63); C455(1.12) | LDD1627 | [14] |
| LDCM0424 | CL35 | HEK-293T | C862(1.10); C741(1.05); C781(0.77); C455(1.17) | LDD1628 | [14] |
| LDCM0425 | CL36 | HEK-293T | C862(1.03); C741(0.98); C781(0.69); C455(1.05) | LDD1629 | [14] |
| LDCM0426 | CL37 | HEK-293T | C862(0.90); C741(0.91); C781(1.10); C455(0.99) | LDD1630 | [14] |
| LDCM0428 | CL39 | HEK-293T | C862(0.97); C741(0.91); C781(0.94); C455(1.09) | LDD1632 | [14] |
| LDCM0429 | CL4 | HEK-293T | C862(0.95); C741(1.01); C781(0.94); C455(1.11) | LDD1633 | [14] |
| LDCM0430 | CL40 | HEK-293T | C862(1.08); C741(1.02); C781(0.91); C455(1.05) | LDD1634 | [14] |
| LDCM0431 | CL41 | HEK-293T | C862(1.04); C741(1.03); C781(0.92); C455(1.05) | LDD1635 | [14] |
| LDCM0432 | CL42 | HEK-293T | C862(1.06); C741(1.00); C781(0.83); C455(1.08) | LDD1636 | [14] |
| LDCM0433 | CL43 | HEK-293T | C862(1.05); C741(1.01); C781(0.64); C455(1.10) | LDD1637 | [14] |
| LDCM0434 | CL44 | HEK-293T | C862(1.13); C741(1.03); C781(0.87); C455(1.14) | LDD1638 | [14] |
| LDCM0435 | CL45 | HEK-293T | C862(1.02); C741(1.01); C781(0.59); C455(1.11) | LDD1639 | [14] |
| LDCM0436 | CL46 | HEK-293T | C862(1.07); C741(1.03); C781(0.61); C455(1.07) | LDD1640 | [14] |
| LDCM0437 | CL47 | HEK-293T | C862(1.08); C741(0.99); C781(0.79); C455(1.22) | LDD1641 | [14] |
| LDCM0438 | CL48 | HEK-293T | C862(1.07); C741(0.96); C781(0.75); C455(0.95) | LDD1642 | [14] |
| LDCM0439 | CL49 | HEK-293T | C862(0.93); C741(0.94); C781(1.17); C455(0.97) | LDD1643 | [14] |
| LDCM0440 | CL5 | HEK-293T | C862(1.00); C741(1.00); C781(1.04); C455(1.00) | LDD1644 | [14] |
| LDCM0441 | CL50 | HEK-293T | C862(0.98); C741(0.92); C781(0.96); C455(1.03) | LDD1645 | [14] |
| LDCM0443 | CL52 | HEK-293T | C862(1.01); C741(0.96); C781(0.91); C455(1.02) | LDD1646 | [14] |
| LDCM0444 | CL53 | HEK-293T | C862(0.95); C741(1.03); C781(0.98); C455(1.07) | LDD1647 | [14] |
| LDCM0445 | CL54 | HEK-293T | C862(0.92); C741(1.00); C781(0.82); C455(0.98) | LDD1648 | [14] |
| LDCM0446 | CL55 | HEK-293T | C862(1.08); C741(1.02); C781(0.61); C455(1.05) | LDD1649 | [14] |
| LDCM0447 | CL56 | HEK-293T | C862(1.06); C741(1.01); C781(0.89); C455(1.10) | LDD1650 | [14] |
| LDCM0448 | CL57 | HEK-293T | C862(1.02); C741(1.17); C781(0.64); C455(1.25) | LDD1651 | [14] |
| LDCM0449 | CL58 | HEK-293T | C862(1.03); C741(1.02); C781(0.63); C455(1.07) | LDD1652 | [14] |
| LDCM0450 | CL59 | HEK-293T | C862(1.05); C741(0.95); C781(0.82); C455(1.16) | LDD1653 | [14] |
| LDCM0451 | CL6 | HEK-293T | C862(0.89); C741(0.91); C781(0.89); C455(0.74) | LDD1654 | [14] |
| LDCM0452 | CL60 | HEK-293T | C862(1.00); C741(0.97); C781(0.70); C455(0.98) | LDD1655 | [14] |
| LDCM0453 | CL61 | HEK-293T | C862(0.88) | LDD1051 | [5] |
| LDCM0454 | CL62 | HEK-293T | C862(1.10) | LDD1052 | [5] |
| LDCM0455 | CL63 | HEK-293T | C862(0.67) | LDD1053 | [5] |
| LDCM0456 | CL64 | HEK-293T | C862(1.43) | LDD1054 | [5] |
| LDCM0457 | CL65 | HEK-293T | C862(0.93) | LDD1055 | [5] |
| LDCM0458 | CL66 | HEK-293T | C862(1.04) | LDD1056 | [5] |
| LDCM0459 | CL67 | HEK-293T | C862(0.61) | LDD1057 | [5] |
| LDCM0460 | CL68 | HEK-293T | C862(0.60) | LDD1058 | [5] |
| LDCM0461 | CL69 | HEK-293T | C862(0.87) | LDD1059 | [5] |
| LDCM0462 | CL7 | HEK-293T | C862(0.98); C741(1.02); C781(0.66); C455(1.04) | LDD1665 | [14] |
| LDCM0463 | CL70 | HEK-293T | C862(1.22) | LDD1061 | [5] |
| LDCM0464 | CL71 | HEK-293T | C862(1.42) | LDD1062 | [5] |
| LDCM0465 | CL72 | HEK-293T | C862(0.98) | LDD1063 | [5] |
| LDCM0466 | CL73 | HEK-293T | C862(0.58) | LDD1064 | [5] |
| LDCM0467 | CL74 | HEK-293T | C862(0.87) | LDD1065 | [5] |
| LDCM0469 | CL76 | HEK-293T | C862(0.82) | LDD1067 | [5] |
| LDCM0470 | CL77 | HEK-293T | C862(0.67) | LDD1068 | [5] |
| LDCM0471 | CL78 | HEK-293T | C862(0.71) | LDD1069 | [5] |
| LDCM0472 | CL79 | HEK-293T | C862(0.89) | LDD1070 | [5] |
| LDCM0473 | CL8 | HEK-293T | C862(0.81); C741(1.03); C781(1.17); C455(1.09) | LDD1676 | [14] |
| LDCM0474 | CL80 | HEK-293T | C862(1.33) | LDD1072 | [5] |
| LDCM0475 | CL81 | HEK-293T | C862(0.68) | LDD1073 | [5] |
| LDCM0476 | CL82 | HEK-293T | C862(0.73) | LDD1074 | [5] |
| LDCM0477 | CL83 | HEK-293T | C862(0.53) | LDD1075 | [5] |
| LDCM0478 | CL84 | HEK-293T | C862(0.73) | LDD1076 | [5] |
| LDCM0479 | CL85 | HEK-293T | C862(0.94) | LDD1077 | [5] |
| LDCM0480 | CL86 | HEK-293T | C862(1.04) | LDD1078 | [5] |
| LDCM0481 | CL87 | HEK-293T | C862(0.70) | LDD1079 | [5] |
| LDCM0482 | CL88 | HEK-293T | C862(1.14) | LDD1080 | [5] |
| LDCM0483 | CL89 | HEK-293T | C862(1.06) | LDD1081 | [5] |
| LDCM0484 | CL9 | HEK-293T | C862(0.91); C741(1.09); C781(0.53); C455(0.99) | LDD1687 | [14] |
| LDCM0485 | CL90 | HEK-293T | C862(0.66) | LDD1083 | [5] |
| LDCM0486 | CL91 | HEK-293T | C862(1.02); C741(0.96); C781(0.86); C455(1.07) | LDD1689 | [14] |
| LDCM0487 | CL92 | HEK-293T | C862(0.98); C741(0.97); C781(0.99); C455(1.06) | LDD1690 | [14] |
| LDCM0488 | CL93 | HEK-293T | C862(0.94); C741(1.06); C781(0.69); C455(1.18) | LDD1691 | [14] |
| LDCM0489 | CL94 | HEK-293T | C862(1.00); C741(0.88); C781(0.77); C455(1.03) | LDD1692 | [14] |
| LDCM0490 | CL95 | HEK-293T | C862(0.95); C741(1.06); C781(0.91); C455(1.08) | LDD1693 | [14] |
| LDCM0491 | CL96 | HEK-293T | C862(0.98); C741(0.80); C781(0.95); C455(0.82) | LDD1694 | [14] |
| LDCM0492 | CL97 | HEK-293T | C862(0.95); C741(0.95); C781(1.18); C455(1.02) | LDD1695 | [14] |
| LDCM0493 | CL98 | HEK-293T | C862(0.99); C741(0.91); C781(1.05); C455(1.00) | LDD1696 | [14] |
| LDCM0494 | CL99 | HEK-293T | C862(0.95); C741(0.89); C781(1.09); C455(1.01) | LDD1697 | [14] |
| LDCM0495 | E2913 | HEK-293T | C862(0.98); C741(0.87); C781(0.98); C455(1.11) | LDD1698 | [14] |
| LDCM0468 | Fragment33 | HEK-293T | C862(0.60) | LDD1066 | [5] |
| LDCM0427 | Fragment51 | HEK-293T | C862(1.04); C741(0.96); C781(0.92); C455(1.03) | LDD1631 | [14] |
| LDCM0022 | KB02 | HCT 116 | C455(1.70); C741(1.40); C1047(1.75) | LDD0080 | [5] |
| LDCM0023 | KB03 | HCT 116 | C455(1.71); C741(1.27); C1047(0.99) | LDD0081 | [5] |
| LDCM0024 | KB05 | HCT 116 | C455(2.40); C741(1.77); C1047(1.59) | LDD0082 | [5] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Other
References
