Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
TH211 Probe Info |
 |
Y627(17.41) | LDD0260 | [2] | |
|
STPyne Probe Info |
 |
K546(6.67); K566(6.49) | LDD0277 | [3] | |
|
Probe 1 Probe Info |
 |
Y1004(13.13) | LDD3495 | [4] | |
|
DBIA Probe Info |
 |
C663(1.21) | LDD3312 | [5] | |
|
AHL-Pu-1 Probe Info |
 |
C601(2.72) | LDD0170 | [6] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [7] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [7] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [8] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [9] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C004 Probe Info |
 |
8.75 | LDD1714 | [10] | |
|
C022 Probe Info |
 |
27.28 | LDD1728 | [10] | |
|
C048 Probe Info |
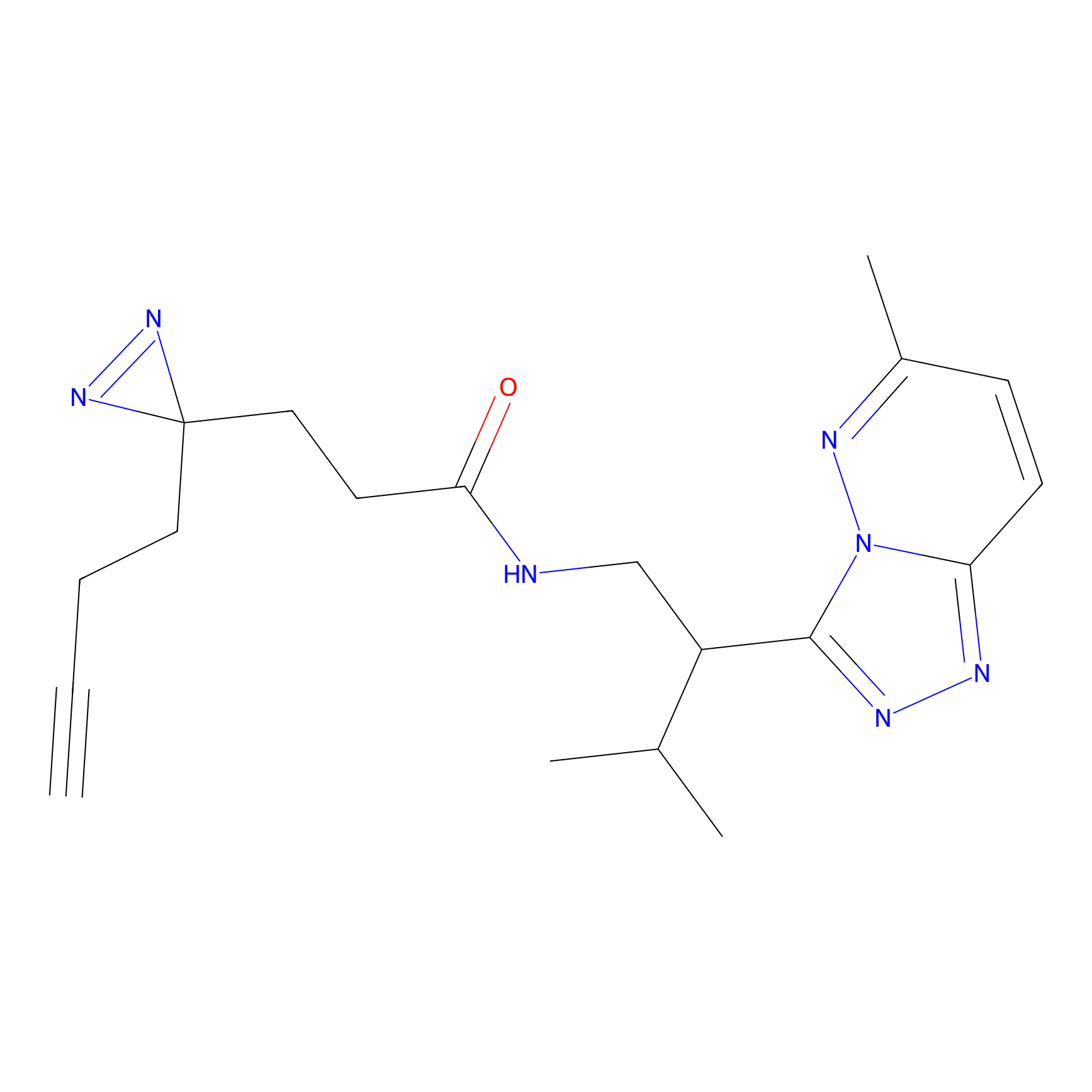 |
7.31 | LDD1746 | [10] | |
|
C106 Probe Info |
 |
59.30 | LDD1793 | [10] | |
|
C153 Probe Info |
 |
19.70 | LDD1834 | [10] | |
|
C159 Probe Info |
 |
11.08 | LDD1839 | [10] | |
|
C161 Probe Info |
 |
16.11 | LDD1841 | [10] | |
|
C169 Probe Info |
 |
23.10 | LDD1849 | [10] | |
|
C210 Probe Info |
 |
66.72 | LDD1884 | [10] | |
|
C282 Probe Info |
 |
46.85 | LDD1952 | [10] | |
|
C305 Probe Info |
 |
24.93 | LDD1974 | [10] | |
|
C313 Probe Info |
 |
22.78 | LDD1980 | [10] | |
|
C326 Probe Info |
 |
9.58 | LDD1990 | [10] | |
|
C349 Probe Info |
 |
10.63 | LDD2010 | [10] | |
|
C350 Probe Info |
 |
43.11 | LDD2011 | [10] | |
|
C353 Probe Info |
 |
18.77 | LDD2014 | [10] | |
|
C361 Probe Info |
 |
24.59 | LDD2022 | [10] | |
|
C397 Probe Info |
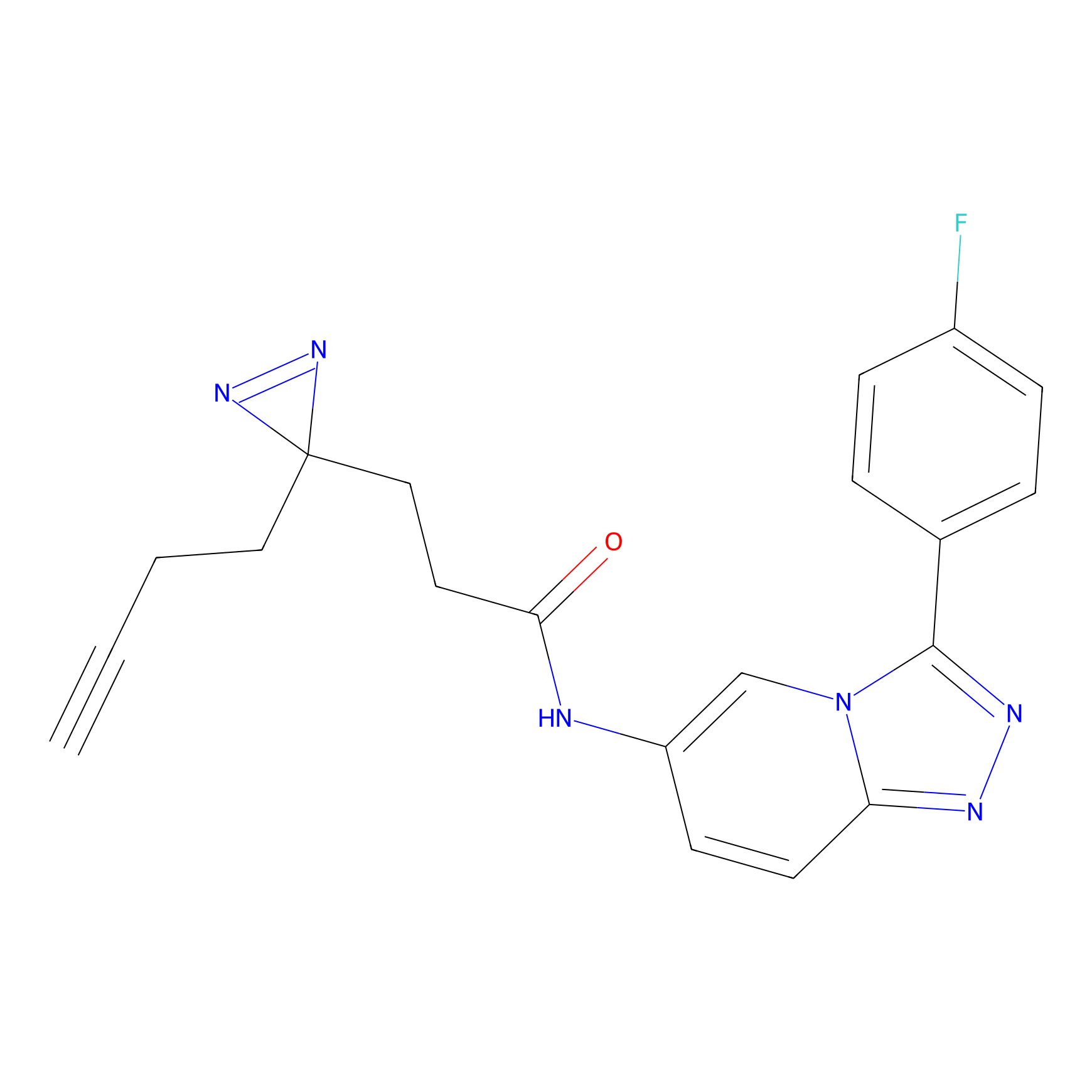 |
15.35 | LDD2056 | [10] | |
|
C399 Probe Info |
 |
7.26 | LDD2058 | [10] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | DM93 | C601(2.72) | LDD0170 | [6] |
| LDCM0026 | 4SU-RNA+native RNA | DM93 | C601(6.41) | LDD0171 | [6] |
| LDCM0214 | AC1 | HEK-293T | C601(1.01) | LDD1507 | [11] |
| LDCM0276 | AC17 | HEK-293T | C601(1.40) | LDD1515 | [11] |
| LDCM0285 | AC25 | HEK-293T | C601(1.39) | LDD1524 | [11] |
| LDCM0294 | AC33 | HEK-293T | C601(1.14) | LDD1533 | [11] |
| LDCM0303 | AC41 | HEK-293T | C601(1.15) | LDD1542 | [11] |
| LDCM0311 | AC49 | HEK-293T | C601(1.11) | LDD1550 | [11] |
| LDCM0320 | AC57 | HEK-293T | C601(0.87) | LDD1559 | [11] |
| LDCM0356 | AKOS034007680 | HEK-293T | C601(1.03) | LDD1570 | [11] |
| LDCM0156 | Aniline | NCI-H1299 | 11.52 | LDD0403 | [1] |
| LDCM0630 | CCW28-3 | 231MFP | C601(1.42) | LDD2214 | [12] |
| LDCM0404 | CL17 | HEK-293T | C601(1.17) | LDD1608 | [11] |
| LDCM0417 | CL29 | HEK-293T | C601(1.08) | LDD1621 | [11] |
| LDCM0431 | CL41 | HEK-293T | C601(1.11) | LDD1635 | [11] |
| LDCM0440 | CL5 | HEK-293T | C601(1.15) | LDD1644 | [11] |
| LDCM0444 | CL53 | HEK-293T | C601(1.27) | LDD1647 | [11] |
| LDCM0457 | CL65 | HEK-293T | C601(1.22) | LDD1660 | [11] |
| LDCM0470 | CL77 | HEK-293T | C601(1.27) | LDD1673 | [11] |
| LDCM0483 | CL89 | HEK-293T | C601(1.70) | LDD1686 | [11] |
| LDCM0022 | KB02 | 22RV1 | C663(1.43) | LDD2243 | [5] |
| LDCM0023 | KB03 | 22RV1 | C663(1.54) | LDD2660 | [5] |
| LDCM0024 | KB05 | HMCB | C663(1.21) | LDD3312 | [5] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
Other
References
