Details of the Target
General Information of Target
| Target ID | LDTP03011 | |||||
|---|---|---|---|---|---|---|
| Target Name | Casein kinase II subunit alpha' (CSNK2A2) | |||||
| Gene Name | CSNK2A2 | |||||
| Gene ID | 1459 | |||||
| Synonyms |
CK2A2; Casein kinase II subunit alpha'; CK II alpha'; EC 2.7.11.1 |
|||||
| 3D Structure | ||||||
| Sequence |
MPGPAAGSRARVYAEVNSLRSREYWDYEAHVPSWGNQDDYQLVRKLGRGKYSEVFEAINI
TNNERVVVKILKPVKKKKIKREVKILENLRGGTNIIKLIDTVKDPVSKTPALVFEYINNT DFKQLYQILTDFDIRFYMYELLKALDYCHSKGIMHRDVKPHNVMIDHQQKKLRLIDWGLA EFYHPAQEYNVRVASRYFKGPELLVDYQMYDYSLDMWSLGCMLASMIFRREPFFHGQDNY DQLVRIAKVLGTEELYGYLKKYHIDLDPHFNDILGQHSRKRWENFIHSENRHLVSPEALD LLDKLLRYDHQQRLTAKEAMEHPYFYPVVKEQSQPCADNAVLSSGLTAAR |
|||||
| Target Type |
Patented-recorded
|
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Protein kinase superfamily, Ser/Thr protein kinase family, CK2 subfamily
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Catalytic subunit of a constitutively active serine/threonine-protein kinase complex that phosphorylates a large number of substrates containing acidic residues C-terminal to the phosphorylated serine or threonine. Regulates numerous cellular processes, such as cell cycle progression, apoptosis and transcription, as well as viral infection. May act as a regulatory node which integrates and coordinates numerous signals leading to an appropriate cellular response. During mitosis, functions as a component of the p53/TP53-dependent spindle assembly checkpoint (SAC) that maintains cyclin-B-CDK1 activity and G2 arrest in response to spindle damage. Also required for p53/TP53-mediated apoptosis, phosphorylating 'Ser-392' of p53/TP53 following UV irradiation. Phosphorylates a number of DNA repair proteins in response to DNA damage, such as MDC1, RAD9A, RAD51 and HTATSF1, promoting their recruitment to DNA damage sites . Can also negatively regulate apoptosis. Phosphorylates the caspases CASP9 and CASP2 and the apoptotic regulator NOL3. Phosphorylation protects CASP9 from cleavage and activation by CASP8, and inhibits the dimerization of CASP2 and activation of CASP8. Regulates transcription by direct phosphorylation of RNA polymerases I, II, III and IV. Also phosphorylates and regulates numerous transcription factors including NF-kappa-B, STAT1, CREB1, IRF1, IRF2, ATF1, SRF, MAX, JUN, FOS, MYC and MYB. Phosphorylates Hsp90 and its co-chaperones FKBP4 and CDC37, which is essential for chaperone function. Regulates Wnt signaling by phosphorylating CTNNB1 and the transcription factor LEF1. Acts as an ectokinase that phosphorylates several extracellular proteins. During viral infection, phosphorylates various proteins involved in the viral life cycles of EBV, HSV, HBV, HCV, HIV, CMV and HPV.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
TH211 Probe Info |
 |
Y308(20.00) | LDD0260 | [2] | |
|
C-Sul Probe Info |
 |
8.38 | LDD0066 | [3] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [4] | |
|
ONAyne Probe Info |
 |
K103(10.00) | LDD0275 | [5] | |
|
BTD Probe Info |
 |
C148(3.25) | LDD1700 | [6] | |
|
D5yne Probe Info |
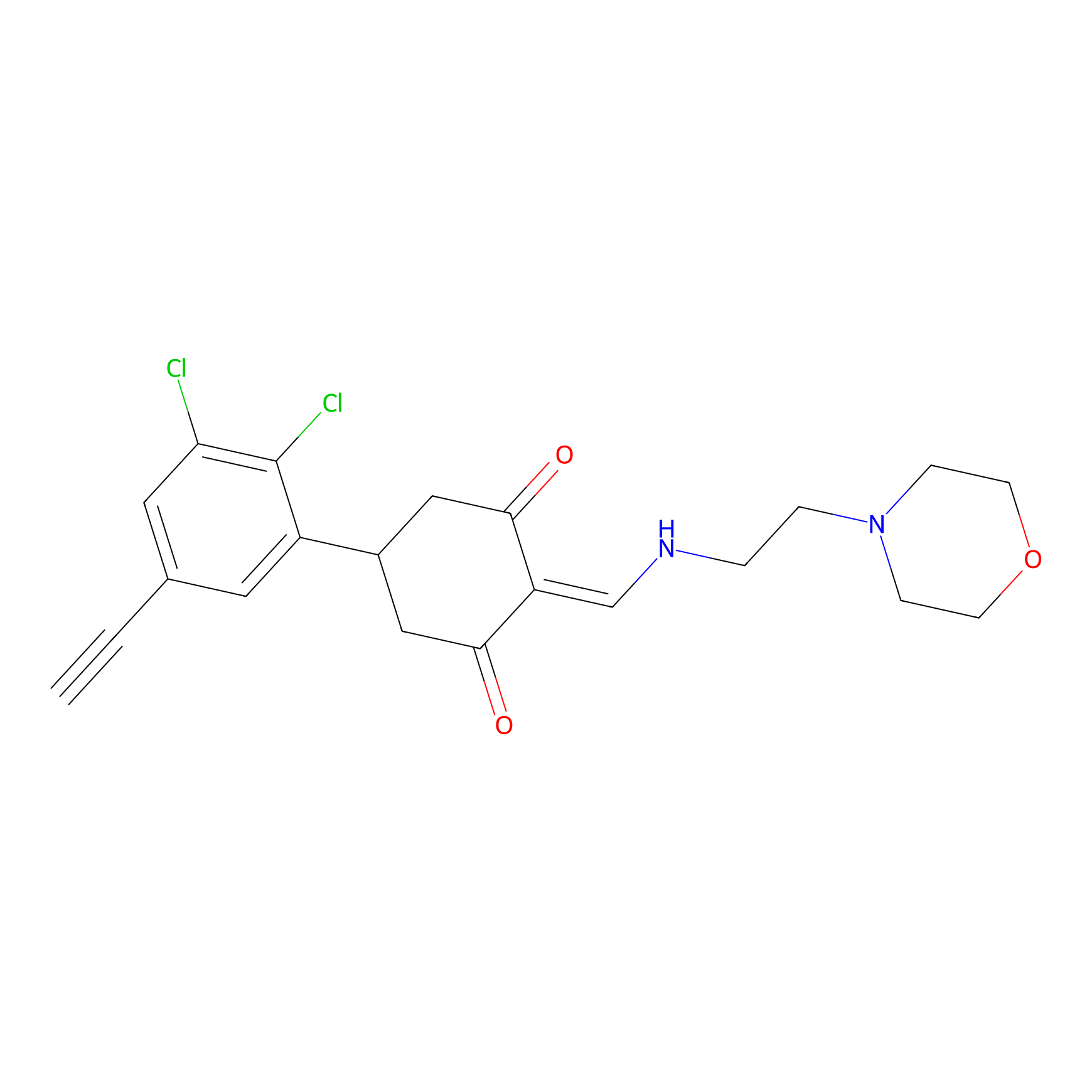 |
1.65 | LDD0055 | [7] | |
|
DA-P3 Probe Info |
 |
6.89 | LDD0183 | [8] | |
|
AHL-Pu-1 Probe Info |
 |
C336(2.26) | LDD0168 | [9] | |
|
Acrolein Probe Info |
 |
H263(0.00); C221(0.00) | LDD0222 | [10] | |
|
DBIA Probe Info |
 |
C336(0.95) | LDD0078 | [11] | |
|
ATP probe Probe Info |
 |
K248(0.00); K103(0.00); K260(0.00); K261(0.00) | LDD0199 | [12] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C148(0.00); C336(0.00) | LDD0038 | [13] | |
|
IA-alkyne Probe Info |
 |
C148(0.00); C336(0.00) | LDD0036 | [13] | |
|
Lodoacetamide azide Probe Info |
 |
C148(0.00); C336(0.00) | LDD0037 | [13] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [14] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [14] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [14] | |
|
NHS Probe Info |
 |
K103(0.00); K260(0.00) | LDD0010 | [15] | |
|
PPMS Probe Info |
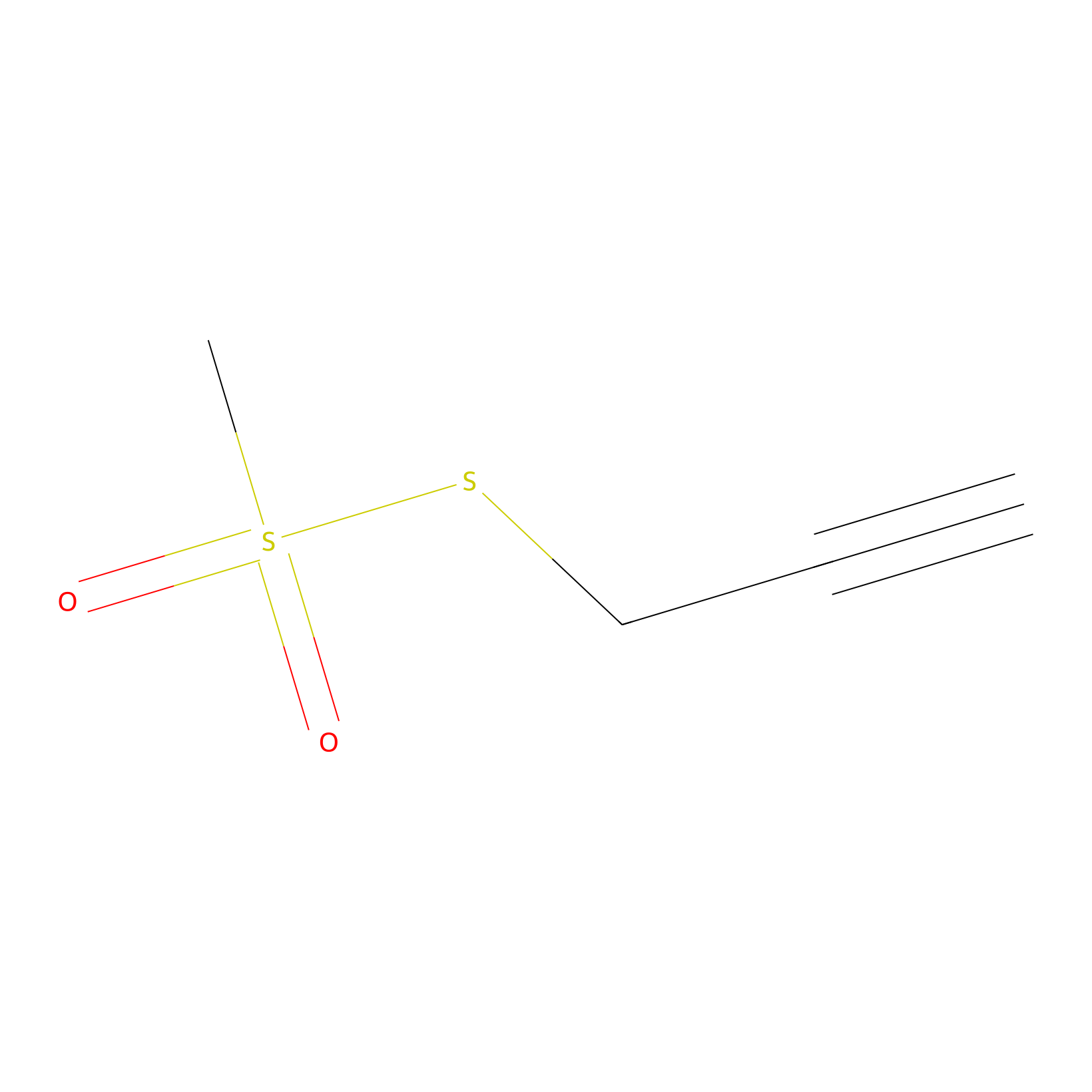 |
N.A. | LDD0008 | [15] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [16] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [15] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [17] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [18] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [10] | |
|
W1 Probe Info |
 |
C336(0.00); K103(0.00); S107(0.00) | LDD0236 | [19] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [20] | |
|
HHS-465 Probe Info |
 |
N.A. | LDD2240 | [21] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe12 Probe Info |
 |
7.62 | LDD0473 | [22] | |
|
FFF probe13 Probe Info |
 |
15.61 | LDD0475 | [22] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0464 | [22] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0137 | [23] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [23] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C148(1.15) | LDD2117 | [6] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C148(1.49) | LDD2152 | [6] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C148(1.00) | LDD2103 | [6] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C148(0.82) | LDD2132 | [6] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C148(1.15) | LDD2131 | [6] |
| LDCM0025 | 4SU-RNA | HEK-293T | C336(2.26) | LDD0168 | [9] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C336(2.04) | LDD0169 | [9] |
| LDCM0214 | AC1 | HCT 116 | C336(1.17) | LDD0531 | [11] |
| LDCM0215 | AC10 | HCT 116 | C336(1.17) | LDD0532 | [11] |
| LDCM0216 | AC100 | HCT 116 | C336(1.04) | LDD0533 | [11] |
| LDCM0217 | AC101 | HCT 116 | C336(1.00) | LDD0534 | [11] |
| LDCM0218 | AC102 | HCT 116 | C336(1.04) | LDD0535 | [11] |
| LDCM0219 | AC103 | HCT 116 | C336(1.22) | LDD0536 | [11] |
| LDCM0220 | AC104 | HCT 116 | C336(1.06) | LDD0537 | [11] |
| LDCM0221 | AC105 | HCT 116 | C336(1.10) | LDD0538 | [11] |
| LDCM0222 | AC106 | HCT 116 | C336(1.14) | LDD0539 | [11] |
| LDCM0223 | AC107 | HCT 116 | C336(0.91) | LDD0540 | [11] |
| LDCM0224 | AC108 | HCT 116 | C336(0.91) | LDD0541 | [11] |
| LDCM0225 | AC109 | HCT 116 | C336(0.80) | LDD0542 | [11] |
| LDCM0226 | AC11 | HCT 116 | C336(1.17) | LDD0543 | [11] |
| LDCM0227 | AC110 | HCT 116 | C336(0.93) | LDD0544 | [11] |
| LDCM0228 | AC111 | HCT 116 | C336(0.96) | LDD0545 | [11] |
| LDCM0229 | AC112 | HCT 116 | C336(0.93) | LDD0546 | [11] |
| LDCM0230 | AC113 | HCT 116 | C336(1.07) | LDD0547 | [11] |
| LDCM0231 | AC114 | HCT 116 | C336(1.07) | LDD0548 | [11] |
| LDCM0232 | AC115 | HCT 116 | C336(1.35) | LDD0549 | [11] |
| LDCM0233 | AC116 | HCT 116 | C336(1.33) | LDD0550 | [11] |
| LDCM0234 | AC117 | HCT 116 | C336(1.25) | LDD0551 | [11] |
| LDCM0235 | AC118 | HCT 116 | C336(1.12) | LDD0552 | [11] |
| LDCM0236 | AC119 | HCT 116 | C336(1.21) | LDD0553 | [11] |
| LDCM0237 | AC12 | HCT 116 | C336(0.95) | LDD0554 | [11] |
| LDCM0238 | AC120 | HCT 116 | C336(1.25) | LDD0555 | [11] |
| LDCM0239 | AC121 | HCT 116 | C336(1.16) | LDD0556 | [11] |
| LDCM0240 | AC122 | HCT 116 | C336(1.20) | LDD0557 | [11] |
| LDCM0241 | AC123 | HCT 116 | C336(1.15) | LDD0558 | [11] |
| LDCM0242 | AC124 | HCT 116 | C336(1.10) | LDD0559 | [11] |
| LDCM0243 | AC125 | HCT 116 | C336(1.00) | LDD0560 | [11] |
| LDCM0244 | AC126 | HCT 116 | C336(1.28) | LDD0561 | [11] |
| LDCM0245 | AC127 | HCT 116 | C336(1.29) | LDD0562 | [11] |
| LDCM0246 | AC128 | HCT 116 | C336(1.01) | LDD0563 | [11] |
| LDCM0247 | AC129 | HCT 116 | C336(0.97) | LDD0564 | [11] |
| LDCM0249 | AC130 | HCT 116 | C336(1.30) | LDD0566 | [11] |
| LDCM0250 | AC131 | HCT 116 | C336(0.92) | LDD0567 | [11] |
| LDCM0251 | AC132 | HCT 116 | C336(1.10) | LDD0568 | [11] |
| LDCM0252 | AC133 | HCT 116 | C336(1.08) | LDD0569 | [11] |
| LDCM0253 | AC134 | HCT 116 | C336(1.26) | LDD0570 | [11] |
| LDCM0254 | AC135 | HCT 116 | C336(1.24) | LDD0571 | [11] |
| LDCM0255 | AC136 | HCT 116 | C336(1.36) | LDD0572 | [11] |
| LDCM0256 | AC137 | HCT 116 | C336(1.36) | LDD0573 | [11] |
| LDCM0257 | AC138 | HCT 116 | C336(1.49) | LDD0574 | [11] |
| LDCM0258 | AC139 | HCT 116 | C336(1.38) | LDD0575 | [11] |
| LDCM0259 | AC14 | HCT 116 | C336(1.08) | LDD0576 | [11] |
| LDCM0260 | AC140 | HCT 116 | C336(1.78) | LDD0577 | [11] |
| LDCM0261 | AC141 | HCT 116 | C336(1.36) | LDD0578 | [11] |
| LDCM0262 | AC142 | HCT 116 | C336(1.12) | LDD0579 | [11] |
| LDCM0263 | AC143 | HCT 116 | C336(1.16) | LDD0580 | [11] |
| LDCM0264 | AC144 | HCT 116 | C336(1.20) | LDD0581 | [11] |
| LDCM0265 | AC145 | HCT 116 | C336(1.23) | LDD0582 | [11] |
| LDCM0266 | AC146 | HCT 116 | C336(1.36) | LDD0583 | [11] |
| LDCM0267 | AC147 | HCT 116 | C336(1.33) | LDD0584 | [11] |
| LDCM0268 | AC148 | HCT 116 | C336(1.54) | LDD0585 | [11] |
| LDCM0269 | AC149 | HCT 116 | C336(1.49) | LDD0586 | [11] |
| LDCM0270 | AC15 | HCT 116 | C336(1.13) | LDD0587 | [11] |
| LDCM0271 | AC150 | HCT 116 | C336(1.12) | LDD0588 | [11] |
| LDCM0272 | AC151 | HCT 116 | C336(1.19) | LDD0589 | [11] |
| LDCM0273 | AC152 | HCT 116 | C336(1.40) | LDD0590 | [11] |
| LDCM0274 | AC153 | HCT 116 | C336(1.67) | LDD0591 | [11] |
| LDCM0621 | AC154 | HCT 116 | C336(1.21) | LDD2158 | [11] |
| LDCM0622 | AC155 | HCT 116 | C336(1.31) | LDD2159 | [11] |
| LDCM0623 | AC156 | HCT 116 | C336(1.04) | LDD2160 | [11] |
| LDCM0624 | AC157 | HCT 116 | C336(1.01) | LDD2161 | [11] |
| LDCM0276 | AC17 | HCT 116 | C336(1.10) | LDD0593 | [11] |
| LDCM0277 | AC18 | HCT 116 | C336(1.49) | LDD0594 | [11] |
| LDCM0278 | AC19 | HCT 116 | C336(1.14) | LDD0595 | [11] |
| LDCM0279 | AC2 | HCT 116 | C336(1.06) | LDD0596 | [11] |
| LDCM0280 | AC20 | HCT 116 | C336(1.13) | LDD0597 | [11] |
| LDCM0281 | AC21 | HCT 116 | C336(1.14) | LDD0598 | [11] |
| LDCM0282 | AC22 | HCT 116 | C336(1.05) | LDD0599 | [11] |
| LDCM0283 | AC23 | HCT 116 | C336(1.04) | LDD0600 | [11] |
| LDCM0284 | AC24 | HCT 116 | C336(1.07) | LDD0601 | [11] |
| LDCM0285 | AC25 | HCT 116 | C336(1.04) | LDD0602 | [11] |
| LDCM0286 | AC26 | HCT 116 | C336(1.13) | LDD0603 | [11] |
| LDCM0287 | AC27 | HCT 116 | C336(1.17) | LDD0604 | [11] |
| LDCM0288 | AC28 | HCT 116 | C336(1.30) | LDD0605 | [11] |
| LDCM0289 | AC29 | HCT 116 | C336(1.27) | LDD0606 | [11] |
| LDCM0290 | AC3 | HCT 116 | C336(0.99) | LDD0607 | [11] |
| LDCM0291 | AC30 | HCT 116 | C336(1.35) | LDD0608 | [11] |
| LDCM0292 | AC31 | HCT 116 | C336(1.16) | LDD0609 | [11] |
| LDCM0293 | AC32 | HCT 116 | C336(1.52) | LDD0610 | [11] |
| LDCM0294 | AC33 | HCT 116 | C336(1.25) | LDD0611 | [11] |
| LDCM0295 | AC34 | HCT 116 | C336(1.39) | LDD0612 | [11] |
| LDCM0296 | AC35 | HCT 116 | C336(0.88) | LDD0613 | [11] |
| LDCM0297 | AC36 | HCT 116 | C336(0.96) | LDD0614 | [11] |
| LDCM0298 | AC37 | HCT 116 | C336(0.89) | LDD0615 | [11] |
| LDCM0299 | AC38 | HCT 116 | C336(0.98) | LDD0616 | [11] |
| LDCM0300 | AC39 | HCT 116 | C336(1.11) | LDD0617 | [11] |
| LDCM0301 | AC4 | HCT 116 | C336(1.12) | LDD0618 | [11] |
| LDCM0302 | AC40 | HCT 116 | C336(1.07) | LDD0619 | [11] |
| LDCM0303 | AC41 | HCT 116 | C336(1.02) | LDD0620 | [11] |
| LDCM0304 | AC42 | HCT 116 | C336(1.05) | LDD0621 | [11] |
| LDCM0305 | AC43 | HCT 116 | C336(1.13) | LDD0622 | [11] |
| LDCM0306 | AC44 | HCT 116 | C336(1.12) | LDD0623 | [11] |
| LDCM0307 | AC45 | HCT 116 | C336(1.15) | LDD0624 | [11] |
| LDCM0308 | AC46 | HCT 116 | C336(1.13) | LDD0625 | [11] |
| LDCM0309 | AC47 | HCT 116 | C336(1.09) | LDD0626 | [11] |
| LDCM0310 | AC48 | HCT 116 | C336(1.17) | LDD0627 | [11] |
| LDCM0311 | AC49 | HCT 116 | C336(1.46) | LDD0628 | [11] |
| LDCM0312 | AC5 | HCT 116 | C336(1.07) | LDD0629 | [11] |
| LDCM0313 | AC50 | HCT 116 | C336(1.51) | LDD0630 | [11] |
| LDCM0314 | AC51 | HCT 116 | C336(0.99) | LDD0631 | [11] |
| LDCM0315 | AC52 | HCT 116 | C336(1.04) | LDD0632 | [11] |
| LDCM0316 | AC53 | HCT 116 | C336(1.19) | LDD0633 | [11] |
| LDCM0317 | AC54 | HCT 116 | C336(1.17) | LDD0634 | [11] |
| LDCM0318 | AC55 | HCT 116 | C336(1.41) | LDD0635 | [11] |
| LDCM0319 | AC56 | HCT 116 | C336(1.74) | LDD0636 | [11] |
| LDCM0320 | AC57 | HCT 116 | C336(1.36) | LDD0637 | [11] |
| LDCM0321 | AC58 | HCT 116 | C336(1.39) | LDD0638 | [11] |
| LDCM0322 | AC59 | HCT 116 | C336(1.58) | LDD0639 | [11] |
| LDCM0323 | AC6 | HCT 116 | C336(1.18) | LDD0640 | [11] |
| LDCM0324 | AC60 | HCT 116 | C336(1.41) | LDD0641 | [11] |
| LDCM0325 | AC61 | HCT 116 | C336(1.24) | LDD0642 | [11] |
| LDCM0326 | AC62 | HCT 116 | C336(1.43) | LDD0643 | [11] |
| LDCM0327 | AC63 | HCT 116 | C336(1.28) | LDD0644 | [11] |
| LDCM0328 | AC64 | HCT 116 | C336(1.47) | LDD0645 | [11] |
| LDCM0329 | AC65 | HCT 116 | C336(1.53) | LDD0646 | [11] |
| LDCM0330 | AC66 | HCT 116 | C336(1.43) | LDD0647 | [11] |
| LDCM0331 | AC67 | HCT 116 | C336(1.61) | LDD0648 | [11] |
| LDCM0332 | AC68 | HCT 116 | C336(1.11) | LDD0649 | [11] |
| LDCM0333 | AC69 | HCT 116 | C336(1.08) | LDD0650 | [11] |
| LDCM0334 | AC7 | HCT 116 | C336(1.01) | LDD0651 | [11] |
| LDCM0335 | AC70 | HCT 116 | C336(1.13) | LDD0652 | [11] |
| LDCM0336 | AC71 | HCT 116 | C336(0.94) | LDD0653 | [11] |
| LDCM0337 | AC72 | HCT 116 | C336(1.12) | LDD0654 | [11] |
| LDCM0338 | AC73 | HCT 116 | C336(1.48) | LDD0655 | [11] |
| LDCM0339 | AC74 | HCT 116 | C336(1.35) | LDD0656 | [11] |
| LDCM0340 | AC75 | HCT 116 | C336(1.43) | LDD0657 | [11] |
| LDCM0341 | AC76 | HCT 116 | C336(1.15) | LDD0658 | [11] |
| LDCM0342 | AC77 | HCT 116 | C336(1.15) | LDD0659 | [11] |
| LDCM0343 | AC78 | HCT 116 | C336(1.05) | LDD0660 | [11] |
| LDCM0344 | AC79 | HCT 116 | C336(1.08) | LDD0661 | [11] |
| LDCM0345 | AC8 | HCT 116 | C336(1.21) | LDD0662 | [11] |
| LDCM0346 | AC80 | HCT 116 | C336(1.05) | LDD0663 | [11] |
| LDCM0347 | AC81 | HCT 116 | C336(1.04) | LDD0664 | [11] |
| LDCM0348 | AC82 | HCT 116 | C336(1.47) | LDD0665 | [11] |
| LDCM0349 | AC83 | HCT 116 | C336(1.55) | LDD0666 | [11] |
| LDCM0350 | AC84 | HCT 116 | C336(1.65) | LDD0667 | [11] |
| LDCM0351 | AC85 | HCT 116 | C336(1.15) | LDD0668 | [11] |
| LDCM0352 | AC86 | HCT 116 | C336(1.15) | LDD0669 | [11] |
| LDCM0353 | AC87 | HCT 116 | C336(1.03) | LDD0670 | [11] |
| LDCM0354 | AC88 | HCT 116 | C336(1.06) | LDD0671 | [11] |
| LDCM0355 | AC89 | HCT 116 | C336(1.09) | LDD0672 | [11] |
| LDCM0357 | AC90 | HCT 116 | C336(1.01) | LDD0674 | [11] |
| LDCM0358 | AC91 | HCT 116 | C336(1.68) | LDD0675 | [11] |
| LDCM0359 | AC92 | HCT 116 | C336(1.38) | LDD0676 | [11] |
| LDCM0360 | AC93 | HCT 116 | C336(1.09) | LDD0677 | [11] |
| LDCM0361 | AC94 | HCT 116 | C336(1.13) | LDD0678 | [11] |
| LDCM0362 | AC95 | HCT 116 | C336(1.10) | LDD0679 | [11] |
| LDCM0363 | AC96 | HCT 116 | C336(1.19) | LDD0680 | [11] |
| LDCM0364 | AC97 | HCT 116 | C336(1.29) | LDD0681 | [11] |
| LDCM0365 | AC98 | HCT 116 | C336(1.41) | LDD0682 | [11] |
| LDCM0366 | AC99 | HCT 116 | C336(0.98) | LDD0683 | [11] |
| LDCM0545 | Acetamide | MDA-MB-231 | C148(0.56) | LDD2138 | [6] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C148(0.98) | LDD2113 | [6] |
| LDCM0248 | AKOS034007472 | HCT 116 | C336(0.94) | LDD0565 | [11] |
| LDCM0356 | AKOS034007680 | HCT 116 | C336(1.06) | LDD0673 | [11] |
| LDCM0275 | AKOS034007705 | HCT 116 | C336(1.65) | LDD0592 | [11] |
| LDCM0156 | Aniline | NCI-H1299 | 15.00 | LDD0403 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C336(0.95) | LDD0078 | [11] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C148(0.57) | LDD2091 | [6] |
| LDCM0108 | Chloroacetamide | HeLa | H263(0.00); C221(0.00) | LDD0222 | [10] |
| LDCM0367 | CL1 | HCT 116 | C336(0.87) | LDD0684 | [11] |
| LDCM0368 | CL10 | HCT 116 | C336(0.87) | LDD0685 | [11] |
| LDCM0369 | CL100 | HCT 116 | C336(1.12) | LDD0686 | [11] |
| LDCM0370 | CL101 | HCT 116 | C336(1.13) | LDD0687 | [11] |
| LDCM0371 | CL102 | HCT 116 | C336(0.95) | LDD0688 | [11] |
| LDCM0372 | CL103 | HCT 116 | C336(0.99) | LDD0689 | [11] |
| LDCM0373 | CL104 | HCT 116 | C336(0.97) | LDD0690 | [11] |
| LDCM0374 | CL105 | HCT 116 | C336(1.23) | LDD0691 | [11] |
| LDCM0375 | CL106 | HCT 116 | C336(1.44) | LDD0692 | [11] |
| LDCM0376 | CL107 | HCT 116 | C336(1.27) | LDD0693 | [11] |
| LDCM0377 | CL108 | HCT 116 | C336(1.39) | LDD0694 | [11] |
| LDCM0378 | CL109 | HCT 116 | C336(1.12) | LDD0695 | [11] |
| LDCM0379 | CL11 | HCT 116 | C336(1.26) | LDD0696 | [11] |
| LDCM0380 | CL110 | HCT 116 | C336(1.14) | LDD0697 | [11] |
| LDCM0381 | CL111 | HCT 116 | C336(1.22) | LDD0698 | [11] |
| LDCM0382 | CL112 | HCT 116 | C336(1.03) | LDD0699 | [11] |
| LDCM0383 | CL113 | HCT 116 | C336(1.31) | LDD0700 | [11] |
| LDCM0384 | CL114 | HCT 116 | C336(1.15) | LDD0701 | [11] |
| LDCM0385 | CL115 | HCT 116 | C336(1.27) | LDD0702 | [11] |
| LDCM0386 | CL116 | HCT 116 | C336(1.21) | LDD0703 | [11] |
| LDCM0387 | CL117 | HCT 116 | C336(1.38) | LDD0704 | [11] |
| LDCM0388 | CL118 | HCT 116 | C336(1.05) | LDD0705 | [11] |
| LDCM0389 | CL119 | HCT 116 | C336(1.12) | LDD0706 | [11] |
| LDCM0390 | CL12 | HCT 116 | C336(1.30) | LDD0707 | [11] |
| LDCM0391 | CL120 | HCT 116 | C336(1.02) | LDD0708 | [11] |
| LDCM0392 | CL121 | HCT 116 | C336(1.01) | LDD0709 | [11] |
| LDCM0393 | CL122 | HCT 116 | C336(1.19) | LDD0710 | [11] |
| LDCM0394 | CL123 | HCT 116 | C336(1.28) | LDD0711 | [11] |
| LDCM0395 | CL124 | HCT 116 | C336(1.29) | LDD0712 | [11] |
| LDCM0396 | CL125 | HCT 116 | C336(1.06) | LDD0713 | [11] |
| LDCM0397 | CL126 | HCT 116 | C336(1.06) | LDD0714 | [11] |
| LDCM0398 | CL127 | HCT 116 | C336(1.13) | LDD0715 | [11] |
| LDCM0399 | CL128 | HCT 116 | C336(1.29) | LDD0716 | [11] |
| LDCM0400 | CL13 | HCT 116 | C336(1.36) | LDD0717 | [11] |
| LDCM0401 | CL14 | HCT 116 | C336(0.84) | LDD0718 | [11] |
| LDCM0402 | CL15 | HCT 116 | C336(0.95) | LDD0719 | [11] |
| LDCM0403 | CL16 | HCT 116 | C336(1.06) | LDD0720 | [11] |
| LDCM0404 | CL17 | HCT 116 | C336(1.01) | LDD0721 | [11] |
| LDCM0405 | CL18 | HCT 116 | C336(1.25) | LDD0722 | [11] |
| LDCM0406 | CL19 | HCT 116 | C336(1.17) | LDD0723 | [11] |
| LDCM0407 | CL2 | HCT 116 | C336(0.79) | LDD0724 | [11] |
| LDCM0408 | CL20 | HCT 116 | C336(1.19) | LDD0725 | [11] |
| LDCM0409 | CL21 | HCT 116 | C336(1.28) | LDD0726 | [11] |
| LDCM0410 | CL22 | HCT 116 | C336(1.76) | LDD0727 | [11] |
| LDCM0411 | CL23 | HCT 116 | C336(1.16) | LDD0728 | [11] |
| LDCM0412 | CL24 | HCT 116 | C336(1.41) | LDD0729 | [11] |
| LDCM0413 | CL25 | HCT 116 | C336(1.19) | LDD0730 | [11] |
| LDCM0414 | CL26 | HCT 116 | C336(1.24) | LDD0731 | [11] |
| LDCM0415 | CL27 | HCT 116 | C336(1.07) | LDD0732 | [11] |
| LDCM0416 | CL28 | HCT 116 | C336(1.20) | LDD0733 | [11] |
| LDCM0417 | CL29 | HCT 116 | C336(1.21) | LDD0734 | [11] |
| LDCM0418 | CL3 | HCT 116 | C336(0.96) | LDD0735 | [11] |
| LDCM0419 | CL30 | HCT 116 | C336(1.01) | LDD0736 | [11] |
| LDCM0420 | CL31 | HCT 116 | C336(0.98) | LDD0737 | [11] |
| LDCM0421 | CL32 | HCT 116 | C336(1.13) | LDD0738 | [11] |
| LDCM0422 | CL33 | HCT 116 | C336(0.95) | LDD0739 | [11] |
| LDCM0423 | CL34 | HCT 116 | C336(1.23) | LDD0740 | [11] |
| LDCM0424 | CL35 | HCT 116 | C336(1.19) | LDD0741 | [11] |
| LDCM0425 | CL36 | HCT 116 | C336(1.04) | LDD0742 | [11] |
| LDCM0426 | CL37 | HCT 116 | C336(1.12) | LDD0743 | [11] |
| LDCM0428 | CL39 | HCT 116 | C336(1.29) | LDD0745 | [11] |
| LDCM0429 | CL4 | HCT 116 | C336(0.99) | LDD0746 | [11] |
| LDCM0430 | CL40 | HCT 116 | C336(1.10) | LDD0747 | [11] |
| LDCM0431 | CL41 | HCT 116 | C336(1.16) | LDD0748 | [11] |
| LDCM0432 | CL42 | HCT 116 | C336(1.45) | LDD0749 | [11] |
| LDCM0433 | CL43 | HCT 116 | C336(1.16) | LDD0750 | [11] |
| LDCM0434 | CL44 | HCT 116 | C336(1.11) | LDD0751 | [11] |
| LDCM0435 | CL45 | HCT 116 | C336(1.23) | LDD0752 | [11] |
| LDCM0436 | CL46 | HCT 116 | C336(0.88) | LDD0753 | [11] |
| LDCM0437 | CL47 | HCT 116 | C336(0.91) | LDD0754 | [11] |
| LDCM0438 | CL48 | HCT 116 | C336(0.84) | LDD0755 | [11] |
| LDCM0439 | CL49 | HCT 116 | C336(0.88) | LDD0756 | [11] |
| LDCM0440 | CL5 | HCT 116 | C336(0.96) | LDD0757 | [11] |
| LDCM0441 | CL50 | HCT 116 | C336(0.87) | LDD0758 | [11] |
| LDCM0442 | CL51 | HCT 116 | C336(0.84) | LDD0759 | [11] |
| LDCM0443 | CL52 | HCT 116 | C336(0.95) | LDD0760 | [11] |
| LDCM0444 | CL53 | HCT 116 | C336(1.00) | LDD0761 | [11] |
| LDCM0445 | CL54 | HCT 116 | C336(0.89) | LDD0762 | [11] |
| LDCM0446 | CL55 | HCT 116 | C336(0.83) | LDD0763 | [11] |
| LDCM0447 | CL56 | HCT 116 | C336(0.94) | LDD0764 | [11] |
| LDCM0448 | CL57 | HCT 116 | C336(0.84) | LDD0765 | [11] |
| LDCM0449 | CL58 | HCT 116 | C336(0.83) | LDD0766 | [11] |
| LDCM0450 | CL59 | HCT 116 | C336(0.84) | LDD0767 | [11] |
| LDCM0451 | CL6 | HCT 116 | C336(1.00) | LDD0768 | [11] |
| LDCM0452 | CL60 | HCT 116 | C336(0.88) | LDD0769 | [11] |
| LDCM0453 | CL61 | HCT 116 | C336(0.96) | LDD0770 | [11] |
| LDCM0454 | CL62 | HCT 116 | C336(0.98) | LDD0771 | [11] |
| LDCM0455 | CL63 | HCT 116 | C336(1.06) | LDD0772 | [11] |
| LDCM0456 | CL64 | HCT 116 | C336(1.13) | LDD0773 | [11] |
| LDCM0457 | CL65 | HCT 116 | C336(0.95) | LDD0774 | [11] |
| LDCM0458 | CL66 | HCT 116 | C336(1.36) | LDD0775 | [11] |
| LDCM0459 | CL67 | HCT 116 | C336(1.05) | LDD0776 | [11] |
| LDCM0460 | CL68 | HCT 116 | C336(1.02) | LDD0777 | [11] |
| LDCM0461 | CL69 | HCT 116 | C336(1.06) | LDD0778 | [11] |
| LDCM0462 | CL7 | HCT 116 | C336(1.23) | LDD0779 | [11] |
| LDCM0463 | CL70 | HCT 116 | C336(1.12) | LDD0780 | [11] |
| LDCM0464 | CL71 | HCT 116 | C336(1.07) | LDD0781 | [11] |
| LDCM0465 | CL72 | HCT 116 | C336(1.08) | LDD0782 | [11] |
| LDCM0466 | CL73 | HCT 116 | C336(1.14) | LDD0783 | [11] |
| LDCM0467 | CL74 | HCT 116 | C336(1.03) | LDD0784 | [11] |
| LDCM0469 | CL76 | HCT 116 | C336(1.11) | LDD0786 | [11] |
| LDCM0470 | CL77 | HCT 116 | C336(1.02) | LDD0787 | [11] |
| LDCM0471 | CL78 | HCT 116 | C336(1.17) | LDD0788 | [11] |
| LDCM0472 | CL79 | HCT 116 | C336(1.32) | LDD0789 | [11] |
| LDCM0473 | CL8 | HCT 116 | C336(1.29) | LDD0790 | [11] |
| LDCM0474 | CL80 | HCT 116 | C336(1.13) | LDD0791 | [11] |
| LDCM0475 | CL81 | HCT 116 | C336(1.09) | LDD0792 | [11] |
| LDCM0476 | CL82 | HCT 116 | C336(1.63) | LDD0793 | [11] |
| LDCM0477 | CL83 | HCT 116 | C336(1.56) | LDD0794 | [11] |
| LDCM0478 | CL84 | HCT 116 | C336(1.87) | LDD0795 | [11] |
| LDCM0479 | CL85 | HCT 116 | C336(1.21) | LDD0796 | [11] |
| LDCM0480 | CL86 | HCT 116 | C336(1.07) | LDD0797 | [11] |
| LDCM0481 | CL87 | HCT 116 | C336(1.21) | LDD0798 | [11] |
| LDCM0482 | CL88 | HCT 116 | C336(1.48) | LDD0799 | [11] |
| LDCM0483 | CL89 | HCT 116 | C336(2.11) | LDD0800 | [11] |
| LDCM0484 | CL9 | HCT 116 | C336(0.99) | LDD0801 | [11] |
| LDCM0485 | CL90 | HCT 116 | C336(0.96) | LDD0802 | [11] |
| LDCM0486 | CL91 | HCT 116 | C336(1.17) | LDD0803 | [11] |
| LDCM0487 | CL92 | HCT 116 | C336(1.09) | LDD0804 | [11] |
| LDCM0488 | CL93 | HCT 116 | C336(1.00) | LDD0805 | [11] |
| LDCM0489 | CL94 | HCT 116 | C336(1.12) | LDD0806 | [11] |
| LDCM0490 | CL95 | HCT 116 | C336(1.12) | LDD0807 | [11] |
| LDCM0491 | CL96 | HCT 116 | C336(1.10) | LDD0808 | [11] |
| LDCM0492 | CL97 | HCT 116 | C336(1.14) | LDD0809 | [11] |
| LDCM0493 | CL98 | HCT 116 | C336(1.21) | LDD0810 | [11] |
| LDCM0494 | CL99 | HCT 116 | C336(1.14) | LDD0811 | [11] |
| LDCM0634 | CY-0357 | Hep-G2 | C336(50.00) | LDD2228 | [20] |
| LDCM0095 | DC-LC3IN-D5 | HeLa | 1.65 | LDD0055 | [7] |
| LDCM0495 | E2913 | HEK-293T | C336(0.96); C148(0.89) | LDD1698 | [24] |
| LDCM0031 | Epigallocatechin gallate | HEK-293T | 6.89 | LDD0183 | [8] |
| LDCM0625 | F8 | Ramos | C336(1.15) | LDD2187 | [25] |
| LDCM0572 | Fragment10 | Ramos | C336(1.07) | LDD2189 | [25] |
| LDCM0574 | Fragment12 | Ramos | C336(0.74) | LDD2191 | [25] |
| LDCM0575 | Fragment13 | Ramos | C336(0.90) | LDD2192 | [25] |
| LDCM0576 | Fragment14 | Ramos | C336(0.53) | LDD2193 | [25] |
| LDCM0579 | Fragment20 | Ramos | C336(0.62) | LDD2194 | [25] |
| LDCM0580 | Fragment21 | Ramos | C336(0.83) | LDD2195 | [25] |
| LDCM0582 | Fragment23 | Ramos | C336(1.00) | LDD2196 | [25] |
| LDCM0578 | Fragment27 | Ramos | C336(0.90) | LDD2197 | [25] |
| LDCM0586 | Fragment28 | Ramos | C336(0.98) | LDD2198 | [25] |
| LDCM0588 | Fragment30 | Ramos | C336(1.02) | LDD2199 | [25] |
| LDCM0589 | Fragment31 | Ramos | C336(0.80) | LDD2200 | [25] |
| LDCM0590 | Fragment32 | Ramos | C336(1.03) | LDD2201 | [25] |
| LDCM0468 | Fragment33 | HCT 116 | C336(0.95) | LDD0785 | [11] |
| LDCM0596 | Fragment38 | Ramos | C336(0.69) | LDD2203 | [25] |
| LDCM0566 | Fragment4 | Ramos | C336(0.82) | LDD2184 | [25] |
| LDCM0427 | Fragment51 | HCT 116 | C336(1.81) | LDD0744 | [11] |
| LDCM0610 | Fragment52 | Ramos | C336(1.15) | LDD2204 | [25] |
| LDCM0614 | Fragment56 | Ramos | C336(1.09) | LDD2205 | [25] |
| LDCM0569 | Fragment7 | Ramos | C336(0.70) | LDD2186 | [25] |
| LDCM0571 | Fragment9 | Ramos | C336(0.78) | LDD2188 | [25] |
| LDCM0022 | KB02 | HCT 116 | C336(1.58) | LDD0080 | [11] |
| LDCM0023 | KB03 | HCT 116 | C336(0.99) | LDD0081 | [11] |
| LDCM0024 | KB05 | HCT 116 | C336(1.09) | LDD0082 | [11] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C148(0.69) | LDD2121 | [6] |
| LDCM0109 | NEM | HeLa | H263(0.00); H292(0.00) | LDD0223 | [10] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C148(1.24) | LDD2093 | [6] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C148(0.50) | LDD2096 | [6] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C148(0.88); C336(0.87) | LDD2098 | [6] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C148(1.89) | LDD2099 | [6] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C148(0.98) | LDD2101 | [6] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C148(1.17) | LDD2107 | [6] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C148(0.68) | LDD2109 | [6] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C148(0.74) | LDD2110 | [6] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C148(1.42) | LDD2111 | [6] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C148(0.47) | LDD2115 | [6] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C148(0.75) | LDD2120 | [6] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C148(0.96) | LDD2123 | [6] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C148(0.93) | LDD2125 | [6] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C148(1.13) | LDD2127 | [6] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C148(0.90) | LDD2128 | [6] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C148(1.19) | LDD2129 | [6] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C148(0.72) | LDD2134 | [6] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C148(1.82) | LDD2135 | [6] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C148(1.44) | LDD2136 | [6] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C148(1.02) | LDD2137 | [6] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C148(3.25) | LDD1700 | [6] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C148(1.23) | LDD2140 | [6] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C148(0.93) | LDD2143 | [6] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C148(2.03) | LDD2144 | [6] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C148(1.20) | LDD2146 | [6] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C148(0.69) | LDD2150 | [6] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C336(1.24) | LDD2153 | [6] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C336(1.06) | LDD2206 | [26] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C336(0.83) | LDD2207 | [26] |
| LDCM0131 | RA190 | MM1.R | C336(1.06) | LDD0304 | [27] |
| LDCM0021 | THZ1 | HCT 116 | C336(1.22); C148(0.93) | LDD2173 | [11] |
| LDCM0111 | W14 | Hep-G2 | R107(0.52) | LDD0238 | [19] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Zinc finger protein 670 (ZNF670) | Krueppel C2H2-type zinc-finger protein family | Q9BS34 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| PRKCA-binding protein (PICK1) | . | Q9NRD5 | |||
The Drug(s) Related To This Target
Approved
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Fostamatinib | Small molecular drug | DB12010 | |||
Investigative
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Pmid22115617c2c | Small molecular drug | D0K6BI | |||
| [1-(6-{6-[(1-methylethyl)Amino]-1h-indazol-1-yl}Pyrazin-2-yl)-1h-pyrrol-3-yl]Acetic Acid | . | DB07546 | |||
Patented
References
