Details of the Target
General Information of Target
| Target ID | LDTP12420 | |||||
|---|---|---|---|---|---|---|
| Target Name | Xaa-Pro aminopeptidase 3 (XPNPEP3) | |||||
| Gene Name | XPNPEP3 | |||||
| Gene ID | 63929 | |||||
| Synonyms |
Xaa-Pro aminopeptidase 3; X-Pro aminopeptidase 3; EC 3.4.11.9; Aminopeptidase P3; APP3 |
|||||
| 3D Structure | ||||||
| Sequence |
MATSTSTEAKSASWWNYFFLYDGSKVKEEGDPTRAGICYFYPSQTLLDQQELLCGQIAGV
VRCVSDISDSPPTLVRLRKLKFAIKVDGDYLWVLGCAVELPDVSCKRFLDQLVGFFNFYN GPVSLAYENCSQEELSTEWDTFIEQILKNTSDLHKIFNSLWNLDQTKVEPLLLLKAARIL QTCQRSPHILAGCILYKGLIVSTQLPPSLTAKVLLHRTAPQEQRLPTGEDAPQEHGAALP PNVQIIPVFVTKEEAISLHEFPVEQMTRSLASPAGLQDGSAQHHPKGGSTSALKENATGH VESMAWTTPDPTSPDEACPDGRKENGCLSGHDLESIRPAGLHNSARGEVLGLSSSLGKEL VFLQEELDLSEIHIPEAQEVEMASGHFAFLHVPVPDGRAPYCKASLSASSSLEPTPPEDT AISSLRPPSAPEMLTQHGAQEQLEDHPGHSSQAPIPRADPLPRRTRRPLLLPRLDPGQRG NKLPTGEQGLDEDVDGVCESHAAPGLECSSGSANCQGAGPSADGISSRLTPAESCMGLVR MNLYTHCVKGLVLSLLAEEPLLGDSAAIEEVYHSSLASLNGLEVHLKETLPRDEAASTSS TYNFTHYDRIQSLLMANLPQVATPQDRRFLQAVSLMHSEFAQLPALYEMTVRNASTAVYA CCNPIQETYFQQLAPAARSSGFPNPQDGAFSLSGKAKQKLLKHGVNLL |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Peptidase M24B family
|
|||||
| Subcellular location |
Cytoplasm; Mitochondrion
|
|||||
| Function |
Catalyzes the removal of a penultimate prolyl residue from the N-termini of peptides, such as Leu-Pro-Ala. Also shows low activity towards peptides with Ala or Ser at the P1 position.; [Isoform 1]: Promotes TNFRSF1B-mediated phosphorylation of MAPK8/JNK1 and MAPK9/JNK2, suggesting a function as an adapter protein for TNFRSF1B; the effect is independent of XPNPEP3 peptidase activity. May inhibit apoptotic cell death induced via TNF-TNFRSF1B signaling.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| Ishikawa (Heraklio) 02 ER | SNV: p.H147Q | . | |||
| LN18 | SNV: p.I83V | . | |||
| MEWO | SNV: p.P112L | . | |||
| MOLT4 | SNV: p.A463V | IA-alkyne Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Alkylaryl probe 2 Probe Info |
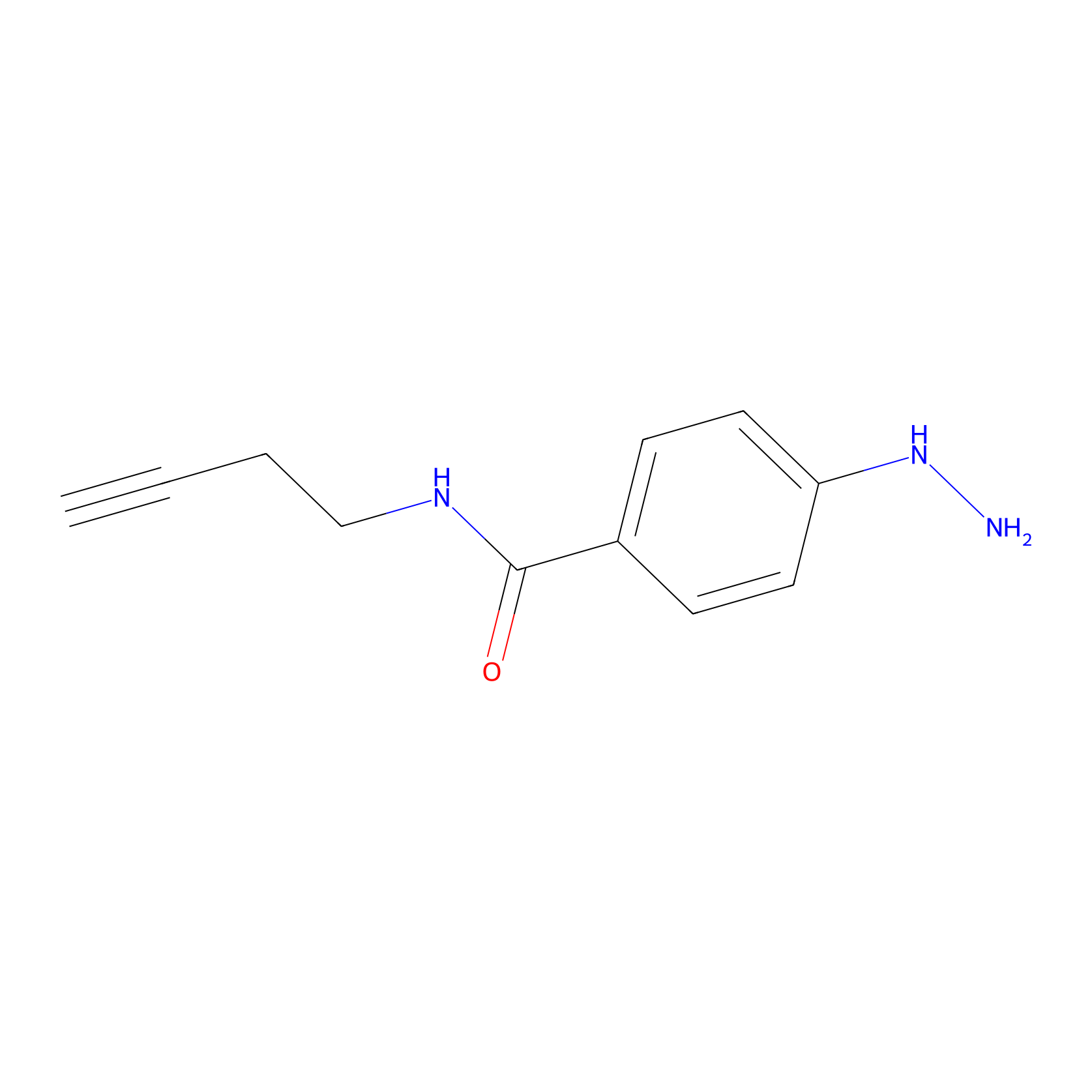 |
20.00 | LDD0389 | [1] | |
|
DBIA Probe Info |
 |
C334(1.02); C338(1.02) | LDD0593 | [2] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [3] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [3] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [4] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [5] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [5] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [5] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [6] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C055 Probe Info |
 |
17.03 | LDD1752 | [7] | |
|
C092 Probe Info |
 |
17.75 | LDD1783 | [7] | |
|
C107 Probe Info |
 |
10.63 | LDD1794 | [7] | |
|
C185 Probe Info |
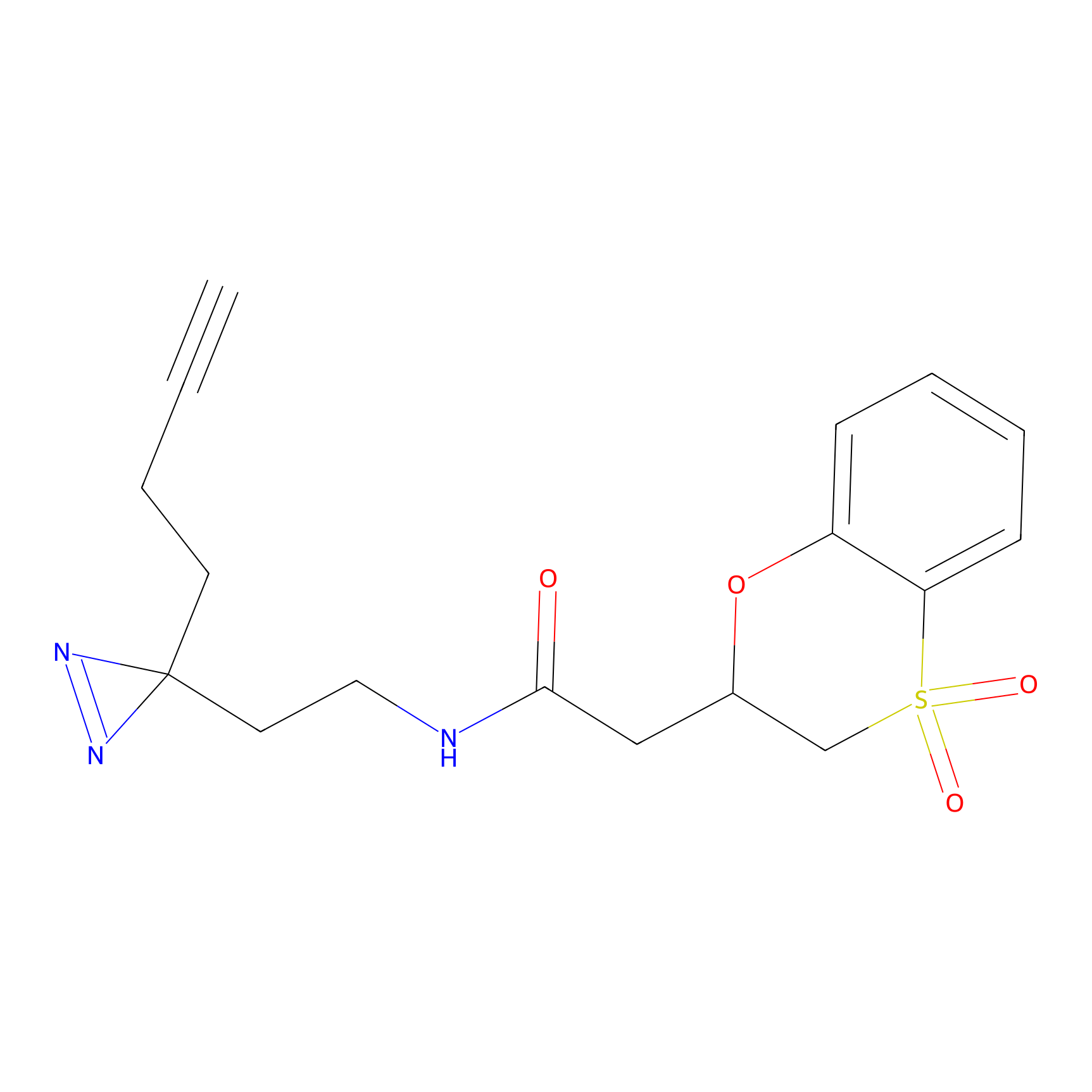 |
9.13 | LDD1863 | [7] | |
|
C350 Probe Info |
 |
36.25 | LDD2011 | [7] | |
|
C356 Probe Info |
 |
8.22 | LDD2017 | [7] | |
|
FFF probe2 Probe Info |
 |
10.05 | LDD0463 | [8] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0465 | [8] | |
|
FFF probe6 Probe Info |
 |
20.00 | LDD0468 | [8] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0136 | [9] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0139 | [9] | |
|
OEA-DA Probe Info |
 |
16.82 | LDD0046 | [10] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0216 | AC100 | HEK-293T | C338(1.15) | LDD0814 | [2] |
| LDCM0217 | AC101 | HEK-293T | C338(0.92) | LDD0815 | [2] |
| LDCM0218 | AC102 | HEK-293T | C338(1.00) | LDD0816 | [2] |
| LDCM0219 | AC103 | HEK-293T | C338(1.44) | LDD0817 | [2] |
| LDCM0220 | AC104 | HEK-293T | C338(1.34) | LDD0818 | [2] |
| LDCM0221 | AC105 | HEK-293T | C338(1.05) | LDD0819 | [2] |
| LDCM0222 | AC106 | HEK-293T | C338(1.48) | LDD0820 | [2] |
| LDCM0223 | AC107 | HEK-293T | C338(1.34) | LDD0821 | [2] |
| LDCM0224 | AC108 | HEK-293T | C338(1.76) | LDD0822 | [2] |
| LDCM0225 | AC109 | HEK-293T | C338(1.02) | LDD0823 | [2] |
| LDCM0227 | AC110 | HEK-293T | C338(1.09) | LDD0825 | [2] |
| LDCM0228 | AC111 | HEK-293T | C338(1.11) | LDD0826 | [2] |
| LDCM0229 | AC112 | HEK-293T | C338(1.24) | LDD0827 | [2] |
| LDCM0230 | AC113 | HEK-293T | C334(1.01) | LDD0828 | [2] |
| LDCM0231 | AC114 | HEK-293T | C334(0.94) | LDD0829 | [2] |
| LDCM0232 | AC115 | HEK-293T | C334(0.98) | LDD0830 | [2] |
| LDCM0233 | AC116 | HEK-293T | C334(0.87) | LDD0831 | [2] |
| LDCM0234 | AC117 | HEK-293T | C334(0.88) | LDD0832 | [2] |
| LDCM0235 | AC118 | HEK-293T | C334(0.88) | LDD0833 | [2] |
| LDCM0236 | AC119 | HEK-293T | C334(0.84) | LDD0834 | [2] |
| LDCM0238 | AC120 | HEK-293T | C334(0.77) | LDD0836 | [2] |
| LDCM0239 | AC121 | HEK-293T | C334(0.75) | LDD0837 | [2] |
| LDCM0240 | AC122 | HEK-293T | C334(0.70) | LDD0838 | [2] |
| LDCM0241 | AC123 | HEK-293T | C334(0.80) | LDD0839 | [2] |
| LDCM0242 | AC124 | HEK-293T | C334(0.74) | LDD0840 | [2] |
| LDCM0243 | AC125 | HEK-293T | C334(0.79) | LDD0841 | [2] |
| LDCM0244 | AC126 | HEK-293T | C334(0.77) | LDD0842 | [2] |
| LDCM0245 | AC127 | HEK-293T | C334(0.73) | LDD0843 | [2] |
| LDCM0246 | AC128 | HEK-293T | C334(1.13) | LDD0844 | [2] |
| LDCM0247 | AC129 | HEK-293T | C334(0.85) | LDD0845 | [2] |
| LDCM0249 | AC130 | HEK-293T | C334(1.02) | LDD0847 | [2] |
| LDCM0250 | AC131 | HEK-293T | C334(1.08) | LDD0848 | [2] |
| LDCM0251 | AC132 | HEK-293T | C334(1.03) | LDD0849 | [2] |
| LDCM0252 | AC133 | HEK-293T | C334(0.90) | LDD0850 | [2] |
| LDCM0253 | AC134 | HEK-293T | C334(0.96) | LDD0851 | [2] |
| LDCM0254 | AC135 | HEK-293T | C334(0.99) | LDD0852 | [2] |
| LDCM0255 | AC136 | HEK-293T | C334(0.86) | LDD0853 | [2] |
| LDCM0256 | AC137 | HEK-293T | C334(0.87) | LDD0854 | [2] |
| LDCM0257 | AC138 | HEK-293T | C334(0.96) | LDD0855 | [2] |
| LDCM0258 | AC139 | HEK-293T | C334(0.87) | LDD0856 | [2] |
| LDCM0260 | AC140 | HEK-293T | C334(1.00) | LDD0858 | [2] |
| LDCM0261 | AC141 | HEK-293T | C334(0.90) | LDD0859 | [2] |
| LDCM0262 | AC142 | HEK-293T | C334(0.92) | LDD0860 | [2] |
| LDCM0276 | AC17 | HCT 116 | C334(1.02); C338(1.02) | LDD0593 | [2] |
| LDCM0277 | AC18 | HCT 116 | C334(0.91); C338(0.91) | LDD0594 | [2] |
| LDCM0278 | AC19 | HCT 116 | C334(0.97); C338(0.97) | LDD0595 | [2] |
| LDCM0280 | AC20 | HCT 116 | C334(0.58); C338(0.58) | LDD0597 | [2] |
| LDCM0281 | AC21 | HCT 116 | C334(0.84); C338(0.84) | LDD0598 | [2] |
| LDCM0282 | AC22 | HCT 116 | C334(0.68); C338(0.68) | LDD0599 | [2] |
| LDCM0283 | AC23 | HCT 116 | C334(0.77); C338(0.77) | LDD0600 | [2] |
| LDCM0284 | AC24 | HCT 116 | C334(0.76); C338(0.76) | LDD0601 | [2] |
| LDCM0285 | AC25 | HEK-293T | C338(0.98) | LDD0883 | [2] |
| LDCM0286 | AC26 | HEK-293T | C338(0.87) | LDD0884 | [2] |
| LDCM0287 | AC27 | HEK-293T | C338(1.26) | LDD0885 | [2] |
| LDCM0288 | AC28 | HEK-293T | C338(2.24) | LDD0886 | [2] |
| LDCM0289 | AC29 | HEK-293T | C338(0.90) | LDD0887 | [2] |
| LDCM0291 | AC30 | HEK-293T | C338(0.95) | LDD0889 | [2] |
| LDCM0292 | AC31 | HEK-293T | C338(1.09) | LDD0890 | [2] |
| LDCM0293 | AC32 | HEK-293T | C338(1.26) | LDD0891 | [2] |
| LDCM0294 | AC33 | HEK-293T | C338(1.06) | LDD0892 | [2] |
| LDCM0295 | AC34 | HEK-293T | C338(0.92) | LDD0893 | [2] |
| LDCM0296 | AC35 | HCT 116 | C334(0.75); C338(0.75) | LDD0613 | [2] |
| LDCM0297 | AC36 | HCT 116 | C334(1.79); C338(1.79) | LDD0614 | [2] |
| LDCM0298 | AC37 | HCT 116 | C334(0.99); C338(0.99) | LDD0615 | [2] |
| LDCM0299 | AC38 | HCT 116 | C334(1.21); C338(1.21) | LDD0616 | [2] |
| LDCM0300 | AC39 | HCT 116 | C334(1.19); C338(1.19) | LDD0617 | [2] |
| LDCM0302 | AC40 | HCT 116 | C334(1.04); C338(1.04) | LDD0619 | [2] |
| LDCM0303 | AC41 | HCT 116 | C334(1.41); C338(1.41) | LDD0620 | [2] |
| LDCM0304 | AC42 | HCT 116 | C334(1.14); C338(1.14) | LDD0621 | [2] |
| LDCM0305 | AC43 | HCT 116 | C334(1.18); C338(1.18) | LDD0622 | [2] |
| LDCM0306 | AC44 | HCT 116 | C334(1.24); C338(1.24) | LDD0623 | [2] |
| LDCM0307 | AC45 | HCT 116 | C334(0.64); C338(0.64) | LDD0624 | [2] |
| LDCM0308 | AC46 | HCT 116 | C334(1.74); C338(1.74) | LDD0625 | [2] |
| LDCM0309 | AC47 | HCT 116 | C334(1.40); C338(1.40) | LDD0626 | [2] |
| LDCM0310 | AC48 | HCT 116 | C334(1.58); C338(1.58) | LDD0627 | [2] |
| LDCM0311 | AC49 | HCT 116 | C334(0.48); C338(0.48) | LDD0628 | [2] |
| LDCM0313 | AC50 | HCT 116 | C334(0.63); C338(0.63) | LDD0630 | [2] |
| LDCM0314 | AC51 | HCT 116 | C334(1.52); C338(1.52) | LDD0631 | [2] |
| LDCM0315 | AC52 | HCT 116 | C334(1.72); C338(1.72) | LDD0632 | [2] |
| LDCM0316 | AC53 | HCT 116 | C334(1.19); C338(1.19) | LDD0633 | [2] |
| LDCM0317 | AC54 | HCT 116 | C334(0.92); C338(0.92) | LDD0634 | [2] |
| LDCM0318 | AC55 | HCT 116 | C334(0.70); C338(0.70) | LDD0635 | [2] |
| LDCM0319 | AC56 | HCT 116 | C334(0.53); C338(0.53) | LDD0636 | [2] |
| LDCM0320 | AC57 | HEK-293T | C334(1.07); C338(1.14) | LDD0918 | [2] |
| LDCM0321 | AC58 | HEK-293T | C334(1.05); C338(1.18) | LDD0919 | [2] |
| LDCM0322 | AC59 | HEK-293T | C334(0.96); C338(1.20) | LDD0920 | [2] |
| LDCM0324 | AC60 | HEK-293T | C334(0.76); C338(1.47) | LDD0922 | [2] |
| LDCM0325 | AC61 | HEK-293T | C334(0.77); C338(1.39) | LDD0923 | [2] |
| LDCM0326 | AC62 | HEK-293T | C334(0.97); C338(1.30) | LDD0924 | [2] |
| LDCM0327 | AC63 | HEK-293T | C334(0.78); C338(1.23) | LDD0925 | [2] |
| LDCM0328 | AC64 | HEK-293T | C334(0.99); C338(1.47) | LDD0926 | [2] |
| LDCM0329 | AC65 | HEK-293T | C334(0.72); C338(1.35) | LDD0927 | [2] |
| LDCM0330 | AC66 | HEK-293T | C334(0.91); C338(1.43) | LDD0928 | [2] |
| LDCM0331 | AC67 | HEK-293T | C334(0.86); C338(1.31) | LDD0929 | [2] |
| LDCM0332 | AC68 | HCT 116 | C334(1.31); C338(1.31) | LDD0649 | [2] |
| LDCM0333 | AC69 | HCT 116 | C334(1.19); C338(1.19) | LDD0650 | [2] |
| LDCM0335 | AC70 | HCT 116 | C334(0.50); C338(0.50) | LDD0652 | [2] |
| LDCM0336 | AC71 | HCT 116 | C334(0.99); C338(0.99) | LDD0653 | [2] |
| LDCM0337 | AC72 | HCT 116 | C334(0.97); C338(0.97) | LDD0654 | [2] |
| LDCM0338 | AC73 | HCT 116 | C334(0.47); C338(0.47) | LDD0655 | [2] |
| LDCM0339 | AC74 | HCT 116 | C334(0.46); C338(0.46) | LDD0656 | [2] |
| LDCM0340 | AC75 | HCT 116 | C334(0.34); C338(0.34) | LDD0657 | [2] |
| LDCM0341 | AC76 | HCT 116 | C334(0.91); C338(0.91) | LDD0658 | [2] |
| LDCM0342 | AC77 | HCT 116 | C334(1.12); C338(1.12) | LDD0659 | [2] |
| LDCM0343 | AC78 | HCT 116 | C334(1.57); C338(1.57) | LDD0660 | [2] |
| LDCM0344 | AC79 | HCT 116 | C334(1.23); C338(1.23) | LDD0661 | [2] |
| LDCM0346 | AC80 | HCT 116 | C334(1.05); C338(1.05) | LDD0663 | [2] |
| LDCM0347 | AC81 | HCT 116 | C334(0.58); C338(0.58) | LDD0664 | [2] |
| LDCM0348 | AC82 | HCT 116 | C334(0.25); C338(0.25) | LDD0665 | [2] |
| LDCM0349 | AC83 | HEK-293T | C334(1.08) | LDD0947 | [2] |
| LDCM0350 | AC84 | HEK-293T | C334(0.85) | LDD0948 | [2] |
| LDCM0351 | AC85 | HEK-293T | C334(0.87) | LDD0949 | [2] |
| LDCM0352 | AC86 | HEK-293T | C334(0.91) | LDD0950 | [2] |
| LDCM0353 | AC87 | HEK-293T | C334(1.04) | LDD0951 | [2] |
| LDCM0354 | AC88 | HEK-293T | C334(0.90) | LDD0952 | [2] |
| LDCM0355 | AC89 | HEK-293T | C334(0.81) | LDD0953 | [2] |
| LDCM0357 | AC90 | HEK-293T | C334(0.71) | LDD0955 | [2] |
| LDCM0358 | AC91 | HEK-293T | C334(0.93) | LDD0956 | [2] |
| LDCM0359 | AC92 | HEK-293T | C334(0.87) | LDD0957 | [2] |
| LDCM0360 | AC93 | HEK-293T | C334(0.81) | LDD0958 | [2] |
| LDCM0361 | AC94 | HEK-293T | C334(0.83) | LDD0959 | [2] |
| LDCM0362 | AC95 | HEK-293T | C334(0.82) | LDD0960 | [2] |
| LDCM0363 | AC96 | HEK-293T | C334(0.76) | LDD0961 | [2] |
| LDCM0364 | AC97 | HEK-293T | C334(0.74) | LDD0962 | [2] |
| LDCM0365 | AC98 | HEK-293T | C338(1.47) | LDD0963 | [2] |
| LDCM0366 | AC99 | HEK-293T | C338(1.18) | LDD0964 | [2] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [5] |
| LDCM0374 | CL105 | HCT 116 | C334(0.76); C338(0.76) | LDD0691 | [2] |
| LDCM0375 | CL106 | HCT 116 | C334(0.92); C338(0.92) | LDD0692 | [2] |
| LDCM0376 | CL107 | HCT 116 | C334(0.58); C338(0.58) | LDD0693 | [2] |
| LDCM0377 | CL108 | HCT 116 | C334(0.42); C338(0.42) | LDD0694 | [2] |
| LDCM0378 | CL109 | HCT 116 | C334(0.70); C338(0.70) | LDD0695 | [2] |
| LDCM0380 | CL110 | HCT 116 | C334(0.85); C338(0.85) | LDD0697 | [2] |
| LDCM0381 | CL111 | HCT 116 | C334(0.77); C338(0.77) | LDD0698 | [2] |
| LDCM0382 | CL112 | HEK-293T | C338(1.09) | LDD0980 | [2] |
| LDCM0383 | CL113 | HEK-293T | C338(1.05) | LDD0981 | [2] |
| LDCM0384 | CL114 | HEK-293T | C338(1.01) | LDD0982 | [2] |
| LDCM0385 | CL115 | HEK-293T | C338(0.95) | LDD0983 | [2] |
| LDCM0386 | CL116 | HEK-293T | C338(1.09) | LDD0984 | [2] |
| LDCM0387 | CL117 | HCT 116 | C334(0.30); C338(0.30) | LDD0704 | [2] |
| LDCM0388 | CL118 | HCT 116 | C334(0.61); C338(0.61) | LDD0705 | [2] |
| LDCM0389 | CL119 | HCT 116 | C334(0.66); C338(0.66) | LDD0706 | [2] |
| LDCM0391 | CL120 | HCT 116 | C334(0.83); C338(0.83) | LDD0708 | [2] |
| LDCM0392 | CL121 | HCT 116 | C334(0.74); C338(0.74) | LDD0709 | [2] |
| LDCM0393 | CL122 | HCT 116 | C334(1.39); C338(1.39) | LDD0710 | [2] |
| LDCM0394 | CL123 | HCT 116 | C334(1.40); C338(1.40) | LDD0711 | [2] |
| LDCM0395 | CL124 | HCT 116 | C334(0.76); C338(0.76) | LDD0712 | [2] |
| LDCM0396 | CL125 | HEK-293T | C334(1.10); C338(1.05) | LDD0994 | [2] |
| LDCM0397 | CL126 | HEK-293T | C334(1.07); C338(1.25) | LDD0995 | [2] |
| LDCM0398 | CL127 | HEK-293T | C334(1.13); C338(1.25) | LDD0996 | [2] |
| LDCM0399 | CL128 | HEK-293T | C334(1.01); C338(1.02) | LDD0997 | [2] |
| LDCM0403 | CL16 | HCT 116 | C334(1.02); C338(1.02) | LDD0720 | [2] |
| LDCM0404 | CL17 | HCT 116 | C334(1.45); C338(1.45) | LDD0721 | [2] |
| LDCM0405 | CL18 | HCT 116 | C334(1.60); C338(1.60) | LDD0722 | [2] |
| LDCM0406 | CL19 | HCT 116 | C334(1.20); C338(1.20) | LDD0723 | [2] |
| LDCM0408 | CL20 | HCT 116 | C334(0.87); C338(0.87) | LDD0725 | [2] |
| LDCM0409 | CL21 | HCT 116 | C334(1.02); C338(1.02) | LDD0726 | [2] |
| LDCM0410 | CL22 | HCT 116 | C334(0.35); C338(0.35) | LDD0727 | [2] |
| LDCM0411 | CL23 | HCT 116 | C334(1.18); C338(1.18) | LDD0728 | [2] |
| LDCM0412 | CL24 | HCT 116 | C334(0.42); C338(0.42) | LDD0729 | [2] |
| LDCM0413 | CL25 | HCT 116 | C334(0.71); C338(0.71) | LDD0730 | [2] |
| LDCM0414 | CL26 | HCT 116 | C334(0.89); C338(0.89) | LDD0731 | [2] |
| LDCM0415 | CL27 | HCT 116 | C334(1.51); C338(1.51) | LDD0732 | [2] |
| LDCM0416 | CL28 | HCT 116 | C334(0.96); C338(0.96) | LDD0733 | [2] |
| LDCM0417 | CL29 | HCT 116 | C334(0.88); C338(0.88) | LDD0734 | [2] |
| LDCM0419 | CL30 | HCT 116 | C334(1.11); C338(1.11) | LDD0736 | [2] |
| LDCM0420 | CL31 | HCT 116 | C334(0.54); C338(0.54) | LDD0737 | [2] |
| LDCM0421 | CL32 | HCT 116 | C334(0.71); C338(0.71) | LDD0738 | [2] |
| LDCM0422 | CL33 | HCT 116 | C334(0.54); C338(0.54) | LDD0739 | [2] |
| LDCM0423 | CL34 | HCT 116 | C334(0.63); C338(0.63) | LDD0740 | [2] |
| LDCM0424 | CL35 | HCT 116 | C334(0.28); C338(0.28) | LDD0741 | [2] |
| LDCM0425 | CL36 | HCT 116 | C334(0.64); C338(0.64) | LDD0742 | [2] |
| LDCM0426 | CL37 | HCT 116 | C334(0.42); C338(0.42) | LDD0743 | [2] |
| LDCM0428 | CL39 | HCT 116 | C334(0.46); C338(0.46) | LDD0745 | [2] |
| LDCM0430 | CL40 | HCT 116 | C334(0.60); C338(0.60) | LDD0747 | [2] |
| LDCM0431 | CL41 | HCT 116 | C334(0.81); C338(0.81) | LDD0748 | [2] |
| LDCM0432 | CL42 | HCT 116 | C334(0.22); C338(0.22) | LDD0749 | [2] |
| LDCM0433 | CL43 | HCT 116 | C334(0.24); C338(0.24) | LDD0750 | [2] |
| LDCM0434 | CL44 | HCT 116 | C334(0.77); C338(0.77) | LDD0751 | [2] |
| LDCM0435 | CL45 | HCT 116 | C334(0.98); C338(0.98) | LDD0752 | [2] |
| LDCM0427 | Fragment51 | HCT 116 | C334(0.23); C338(0.23) | LDD0744 | [2] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [5] |
| LDCM0022 | KB02 | HEK-293T | C492(0.97) | LDD1492 | [11] |
| LDCM0023 | KB03 | HEK-293T | C492(0.96) | LDD1497 | [11] |
| LDCM0024 | KB05 | HEK-293T | C492(0.98) | LDD1502 | [11] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0224 | [5] |
| LDCM0099 | Phenelzine | HEK-293T | 4.00 | LDD0390 | [1] |
References
