Details of the Target
General Information of Target
| Target ID | LDTP04434 | |||||
|---|---|---|---|---|---|---|
| Target Name | Host cell factor 1 (HCFC1) | |||||
| Gene Name | HCFC1 | |||||
| Gene ID | 3054 | |||||
| Synonyms |
HCF1; HFC1; Host cell factor 1; HCF; HCF-1; C1 factor; CFF; VCAF; VP16 accessory protein) [Cleaved into: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N-terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C-terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]
|
|||||
| 3D Structure | ||||||
| Sequence |
MASAVSPANLPAVLLQPRWKRVVGWSGPVPRPRHGHRAVAIKELIVVFGGGNEGIVDELH
VYNTATNQWFIPAVRGDIPPGCAAYGFVCDGTRLLVFGGMVEYGKYSNDLYELQASRWEW KRLKAKTPKNGPPPCPRLGHSFSLVGNKCYLFGGLANDSEDPKNNIPRYLNDLYILELRP GSGVVAWDIPITYGVLPPPRESHTAVVYTEKDNKKSKLVIYGGMSGCRLGDLWTLDIDTL TWNKPSLSGVAPLPRSLHSATTIGNKMYVFGGWVPLVMDDVKVATHEKEWKCTNTLACLN LDTMAWETILMDTLEDNIPRARAGHCAVAINTRLYIWSGRDGYRKAWNNQVCCKDLWYLE TEKPPPPARVQLVRANTNSLEVSWGAVATADSYLLQLQKYDIPATAATATSPTPNPVPSV PANPPKSPAPAAAAPAVQPLTQVGITLLPQAAPAPPTTTTIQVLPTVPGSSISVPTAART QGVPAVLKVTGPQATTGTPLVTMRPASQAGKAPVTVTSLPAGVRMVVPTQSAQGTVIGSS PQMSGMAALAAAAAATQKIPPSSAPTVLSVPAGTTIVKTMAVTPGTTTLPATVKVASSPV MVSNPATRMLKTAAAQVGTSVSSATNTSTRPIITVHKSGTVTVAQQAQVVTTVVGGVTKT ITLVKSPISVPGGSALISNLGKVMSVVQTKPVQTSAVTGQASTGPVTQIIQTKGPLPAGT ILKLVTSADGKPTTIITTTQASGAGTKPTILGISSVSPSTTKPGTTTIIKTIPMSAIITQ AGATGVTSSPGIKSPITIITTKVMTSGTGAPAKIITAVPKIATGHGQQGVTQVVLKGAPG QPGTILRTVPMGGVRLVTPVTVSAVKPAVTTLVVKGTTGVTTLGTVTGTVSTSLAGAGGH STSASLATPITTLGTIATLSSQVINPTAITVSAAQTTLTAAGGLTTPTITMQPVSQPTQV TLITAPSGVEAQPVHDLPVSILASPTTEQPTATVTIADSGQGDVQPGTVTLVCSNPPCET HETGTTNTATTTVVANLGGHPQPTQVQFVCDRQEAAASLVTSTVGQQNGSVVRVCSNPPC ETHETGTTNTATTATSNMAGQHGCSNPPCETHETGTTNTATTAMSSVGANHQRDARRACA AGTPAVIRISVATGALEAAQGSKSQCQTRQTSATSTTMTVMATGAPCSAGPLLGPSMARE PGGRSPAFVQLAPLSSKVRLSSPSIKDLPAGRHSHAVSTAAMTRSSVGAGEPRMAPVCES LQGGSPSTTVTVTALEALLCPSATVTQVCSNPPCETHETGTTNTATTSNAGSAQRVCSNP PCETHETGTTHTATTATSNGGTGQPEGGQQPPAGRPCETHQTTSTGTTMSVSVGALLPDA TSSHRTVESGLEVAAAPSVTPQAGTALLAPFPTQRVCSNPPCETHETGTTHTATTVTSNM SSNQDPPPAASDQGEVESTQGDSVNITSSSAITTTVSSTLTRAVTTVTQSTPVPGPSVPP PEELQVSPGPRQQLPPRQLLQSASTALMGESAEVLSASQTPELPAAVDLSSTGEPSSGQE SAGSAVVATVVVQPPPPTQSEVDQLSLPQELMAEAQAGTTTLMVTGLTPEELAVTAAAEA AAQAAATEEAQALAIQAVLQAAQQAVMGTGEPMDTSEAAATVTQAELGHLSAEGQEGQAT TIPIVLTQQELAALVQQQQLQEAQAQQQHHHLPTEALAPADSLNDPAIESNCLNELAGTV PSTVALLPSTATESLAPSNTFVAPQPVVVASPAKLQAAATLTEVANGIESLGVKPDLPPP PSKAPMKKENQWFDVGVIKGTNVMVTHYFLPPDDAVPSDDDLGTVPDYNQLKKQELQPGT AYKFRVAGINACGRGPFSEISAFKTCLPGFPGAPCAIKISKSPDGAHLTWEPPSVTSGKI IEYSVYLAIQSSQAGGELKSSTPAQLAFMRVYCGPSPSCLVQSSSLSNAHIDYTTKPAII FRIAARNEKGYGPATQVRWLQETSKDSSGTKPANKRPMSSPEMKSAPKKSKADGQ |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Transcriptional coregulator. Involved in control of the cell cycle. Also antagonizes transactivation by ZBTB17 and GABP2; represses ZBTB17 activation of the p15(INK4b) promoter and inhibits its ability to recruit p300. Coactivator for EGR2 and GABP2. Tethers the chromatin modifying Set1/Ash2 histone H3 'Lys-4' methyltransferase (H3K4me) and Sin3 histone deacetylase (HDAC) complexes (involved in the activation and repression of transcription, respectively) together. Component of a THAP1/THAP3-HCFC1-OGT complex that is required for the regulation of the transcriptional activity of RRM1. As part of the NSL complex it may be involved in acetylation of nucleosomal histone H4 on several lysine residues. Recruits KMT2E/MLL5 to E2F1 responsive promoters promoting transcriptional activation and thereby facilitates G1 to S phase transition. Modulates expression of homeobox protein PDX1, perhaps acting in concert with transcription factor E2F1, thereby regulating pancreatic beta-cell growth and glucose-stimulated insulin secretion. May negatively modulate transcriptional activity of FOXO3.; (Microbial infection) In case of human herpes simplex virus (HSV) infection, HCFC1 forms a multiprotein-DNA complex with the viral transactivator protein VP16 and POU2F1 thereby enabling the transcription of the viral immediate early genes.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| C32 | SNV: p.Q1518K | DBIA Probe Info | |||
| COLO792 | SNV: p.P254S Substitution: p.P1017F |
DBIA Probe Info | |||
| CORL23 | SNV: p.G538V | DBIA Probe Info | |||
| EFO27 | SNV: p.R333C | DBIA Probe Info | |||
| HCT15 | SNV: p.R1986C | DBIA Probe Info | |||
| HEC1 | SNV: p.L1700P | DBIA Probe Info | |||
| HT | SNV: p.T738A | DBIA Probe Info | |||
| HT115 | SNV: p.G1787D | DBIA Probe Info | |||
| IGR1 | SNV: p.S1958F | DBIA Probe Info | |||
| JURKAT | SNV: p.T784M; p.Q1053E; p.G1300S | Compound 10 Probe Info | |||
| KNS42 | SNV: p.G1038R; p.D1583H | DBIA Probe Info | |||
| LNCaP clone FGC | SNV: p.P1798L | DBIA Probe Info | |||
| MDAMB436 | SNV: p.P1766Q | DBIA Probe Info | |||
| MELHO | SNV: p.M1528I | DBIA Probe Info | |||
| MEWO | SNV: p.P1500L | DBIA Probe Info | |||
| MFE319 | SNV: p.E1789G; p.V1915M | DBIA Probe Info | |||
| MOLT4 | SNV: p.P435R; p.P439S | IA-alkyne Probe Info | |||
| NCIH1048 | SNV: p.T767A | DBIA Probe Info | |||
| NCIH1155 | SNV: p.F1829L | DBIA Probe Info | |||
| NCIH1666 | SNV: p.G1606W | DBIA Probe Info | |||
| NUGC3 | SNV: p.P985L | DBIA Probe Info | |||
| OVCAR8 | SNV: p.C149Y | DBIA Probe Info | |||
| PF382 | SNV: p.I1980V | DBIA Probe Info | |||
| SUDHL6 | Deletion: p.I1980del | DBIA Probe Info | |||
| T24 | SNV: p.A1353S | DBIA Probe Info | |||
| TOV21G | SNV: p.P985Q | DBIA Probe Info | |||
| WM2664 | Insertion: p.N378KfsTer24 | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
AZ-9 Probe Info |
 |
5.76 | LDD0393 | [2] | |
|
TH211 Probe Info |
 |
Y268(6.76) | LDD0260 | [3] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [4] | |
|
STPyne Probe Info |
 |
K1853(6.67); K1863(7.70); K1884(5.88); K1989(3.70) | LDD0277 | [5] | |
|
ONAyne Probe Info |
 |
N.A. | LDD0273 | [5] | |
|
Probe 1 Probe Info |
 |
Y106(21.58); Y111(20.03); Y208(13.53); Y1862(14.25) | LDD3495 | [6] | |
|
P11 Probe Info |
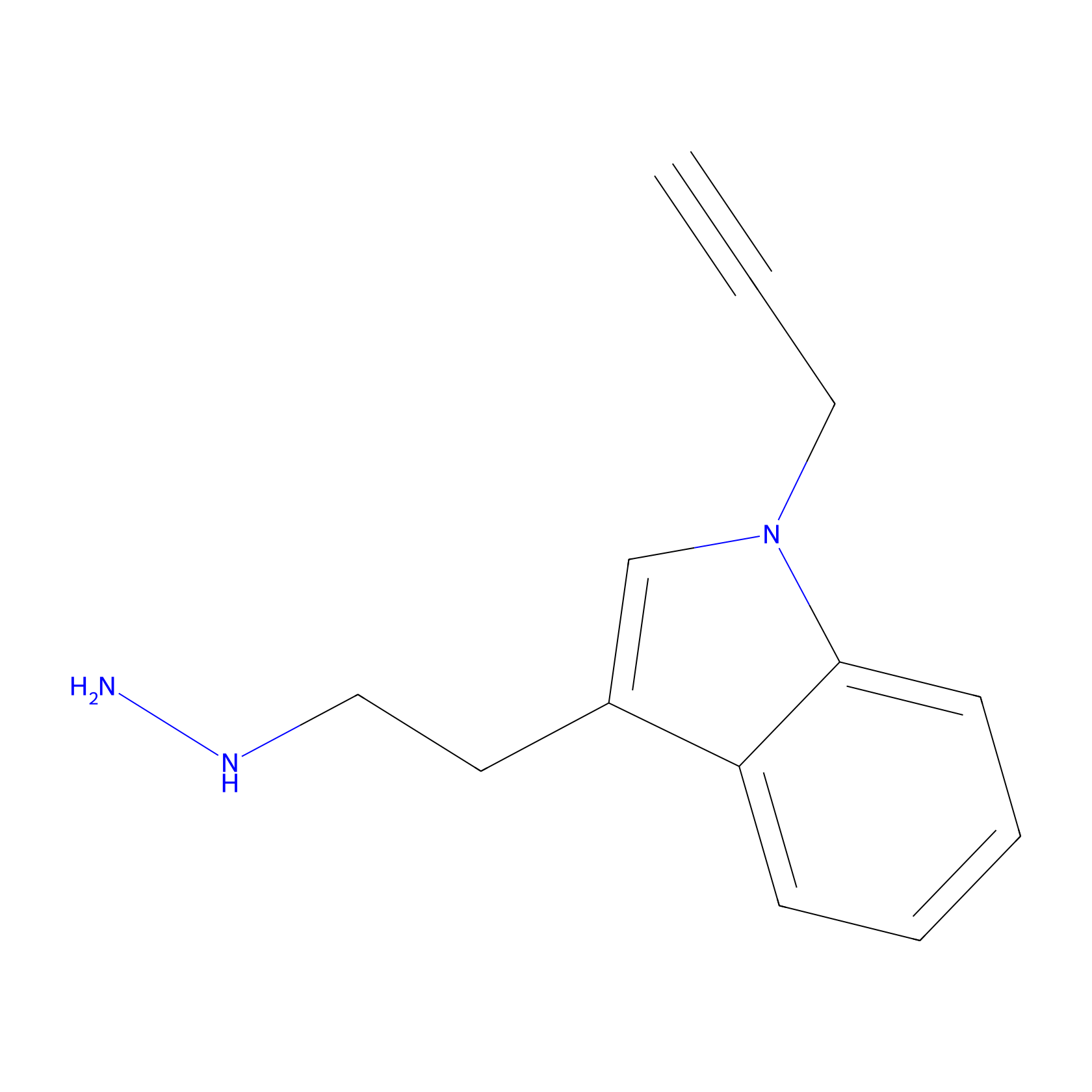 |
5.49 | LDD0201 | [7] | |
|
BTD Probe Info |
 |
C326(2.29); C1166(1.17) | LDD1700 | [8] | |
|
Sulforaphane-probe2 Probe Info |
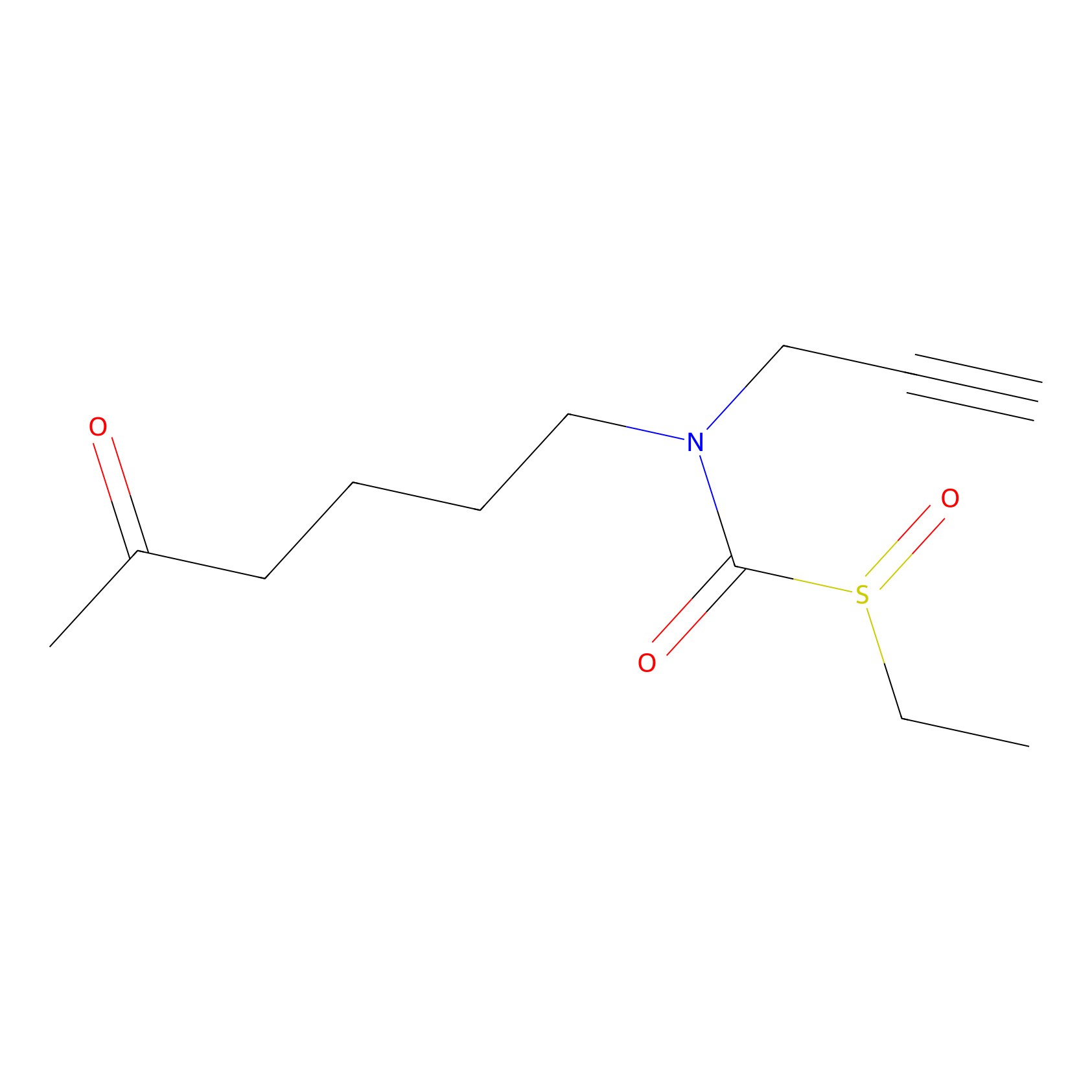 |
1.98 | LDD0160 | [9] | |
|
AHL-Pu-1 Probe Info |
 |
C227(2.56); C1166(3.41) | LDD0169 | [10] | |
|
DBIA Probe Info |
 |
C1166(8.03); C1139(20.67) | LDD0209 | [11] | |
|
HHS-465 Probe Info |
 |
Y103(10.00) | LDD2237 | [12] | |
|
5E-2FA Probe Info |
 |
H825(0.00); H1907(0.00); H1970(0.00) | LDD2235 | [13] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [14] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C1139(0.00); C326(0.00); C149(0.00); C227(0.00) | LDD0038 | [15] | |
|
IA-alkyne Probe Info |
 |
C227(0.00); C149(0.00) | LDD0032 | [16] | |
|
IPIAA_L Probe Info |
 |
C1953(0.00); C149(0.00) | LDD0031 | [17] | |
|
Lodoacetamide azide Probe Info |
 |
C1139(0.00); C326(0.00); C149(0.00); C227(0.00) | LDD0037 | [15] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [18] | |
|
NAIA_4 Probe Info |
 |
C227(0.00); C1050(0.00); C1139(0.00); C1872(0.00) | LDD2226 | [19] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [18] | |
|
WYneN Probe Info |
 |
C1166(0.00); C1139(0.00) | LDD0021 | [20] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [20] | |
|
Compound 10 Probe Info |
 |
C1139(0.00); C1166(0.00); C1872(0.00); C1886(0.00) | LDD2216 | [21] | |
|
Compound 11 Probe Info |
 |
C1139(0.00); C1872(0.00); C1886(0.00); C227(0.00) | LDD2213 | [21] | |
|
ENE Probe Info |
 |
C1872(0.00); C1886(0.00); C353(0.00) | LDD0006 | [20] | |
|
IPM Probe Info |
 |
C1139(0.00); C1166(0.00); C1886(0.00); C227(0.00) | LDD0005 | [20] | |
|
NHS Probe Info |
 |
K1989(0.00); K665(0.00) | LDD0010 | [20] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0017 | [22] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [23] | |
|
VSF Probe Info |
 |
C135(0.00); C1886(0.00) | LDD0007 | [20] | |
|
Phosphinate-6 Probe Info |
 |
C1872(0.00); C1166(0.00); C1895(0.00); C1886(0.00) | LDD0018 | [24] | |
|
1c-yne Probe Info |
 |
K1989(0.00); K1901(0.00); K820(0.00); K665(0.00) | LDD0228 | [25] | |
|
Acrolein Probe Info |
 |
H258(0.00); C1872(0.00); C1895(0.00); C227(0.00) | LDD0217 | [26] | |
|
Crotonaldehyde Probe Info |
 |
C1872(0.00); C1895(0.00); C227(0.00) | LDD0219 | [26] | |
|
Methacrolein Probe Info |
 |
C326(0.00); C1872(0.00); C227(0.00); C1895(0.00) | LDD0218 | [26] | |
|
W1 Probe Info |
 |
C1886(0.00); C1895(0.00); C352(0.00); C353(0.00) | LDD0236 | [27] | |
|
AOyne Probe Info |
 |
10.70 | LDD0443 | [28] | |
|
NAIA_5 Probe Info |
 |
C1139(0.00); C326(0.00); C149(0.00); C1872(0.00) | LDD2223 | [19] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-1 Probe Info |
 |
N.A. | LDD0137 | [29] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [30] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C326(0.69) | LDD2112 | [8] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C326(0.45) | LDD2095 | [8] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C1872(1.17) | LDD2130 | [8] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C326(0.86); C135(1.54); C1872(0.55) | LDD2117 | [8] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C326(1.19); C135(3.20); C1872(1.07) | LDD2152 | [8] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C135(1.22) | LDD2103 | [8] |
| LDCM0025 | 4SU-RNA | DM93 | C1895(3.24) | LDD0170 | [10] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C227(2.56); C1166(3.41) | LDD0169 | [10] |
| LDCM0214 | AC1 | HEK-293T | C1872(0.95); C1139(1.00); C227(0.98); C1187(1.08) | LDD1507 | [31] |
| LDCM0215 | AC10 | HEK-293T | C135(1.22); C326(0.98); C1872(1.00); C1139(1.05) | LDD1508 | [31] |
| LDCM0226 | AC11 | HEK-293T | C135(1.11); C1872(0.96); C1139(1.01); C227(0.89) | LDD1509 | [31] |
| LDCM0237 | AC12 | HEK-293T | C135(1.10); C326(0.96); C1872(0.90); C1139(1.02) | LDD1510 | [31] |
| LDCM0259 | AC14 | HEK-293T | C135(0.98); C326(0.97); C1872(0.80); C1139(1.05) | LDD1512 | [31] |
| LDCM0270 | AC15 | HEK-293T | C135(1.26); C1872(0.92); C1139(0.97); C227(0.93) | LDD1513 | [31] |
| LDCM0276 | AC17 | HEK-293T | C1872(1.00); C1139(1.05); C227(1.02); C1187(1.03) | LDD1515 | [31] |
| LDCM0277 | AC18 | HEK-293T | C135(1.14); C326(1.17); C1872(0.97); C1139(0.98) | LDD1516 | [31] |
| LDCM0278 | AC19 | HEK-293T | C135(1.30); C1872(1.49); C1139(1.68); C227(0.85) | LDD1517 | [31] |
| LDCM0279 | AC2 | HEK-293T | C135(1.12); C326(1.06); C1872(1.04); C1139(1.00) | LDD1518 | [31] |
| LDCM0280 | AC20 | HEK-293T | C135(1.06); C326(1.21); C1872(0.97); C1139(1.05) | LDD1519 | [31] |
| LDCM0281 | AC21 | HEK-293T | C135(1.01); C326(1.01); C1872(1.00); C1139(1.01) | LDD1520 | [31] |
| LDCM0282 | AC22 | HEK-293T | C135(0.96); C326(1.00); C1872(0.86); C1139(1.00) | LDD1521 | [31] |
| LDCM0283 | AC23 | HEK-293T | C135(1.15); C1872(0.93); C1139(0.95); C227(0.96) | LDD1522 | [31] |
| LDCM0284 | AC24 | HEK-293T | C135(1.15); C326(1.27); C1872(1.04); C1139(1.00) | LDD1523 | [31] |
| LDCM0285 | AC25 | HEK-293T | C1872(1.01); C1139(1.05); C227(1.01); C1187(1.01) | LDD1524 | [31] |
| LDCM0286 | AC26 | HEK-293T | C135(1.18); C326(1.03); C1872(1.07); C1139(1.03) | LDD1525 | [31] |
| LDCM0287 | AC27 | HEK-293T | C135(1.04); C1872(1.06); C1139(1.03); C227(1.02) | LDD1526 | [31] |
| LDCM0288 | AC28 | HEK-293T | C135(1.19); C326(1.02); C1872(0.99); C1139(1.03) | LDD1527 | [31] |
| LDCM0289 | AC29 | HEK-293T | C135(1.09); C326(0.97); C1872(0.87); C1139(1.03) | LDD1528 | [31] |
| LDCM0290 | AC3 | HEK-293T | C135(1.10); C1872(0.95); C1139(1.03); C227(0.90) | LDD1529 | [31] |
| LDCM0291 | AC30 | HEK-293T | C135(1.08); C326(0.99); C1872(1.01); C1139(1.06) | LDD1530 | [31] |
| LDCM0292 | AC31 | HEK-293T | C135(1.18); C1872(0.95); C1139(0.96); C227(0.95) | LDD1531 | [31] |
| LDCM0293 | AC32 | HEK-293T | C135(1.09); C326(1.53); C1872(1.02); C1139(1.03) | LDD1532 | [31] |
| LDCM0294 | AC33 | HEK-293T | C1872(1.02); C1139(1.02); C227(1.00); C1187(1.11) | LDD1533 | [31] |
| LDCM0295 | AC34 | HEK-293T | C135(1.15); C326(1.09); C1872(1.07); C1139(1.04) | LDD1534 | [31] |
| LDCM0296 | AC35 | HEK-293T | C135(1.20); C1872(1.02); C1139(0.99); C227(0.97) | LDD1535 | [31] |
| LDCM0297 | AC36 | HEK-293T | C135(1.12); C326(1.09); C1872(0.93); C1139(1.00) | LDD1536 | [31] |
| LDCM0298 | AC37 | HEK-293T | C135(0.98); C326(1.07); C1872(0.82); C1139(1.01) | LDD1537 | [31] |
| LDCM0299 | AC38 | HEK-293T | C135(1.02); C326(0.99); C1872(0.90); C1139(0.99) | LDD1538 | [31] |
| LDCM0300 | AC39 | HEK-293T | C135(1.12); C1872(0.99); C1139(1.00); C227(0.93) | LDD1539 | [31] |
| LDCM0301 | AC4 | HEK-293T | C135(1.15); C326(1.16); C1872(0.87); C1139(1.03) | LDD1540 | [31] |
| LDCM0302 | AC40 | HEK-293T | C135(1.09); C326(1.48); C1872(1.07); C1139(1.03) | LDD1541 | [31] |
| LDCM0303 | AC41 | HEK-293T | C1872(0.97); C1139(1.04); C227(1.01); C1187(1.11) | LDD1542 | [31] |
| LDCM0304 | AC42 | HEK-293T | C135(1.27); C326(1.02); C1872(1.04); C1139(1.11) | LDD1543 | [31] |
| LDCM0305 | AC43 | HEK-293T | C135(1.22); C1872(0.93); C1139(0.96); C227(0.95) | LDD1544 | [31] |
| LDCM0306 | AC44 | HEK-293T | C135(1.24); C326(1.16); C1872(0.98); C1139(1.01) | LDD1545 | [31] |
| LDCM0307 | AC45 | HEK-293T | C135(0.96); C326(1.06); C1872(1.06); C1139(1.08) | LDD1546 | [31] |
| LDCM0308 | AC46 | HEK-293T | C135(1.00); C326(0.92); C1872(0.96); C1139(1.05) | LDD1547 | [31] |
| LDCM0309 | AC47 | HEK-293T | C135(1.22); C1872(0.93); C1139(0.96); C227(0.95) | LDD1548 | [31] |
| LDCM0310 | AC48 | HEK-293T | C135(1.15); C326(1.20); C1872(1.00); C1139(1.08) | LDD1549 | [31] |
| LDCM0311 | AC49 | HEK-293T | C1872(1.03); C1139(1.02); C227(1.02); C1187(1.14) | LDD1550 | [31] |
| LDCM0312 | AC5 | HEK-293T | C135(0.99); C326(1.07); C1872(0.75); C1139(0.80) | LDD1551 | [31] |
| LDCM0313 | AC50 | HEK-293T | C135(1.28); C326(1.11); C1872(1.06); C1139(1.00) | LDD1552 | [31] |
| LDCM0314 | AC51 | HEK-293T | C135(1.11); C1872(1.00); C1139(0.97); C227(0.92) | LDD1553 | [31] |
| LDCM0315 | AC52 | HEK-293T | C135(1.29); C326(1.21); C1872(0.86); C1139(1.03) | LDD1554 | [31] |
| LDCM0316 | AC53 | HEK-293T | C135(1.01); C326(1.08); C1872(1.12); C1139(0.97) | LDD1555 | [31] |
| LDCM0317 | AC54 | HEK-293T | C135(1.03); C326(1.00); C1872(0.98); C1139(1.02) | LDD1556 | [31] |
| LDCM0318 | AC55 | HEK-293T | C135(1.17); C1872(0.91); C1139(1.00); C227(0.92) | LDD1557 | [31] |
| LDCM0319 | AC56 | HEK-293T | C135(1.16); C326(1.03); C1872(0.96); C1139(1.02) | LDD1558 | [31] |
| LDCM0320 | AC57 | HEK-293T | C1872(0.98); C1139(1.04); C227(1.06); C1187(1.16) | LDD1559 | [31] |
| LDCM0321 | AC58 | HEK-293T | C135(1.17); C326(1.00); C1872(0.88); C1139(1.02) | LDD1560 | [31] |
| LDCM0322 | AC59 | HEK-293T | C135(1.12); C1872(1.00); C1139(0.98); C227(0.92) | LDD1561 | [31] |
| LDCM0323 | AC6 | HEK-293T | C135(1.01); C326(0.96); C1872(0.90); C1139(1.04) | LDD1562 | [31] |
| LDCM0324 | AC60 | HEK-293T | C135(1.07); C326(0.97); C1872(0.86); C1139(1.01) | LDD1563 | [31] |
| LDCM0325 | AC61 | HEK-293T | C135(1.00); C326(1.07); C1872(0.92); C1139(0.97) | LDD1564 | [31] |
| LDCM0326 | AC62 | HEK-293T | C135(1.03); C326(0.97); C1872(0.86); C1139(1.06) | LDD1565 | [31] |
| LDCM0327 | AC63 | HEK-293T | C135(1.17); C1872(0.97); C1139(1.00); C227(0.88) | LDD1566 | [31] |
| LDCM0328 | AC64 | HEK-293T | C135(1.22); C326(1.00); C1872(1.13); C1139(1.03) | LDD1567 | [31] |
| LDCM0334 | AC7 | HEK-293T | C135(1.07); C1872(0.95); C1139(0.94); C227(0.90) | LDD1568 | [31] |
| LDCM0345 | AC8 | HEK-293T | C135(1.02); C326(1.20); C1872(1.16); C1139(1.07) | LDD1569 | [31] |
| LDCM0545 | Acetamide | MDA-MB-231 | C1872(0.32) | LDD2138 | [8] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C326(0.89); C135(1.81); C353(0.94) | LDD2113 | [8] |
| LDCM0248 | AKOS034007472 | HEK-293T | C135(0.99); C326(0.96); C1872(0.78); C1139(1.16) | LDD1511 | [31] |
| LDCM0356 | AKOS034007680 | HEK-293T | C1872(0.99); C1139(1.03); C227(1.01); C1187(1.08) | LDD1570 | [31] |
| LDCM0275 | AKOS034007705 | HEK-293T | C135(1.21); C326(1.05); C1872(0.99); C1139(0.99) | LDD1514 | [31] |
| LDCM0156 | Aniline | NCI-H1299 | C1139(0.00); C1872(0.00); C227(0.00); C1886(0.00) | LDD0404 | [1] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C135(0.40); C1872(0.50) | LDD2091 | [8] |
| LDCM0088 | C45 | HEK-293T | 5.49 | LDD0201 | [7] |
| LDCM0630 | CCW28-3 | 231MFP | C1886(1.22); C227(1.19) | LDD2214 | [32] |
| LDCM0108 | Chloroacetamide | HeLa | C227(0.00); C1872(0.00); H258(0.00); C352(0.00) | LDD0222 | [26] |
| LDCM0632 | CL-Sc | Hep-G2 | C326(2.12); C1139(1.16); C227(0.57) | LDD2227 | [19] |
| LDCM0367 | CL1 | HEK-293T | C135(1.08); C326(1.03); C1872(0.91); C1139(0.95) | LDD1571 | [31] |
| LDCM0368 | CL10 | HEK-293T | C135(0.97); C326(0.76); C1872(1.01); C1139(1.13) | LDD1572 | [31] |
| LDCM0369 | CL100 | HEK-293T | C326(0.99); C1872(0.96); C1139(1.01); C227(0.94) | LDD1573 | [31] |
| LDCM0370 | CL101 | HEK-293T | C135(1.01); C326(0.96); C1872(0.96); C1139(1.04) | LDD1574 | [31] |
| LDCM0371 | CL102 | HEK-293T | C135(1.13); C1872(0.99); C1139(1.03); C227(0.89) | LDD1575 | [31] |
| LDCM0372 | CL103 | HEK-293T | C135(1.06); C326(1.05); C1872(1.06); C1139(1.03) | LDD1576 | [31] |
| LDCM0373 | CL104 | HEK-293T | C326(1.07); C1872(0.89); C1139(1.01); C227(0.98) | LDD1577 | [31] |
| LDCM0374 | CL105 | HEK-293T | C135(0.96); C326(0.97); C1872(1.02); C1139(0.98) | LDD1578 | [31] |
| LDCM0375 | CL106 | HEK-293T | C135(1.09); C1872(1.00); C1139(0.96); C227(0.94) | LDD1579 | [31] |
| LDCM0376 | CL107 | HEK-293T | C135(1.13); C326(1.04); C1872(1.05); C1139(0.97) | LDD1580 | [31] |
| LDCM0377 | CL108 | HEK-293T | C326(0.99); C1872(1.06); C1139(0.99); C227(1.01) | LDD1581 | [31] |
| LDCM0378 | CL109 | HEK-293T | C135(1.07); C326(0.85); C1872(1.03); C1139(0.93) | LDD1582 | [31] |
| LDCM0379 | CL11 | HEK-293T | C135(0.90); C1872(0.77); C1139(1.18); C227(0.87) | LDD1583 | [31] |
| LDCM0380 | CL110 | HEK-293T | C135(0.99); C1872(1.02); C1139(1.00); C227(0.92) | LDD1584 | [31] |
| LDCM0381 | CL111 | HEK-293T | C135(0.97); C326(1.06); C1872(1.04); C1139(0.98) | LDD1585 | [31] |
| LDCM0382 | CL112 | HEK-293T | C326(0.90); C1872(1.01); C1139(1.04); C227(0.98) | LDD1586 | [31] |
| LDCM0383 | CL113 | HEK-293T | C135(1.11); C326(0.97); C1872(0.95); C1139(1.06) | LDD1587 | [31] |
| LDCM0384 | CL114 | HEK-293T | C135(1.06); C1872(0.98); C1139(0.99); C227(0.84) | LDD1588 | [31] |
| LDCM0385 | CL115 | HEK-293T | C135(1.12); C326(1.09); C1872(1.07); C1139(0.98) | LDD1589 | [31] |
| LDCM0386 | CL116 | HEK-293T | C326(0.99); C1872(1.01); C1139(0.98); C227(0.98) | LDD1590 | [31] |
| LDCM0387 | CL117 | HEK-293T | C135(1.15); C326(1.04); C1872(1.02); C1139(1.01) | LDD1591 | [31] |
| LDCM0388 | CL118 | HEK-293T | C135(1.15); C1872(1.10); C1139(1.12); C227(0.92) | LDD1592 | [31] |
| LDCM0389 | CL119 | HEK-293T | C135(1.20); C326(1.03); C1872(1.03); C1139(0.97) | LDD1593 | [31] |
| LDCM0390 | CL12 | HEK-293T | C135(0.97); C326(0.84); C1872(0.90); C1139(1.22) | LDD1594 | [31] |
| LDCM0391 | CL120 | HEK-293T | C326(0.89); C1872(0.95); C1139(1.07); C227(0.97) | LDD1595 | [31] |
| LDCM0392 | CL121 | HEK-293T | C135(1.13); C326(0.89); C1872(0.99); C1139(1.03) | LDD1596 | [31] |
| LDCM0393 | CL122 | HEK-293T | C135(1.14); C1872(0.96); C1139(1.00); C227(0.87) | LDD1597 | [31] |
| LDCM0394 | CL123 | HEK-293T | C135(0.97); C326(0.63); C1872(1.02); C1139(0.93) | LDD1598 | [31] |
| LDCM0395 | CL124 | HEK-293T | C326(0.96); C1872(0.99); C1139(1.00); C227(0.92) | LDD1599 | [31] |
| LDCM0396 | CL125 | HEK-293T | C135(1.09); C326(1.02); C1872(1.01); C1139(0.98) | LDD1600 | [31] |
| LDCM0397 | CL126 | HEK-293T | C135(1.04); C1872(1.04); C1139(1.01); C227(0.90) | LDD1601 | [31] |
| LDCM0398 | CL127 | HEK-293T | C135(1.04); C326(1.07); C1872(1.02); C1139(0.96) | LDD1602 | [31] |
| LDCM0399 | CL128 | HEK-293T | C326(1.01); C1872(0.98); C1139(1.03); C227(0.99) | LDD1603 | [31] |
| LDCM0400 | CL13 | HEK-293T | C135(1.06); C326(1.04); C1872(1.03); C1139(1.12) | LDD1604 | [31] |
| LDCM0401 | CL14 | HEK-293T | C135(1.23); C1872(0.88); C1139(0.97); C227(0.89) | LDD1605 | [31] |
| LDCM0402 | CL15 | HEK-293T | C135(0.97); C326(1.00); C1872(1.07); C1139(0.94) | LDD1606 | [31] |
| LDCM0403 | CL16 | HEK-293T | C326(1.09); C1872(0.85); C1139(0.94); C227(0.95) | LDD1607 | [31] |
| LDCM0404 | CL17 | HEK-293T | C1872(0.86); C1139(1.04); C227(0.93); C1187(1.16) | LDD1608 | [31] |
| LDCM0405 | CL18 | HEK-293T | C135(1.20); C326(1.00); C1872(0.82); C1139(1.04) | LDD1609 | [31] |
| LDCM0406 | CL19 | HEK-293T | C135(1.17); C1872(0.81); C1139(0.99); C227(0.85) | LDD1610 | [31] |
| LDCM0407 | CL2 | HEK-293T | C135(0.92); C1872(0.84); C1139(1.01); C227(0.87) | LDD1611 | [31] |
| LDCM0408 | CL20 | HEK-293T | C135(1.07); C326(0.91); C1872(0.80); C1139(1.02) | LDD1612 | [31] |
| LDCM0409 | CL21 | HEK-293T | C135(0.98); C326(0.81); C1872(0.76); C1139(1.17) | LDD1613 | [31] |
| LDCM0410 | CL22 | HEK-293T | C135(0.95); C326(0.91); C1872(0.85); C1139(1.10) | LDD1614 | [31] |
| LDCM0411 | CL23 | HEK-293T | C135(1.05); C1872(0.86); C1139(1.14); C227(0.90) | LDD1615 | [31] |
| LDCM0412 | CL24 | HEK-293T | C135(0.86); C326(1.00); C1872(0.79); C1139(1.19) | LDD1616 | [31] |
| LDCM0413 | CL25 | HEK-293T | C135(0.97); C326(0.91); C1872(0.96); C1139(0.89) | LDD1617 | [31] |
| LDCM0414 | CL26 | HEK-293T | C135(1.03); C1872(0.96); C1139(0.98); C227(0.92) | LDD1618 | [31] |
| LDCM0415 | CL27 | HEK-293T | C135(1.05); C326(1.11); C1872(1.01); C1139(0.96) | LDD1619 | [31] |
| LDCM0416 | CL28 | HEK-293T | C326(0.81); C1872(0.95); C1139(0.96); C227(0.93) | LDD1620 | [31] |
| LDCM0417 | CL29 | HEK-293T | C1872(1.02); C1139(1.03); C227(0.98); C1187(1.12) | LDD1621 | [31] |
| LDCM0418 | CL3 | HEK-293T | C135(1.02); C326(0.89); C1872(1.02); C1139(0.93) | LDD1622 | [31] |
| LDCM0419 | CL30 | HEK-293T | C135(1.07); C326(0.94); C1872(0.84); C1139(1.02) | LDD1623 | [31] |
| LDCM0420 | CL31 | HEK-293T | C135(1.10); C1872(1.03); C1139(1.10); C227(0.94) | LDD1624 | [31] |
| LDCM0421 | CL32 | HEK-293T | C135(1.19); C326(0.76); C1872(0.86); C1139(1.04) | LDD1625 | [31] |
| LDCM0422 | CL33 | HEK-293T | C135(0.98); C326(0.91); C1872(0.90); C1139(0.93) | LDD1626 | [31] |
| LDCM0423 | CL34 | HEK-293T | C135(1.02); C326(0.81); C1872(0.93); C1139(1.10) | LDD1627 | [31] |
| LDCM0424 | CL35 | HEK-293T | C135(1.08); C1872(1.06); C1139(1.17); C227(1.03) | LDD1628 | [31] |
| LDCM0425 | CL36 | HEK-293T | C135(0.96); C326(1.19); C1872(0.93); C1139(1.14) | LDD1629 | [31] |
| LDCM0426 | CL37 | HEK-293T | C135(0.90); C326(1.10); C1872(0.86); C1139(0.90) | LDD1630 | [31] |
| LDCM0428 | CL39 | HEK-293T | C135(1.11); C326(1.08); C1872(0.98); C1139(0.94) | LDD1632 | [31] |
| LDCM0429 | CL4 | HEK-293T | C326(0.87); C1872(0.95); C1139(0.97); C227(0.95) | LDD1633 | [31] |
| LDCM0430 | CL40 | HEK-293T | C326(1.12); C1872(0.95); C1139(0.98); C227(0.98) | LDD1634 | [31] |
| LDCM0431 | CL41 | HEK-293T | C1872(1.06); C1139(1.02); C227(1.00); C1187(1.06) | LDD1635 | [31] |
| LDCM0432 | CL42 | HEK-293T | C135(1.07); C326(1.15); C1872(0.90); C1139(1.08) | LDD1636 | [31] |
| LDCM0433 | CL43 | HEK-293T | C135(1.08); C1872(0.98); C1139(1.04); C227(0.89) | LDD1637 | [31] |
| LDCM0434 | CL44 | HEK-293T | C135(1.19); C326(0.95); C1872(0.92); C1139(1.07) | LDD1638 | [31] |
| LDCM0435 | CL45 | HEK-293T | C135(0.98); C326(0.67); C1872(1.01); C1139(1.11) | LDD1639 | [31] |
| LDCM0436 | CL46 | HEK-293T | C135(1.05); C326(0.91); C1872(0.81); C1139(1.10) | LDD1640 | [31] |
| LDCM0437 | CL47 | HEK-293T | C135(1.18); C1872(0.91); C1139(1.17); C227(0.96) | LDD1641 | [31] |
| LDCM0438 | CL48 | HEK-293T | C135(1.03); C326(1.08); C1872(1.13); C1139(1.21) | LDD1642 | [31] |
| LDCM0439 | CL49 | HEK-293T | C135(1.11); C326(1.24); C1872(0.81); C1139(1.00) | LDD1643 | [31] |
| LDCM0440 | CL5 | HEK-293T | C1872(0.87); C1139(0.99); C227(0.99); C1187(1.07) | LDD1644 | [31] |
| LDCM0441 | CL50 | HEK-293T | C135(0.99); C1872(0.79); C1139(1.04); C227(0.87) | LDD1645 | [31] |
| LDCM0443 | CL52 | HEK-293T | C326(0.95); C1872(0.83); C1139(1.04); C227(0.97) | LDD1646 | [31] |
| LDCM0444 | CL53 | HEK-293T | C1872(0.95); C1139(1.05); C227(0.89); C1187(1.12) | LDD1647 | [31] |
| LDCM0445 | CL54 | HEK-293T | C135(1.14); C326(0.79); C1872(0.83); C1139(1.25) | LDD1648 | [31] |
| LDCM0446 | CL55 | HEK-293T | C135(1.19); C1872(0.90); C1139(1.03); C227(0.91) | LDD1649 | [31] |
| LDCM0447 | CL56 | HEK-293T | C135(1.20); C326(1.15); C1872(0.89); C1139(1.13) | LDD1650 | [31] |
| LDCM0448 | CL57 | HEK-293T | C135(1.00); C326(0.87); C1872(0.80); C1139(1.44) | LDD1651 | [31] |
| LDCM0449 | CL58 | HEK-293T | C135(0.97); C326(0.95); C1872(0.78); C1139(1.06) | LDD1652 | [31] |
| LDCM0450 | CL59 | HEK-293T | C135(1.06); C1872(0.85); C1139(1.12); C227(0.95) | LDD1653 | [31] |
| LDCM0451 | CL6 | HEK-293T | C135(1.05); C326(1.00); C1872(0.80); C1139(1.24) | LDD1654 | [31] |
| LDCM0452 | CL60 | HEK-293T | C135(0.98); C326(1.05); C1872(0.89); C1139(1.21) | LDD1655 | [31] |
| LDCM0453 | CL61 | HEK-293T | C135(0.98); C326(1.10); C1872(0.82); C1139(1.00) | LDD1656 | [31] |
| LDCM0454 | CL62 | HEK-293T | C135(1.11); C1872(0.93); C1139(1.03); C227(0.91) | LDD1657 | [31] |
| LDCM0455 | CL63 | HEK-293T | C135(1.13); C326(1.15); C1872(1.04); C1139(0.96) | LDD1658 | [31] |
| LDCM0456 | CL64 | HEK-293T | C326(0.76); C1872(0.86); C1139(1.04); C227(0.90) | LDD1659 | [31] |
| LDCM0457 | CL65 | HEK-293T | C1872(0.87); C1139(1.03); C227(1.00); C1187(1.19) | LDD1660 | [31] |
| LDCM0458 | CL66 | HEK-293T | C135(1.10); C326(1.08); C1872(0.86); C1139(1.04) | LDD1661 | [31] |
| LDCM0459 | CL67 | HEK-293T | C135(1.21); C1872(0.91); C1139(1.01); C227(0.97) | LDD1662 | [31] |
| LDCM0460 | CL68 | HEK-293T | C135(1.10); C326(0.92); C1872(0.89); C1139(1.09) | LDD1663 | [31] |
| LDCM0461 | CL69 | HEK-293T | C135(1.00); C326(0.87); C1872(0.80); C1139(1.13) | LDD1664 | [31] |
| LDCM0462 | CL7 | HEK-293T | C135(1.12); C1872(0.91); C1139(1.00); C227(0.88) | LDD1665 | [31] |
| LDCM0463 | CL70 | HEK-293T | C135(1.00); C326(0.89); C1872(0.89); C1139(1.06) | LDD1666 | [31] |
| LDCM0464 | CL71 | HEK-293T | C135(1.13); C1872(0.88); C1139(1.13); C227(0.96) | LDD1667 | [31] |
| LDCM0465 | CL72 | HEK-293T | C135(0.97); C326(0.94); C1872(0.95); C1139(1.21) | LDD1668 | [31] |
| LDCM0466 | CL73 | HEK-293T | C135(1.01); C326(0.96); C1872(0.88); C1139(0.94) | LDD1669 | [31] |
| LDCM0467 | CL74 | HEK-293T | C135(1.06); C1872(0.85); C1139(1.03); C227(0.87) | LDD1670 | [31] |
| LDCM0469 | CL76 | HEK-293T | C326(1.06); C1872(0.97); C1139(0.97); C227(0.97) | LDD1672 | [31] |
| LDCM0470 | CL77 | HEK-293T | C1872(0.99); C1139(1.10); C227(0.99); C1187(1.06) | LDD1673 | [31] |
| LDCM0471 | CL78 | HEK-293T | C135(1.11); C326(1.16); C1872(0.90); C1139(1.00) | LDD1674 | [31] |
| LDCM0472 | CL79 | HEK-293T | C135(1.20); C1872(0.94); C1139(0.97); C227(0.85) | LDD1675 | [31] |
| LDCM0473 | CL8 | HEK-293T | C135(0.98); C326(0.98); C1872(0.87); C1139(1.25) | LDD1676 | [31] |
| LDCM0474 | CL80 | HEK-293T | C135(1.19); C326(0.91); C1872(0.82); C1139(0.99) | LDD1677 | [31] |
| LDCM0475 | CL81 | HEK-293T | C135(1.01); C326(0.99); C1872(0.86); C1139(1.05) | LDD1678 | [31] |
| LDCM0476 | CL82 | HEK-293T | C135(1.04); C326(0.93); C1872(0.85); C1139(1.09) | LDD1679 | [31] |
| LDCM0477 | CL83 | HEK-293T | C135(1.16); C1872(0.97); C1139(1.09); C227(0.90) | LDD1680 | [31] |
| LDCM0478 | CL84 | HEK-293T | C135(1.01); C326(1.05); C1872(0.94); C1139(1.18) | LDD1681 | [31] |
| LDCM0479 | CL85 | HEK-293T | C135(0.97); C326(1.46); C1872(0.85); C1139(0.94) | LDD1682 | [31] |
| LDCM0480 | CL86 | HEK-293T | C135(0.95); C1872(0.86); C1139(1.02); C227(0.86) | LDD1683 | [31] |
| LDCM0481 | CL87 | HEK-293T | C135(1.06); C326(1.07); C1872(1.01); C1139(0.97) | LDD1684 | [31] |
| LDCM0482 | CL88 | HEK-293T | C326(1.14); C1872(0.89); C1139(1.02); C227(0.97) | LDD1685 | [31] |
| LDCM0483 | CL89 | HEK-293T | C1872(0.88); C1139(0.98); C227(1.00); C1187(1.09) | LDD1686 | [31] |
| LDCM0484 | CL9 | HEK-293T | C135(0.97); C326(0.77); C1872(0.79); C1139(1.11) | LDD1687 | [31] |
| LDCM0485 | CL90 | HEK-293T | C135(1.00); C326(1.04); C1872(0.80); C1139(1.62) | LDD1688 | [31] |
| LDCM0486 | CL91 | HEK-293T | C135(1.18); C1872(0.85); C1139(0.98); C227(0.84) | LDD1689 | [31] |
| LDCM0487 | CL92 | HEK-293T | C135(1.12); C326(1.00); C1872(0.83); C1139(1.04) | LDD1690 | [31] |
| LDCM0488 | CL93 | HEK-293T | C135(1.02); C326(0.76); C1872(0.87); C1139(0.95) | LDD1691 | [31] |
| LDCM0489 | CL94 | HEK-293T | C135(0.92); C326(0.82); C1872(0.86); C1139(1.13) | LDD1692 | [31] |
| LDCM0490 | CL95 | HEK-293T | C135(1.00); C1872(1.03); C1139(1.47); C227(0.90) | LDD1693 | [31] |
| LDCM0491 | CL96 | HEK-293T | C135(0.91); C326(0.98); C1872(0.83); C1139(1.12) | LDD1694 | [31] |
| LDCM0492 | CL97 | HEK-293T | C135(0.97); C326(0.81); C1872(0.98); C1139(1.00) | LDD1695 | [31] |
| LDCM0493 | CL98 | HEK-293T | C135(1.01); C1872(0.95); C1139(1.04); C227(0.87) | LDD1696 | [31] |
| LDCM0494 | CL99 | HEK-293T | C135(0.96); C326(1.06); C1872(1.07); C1139(1.02) | LDD1697 | [31] |
| LDCM0495 | E2913 | HEK-293T | C135(1.12); C326(1.19); C1872(1.01); C1139(1.00) | LDD1698 | [31] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C135(2.45); C1139(1.76); C1886(1.11); C227(1.10) | LDD1702 | [8] |
| LDCM0625 | F8 | Ramos | C326(1.31); C227(2.08); C1139(0.97); C149(0.79) | LDD2187 | [33] |
| LDCM0572 | Fragment10 | MDA-MB-231 | C1872(1.51) | LDD1465 | [34] |
| LDCM0573 | Fragment11 | MDA-MB-231 | C1872(0.54) | LDD1467 | [34] |
| LDCM0574 | Fragment12 | Ramos | C326(0.31); C227(0.69); C1139(1.03); C149(0.39) | LDD2191 | [33] |
| LDCM0575 | Fragment13 | MDA-MB-231 | C1872(1.04) | LDD1469 | [34] |
| LDCM0576 | Fragment14 | MDA-MB-231 | C1872(1.10) | LDD1471 | [34] |
| LDCM0579 | Fragment20 | Ramos | C326(0.69); C227(2.12); C1139(0.94); 0.49 | LDD2194 | [33] |
| LDCM0580 | Fragment21 | Ramos | C326(0.69); C227(0.95); C1139(0.98); C149(0.64) | LDD2195 | [33] |
| LDCM0582 | Fragment23 | Ramos | C326(0.36); C227(0.78); C1139(1.33); 0.42 | LDD2196 | [33] |
| LDCM0578 | Fragment27 | Ramos | C326(1.47); C227(1.04); C1139(0.97) | LDD2197 | [33] |
| LDCM0586 | Fragment28 | Ramos | C326(0.67); C227(1.27); C1139(1.00); 1.22 | LDD2198 | [33] |
| LDCM0588 | Fragment30 | Ramos | C326(1.20); C227(0.73); C1139(1.38); 1.13 | LDD2199 | [33] |
| LDCM0589 | Fragment31 | Ramos | C326(1.60); C227(1.12); C1139(1.23); 1.21 | LDD2200 | [33] |
| LDCM0590 | Fragment32 | Ramos | C326(0.52); C227(0.63); C1139(2.83); C149(0.47) | LDD2201 | [33] |
| LDCM0468 | Fragment33 | HEK-293T | C135(1.08); C326(0.94); C1872(1.03); C1139(0.91) | LDD1671 | [31] |
| LDCM0596 | Fragment38 | MDA-MB-231 | C1872(1.28) | LDD1480 | [34] |
| LDCM0566 | Fragment4 | Ramos | C326(0.90); C227(1.18); C1139(1.03); 1.13 | LDD2184 | [33] |
| LDCM0599 | Fragment41 | MDA-MB-231 | C1872(0.47) | LDD1481 | [34] |
| LDCM0427 | Fragment51 | HEK-293T | C135(1.07); C1872(0.90); C1139(1.03); C227(0.91) | LDD1631 | [31] |
| LDCM0610 | Fragment52 | Ramos | C326(1.58); C227(0.63); C1139(1.15); 3.68 | LDD2204 | [33] |
| LDCM0614 | Fragment56 | Ramos | C326(2.17); C227(0.83); C1139(1.52); C149(1.26) | LDD2205 | [33] |
| LDCM0569 | Fragment7 | Ramos | C326(0.53); C227(0.73); C1139(1.51); 0.80 | LDD2186 | [33] |
| LDCM0571 | Fragment9 | Ramos | C1872(1.02) | LDD1464 | [34] |
| LDCM0107 | IAA | HeLa | H258(0.00); C1895(0.00) | LDD0221 | [26] |
| LDCM0022 | KB02 | MDA-MB-231 | C1872(4.07) | LDD1460 | [34] |
| LDCM0023 | KB03 | Jurkat | C1166(8.03); C1139(20.67) | LDD0209 | [11] |
| LDCM0024 | KB05 | COLO792 | C149(3.33); C227(2.32); C135(2.57); C1916(5.58) | LDD3310 | [35] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C135(1.61) | LDD2102 | [8] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C326(0.66); C135(0.87); C1872(0.62) | LDD2121 | [8] |
| LDCM0109 | NEM | HeLa | H258(0.00); H203(0.00); H140(0.00); H1235(0.00) | LDD0223 | [26] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C1139(0.34) | LDD2089 | [8] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C135(1.14) | LDD2090 | [8] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C326(1.16); C1139(1.06) | LDD2092 | [8] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C326(0.92); C135(2.93); C1139(0.87) | LDD2093 | [8] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C326(0.26) | LDD2096 | [8] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C326(0.97); C1872(0.65) | LDD2097 | [8] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C326(0.67); C135(0.72) | LDD2098 | [8] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C326(1.15); C1166(0.72); C1872(0.99); C1139(0.82) | LDD2099 | [8] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C326(0.37); C135(0.40); C1166(0.59) | LDD2100 | [8] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C326(1.22); C1166(1.40); C1872(1.23) | LDD2101 | [8] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C326(0.56); C135(0.83) | LDD2104 | [8] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C326(1.59) | LDD2105 | [8] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C326(0.83); C135(1.57); C1872(0.77); C1139(0.68) | LDD2107 | [8] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C326(0.39) | LDD2108 | [8] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C326(0.46); C135(1.78); C1166(0.40); C1872(0.51) | LDD2109 | [8] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C326(0.85); C135(2.37); C1872(0.80) | LDD2111 | [8] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C326(0.46); C1166(0.58); C1872(0.58) | LDD2115 | [8] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C326(0.67) | LDD2116 | [8] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C1166(1.07) | LDD2118 | [8] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C326(1.31); C135(1.37); C1872(1.82) | LDD2119 | [8] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C326(0.58); C1872(1.01) | LDD2120 | [8] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C326(0.79); C135(1.72); C1166(0.69); C1872(0.61) | LDD2123 | [8] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C326(0.47) | LDD2124 | [8] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C326(0.84); C1872(0.59); C1139(0.80) | LDD2125 | [8] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C326(0.40); C1872(1.15) | LDD2126 | [8] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C326(0.73); C1166(0.52); C1872(0.67) | LDD2127 | [8] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C326(0.46); C135(1.11); C1872(1.20) | LDD2128 | [8] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C326(1.01); C135(1.96); C1166(0.75) | LDD2129 | [8] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C326(0.62); C1166(1.16) | LDD2133 | [8] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C326(0.53); C135(1.69) | LDD2134 | [8] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C326(1.20); C135(3.35); C1872(0.95) | LDD2135 | [8] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C326(0.86); C135(1.50); C1166(0.66); C1872(0.87) | LDD2136 | [8] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C326(0.86); C135(1.81); C1872(0.75) | LDD2137 | [8] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C326(2.29); C1166(1.17) | LDD1700 | [8] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C326(0.84); C1872(0.59); C1139(0.41) | LDD2140 | [8] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C326(0.56); C1872(1.13) | LDD2143 | [8] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C326(1.94); C135(0.64); C1872(3.64) | LDD2144 | [8] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C326(0.77) | LDD2145 | [8] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C326(0.75); C135(1.03) | LDD2146 | [8] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C135(2.74) | LDD2148 | [8] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C326(0.37); C1872(1.08) | LDD2149 | [8] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C135(0.66); C1872(0.56); C1139(0.25) | LDD2150 | [8] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C326(0.27) | LDD2151 | [8] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C135(1.87) | LDD2153 | [8] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C227(1.01) | LDD2206 | [36] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C135(1.20); C227(0.80); C1872(0.73); C1139(0.70) | LDD2207 | [36] |
| LDCM0131 | RA190 | MM1.R | C1139(2.24); C1872(1.36); C1886(1.31); C1895(1.31) | LDD0304 | [37] |
| LDCM0170 | Structure8 | Ramos | 7.08 | LDD0433 | [38] |
| LDCM0003 | Sulforaphane | MDA-MB-231 | 1.98 | LDD0160 | [9] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transcription factor
Other
References
