Details of the Target
General Information of Target
| Target ID | LDTP01986 | |||||
|---|---|---|---|---|---|---|
| Target Name | NADH-ubiquinone oxidoreductase chain 5 (MT-ND5) | |||||
| Gene Name | MT-ND5 | |||||
| Gene ID | 4540 | |||||
| Synonyms |
MTND5; NADH5; ND5; NADH-ubiquinone oxidoreductase chain 5; EC 7.1.1.2; NADH dehydrogenase subunit 5 |
|||||
| 3D Structure | ||||||
| Sequence |
MTMHTTMTTLTLTSLIPPILTTLVNPNKKNSYPHYVKSIVASTFIISLFPTTMFMCLDQE
VIISNWHWATTQTTQLSLSFKLDYFSMMFIPVALFVTWSIMEFSLWYMNSDPNINQFFKY LLIFLITMLILVTANNLFQLFIGWEGVGIMSFLLISWWYARADANTAAIQAILYNRIGDI GFILALAWFILHSNSWDPQQMALLNANPSLTPLLGLLLAAAGKSAQLGLHPWLPSAMEGP TPVSALLHSSTMVVAGIFLLIRFHPLAENSPLIQTLTLCLGAITTLFAAVCALTQNDIKK IVAFSTSSQLGLMMVTIGINQPHLAFLHICTHAFFKAMLFMCSGSIIHNLNNEQDIRKMG GLLKTMPLTSTSLTIGSLALAGMPFLTGFYSKDHIIETANMSYTNAWALSITLIATSLTS AYSTRMILLTLTGQPRFPTLTNINENNPTLLNPIKRLAAGSLFAGFLITNNISPASPFQT TIPLYLKLTALAVTFLGLLTALDLNYLTNKLKMKSPLCTFYFSNMLGFYPSITHRTIPYL GLLTSQNLPLLLLDLTWLEKLLPKTISQHQISTSIITSTQKGMIKLYFLSFFFPLILTLL LIT |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Complex I subunit 5 family
|
|||||
| Subcellular location |
Mitochondrion inner membrane
|
|||||
| Function |
Core subunit of the mitochondrial membrane respiratory chain NADH dehydrogenase (Complex I) which catalyzes electron transfer from NADH through the respiratory chain, using ubiquinone as an electron acceptor. Essential for the catalytic activity and assembly of complex I.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Jackson_1 Probe Info |
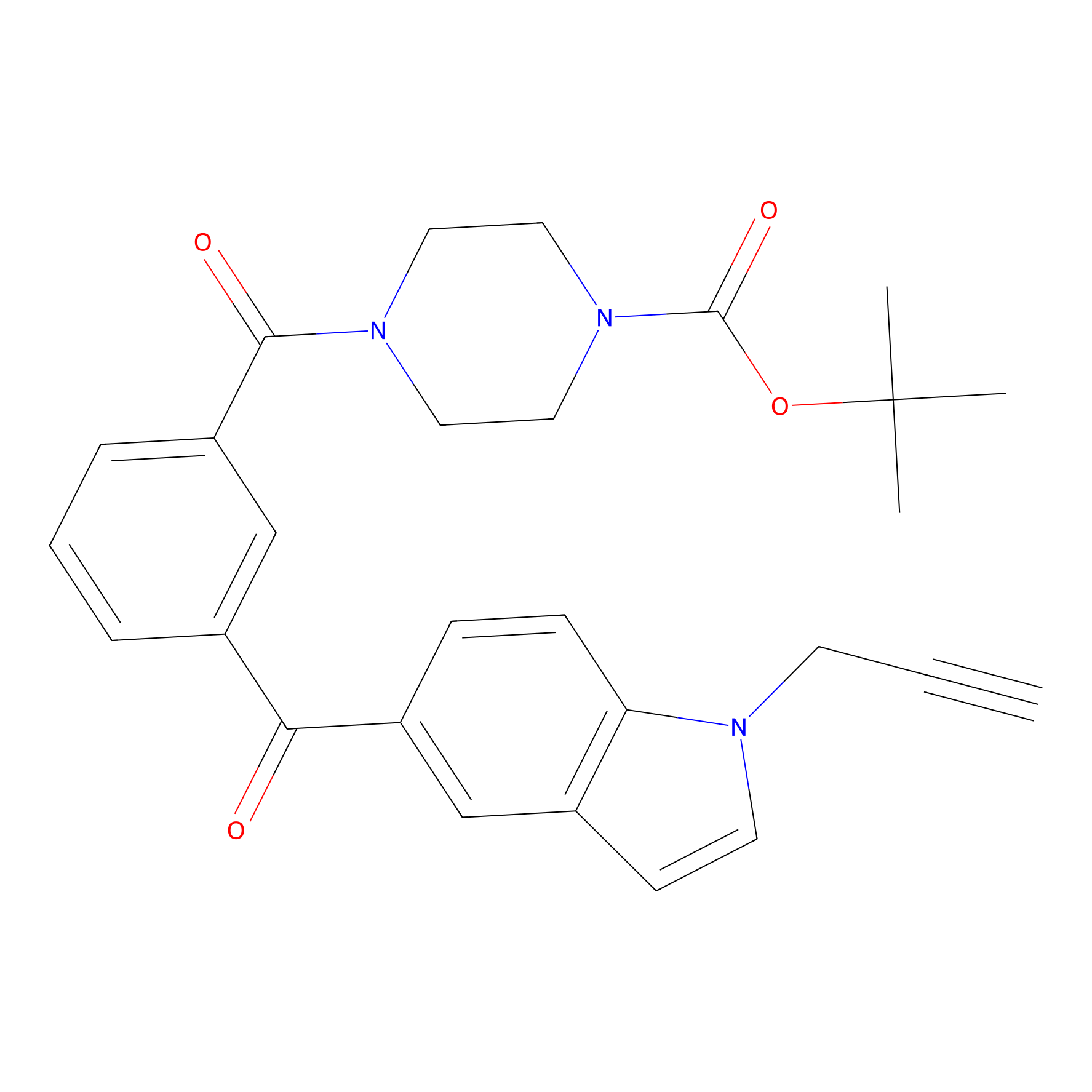 |
10.99 | LDD0122 | [1] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [2] | |
|
IPM Probe Info |
 |
N.A. | LDD0241 | [3] | |
|
Johansson_61 Probe Info |
 |
_(20.00) | LDD1487 | [4] | |
|
YY4-yne Probe Info |
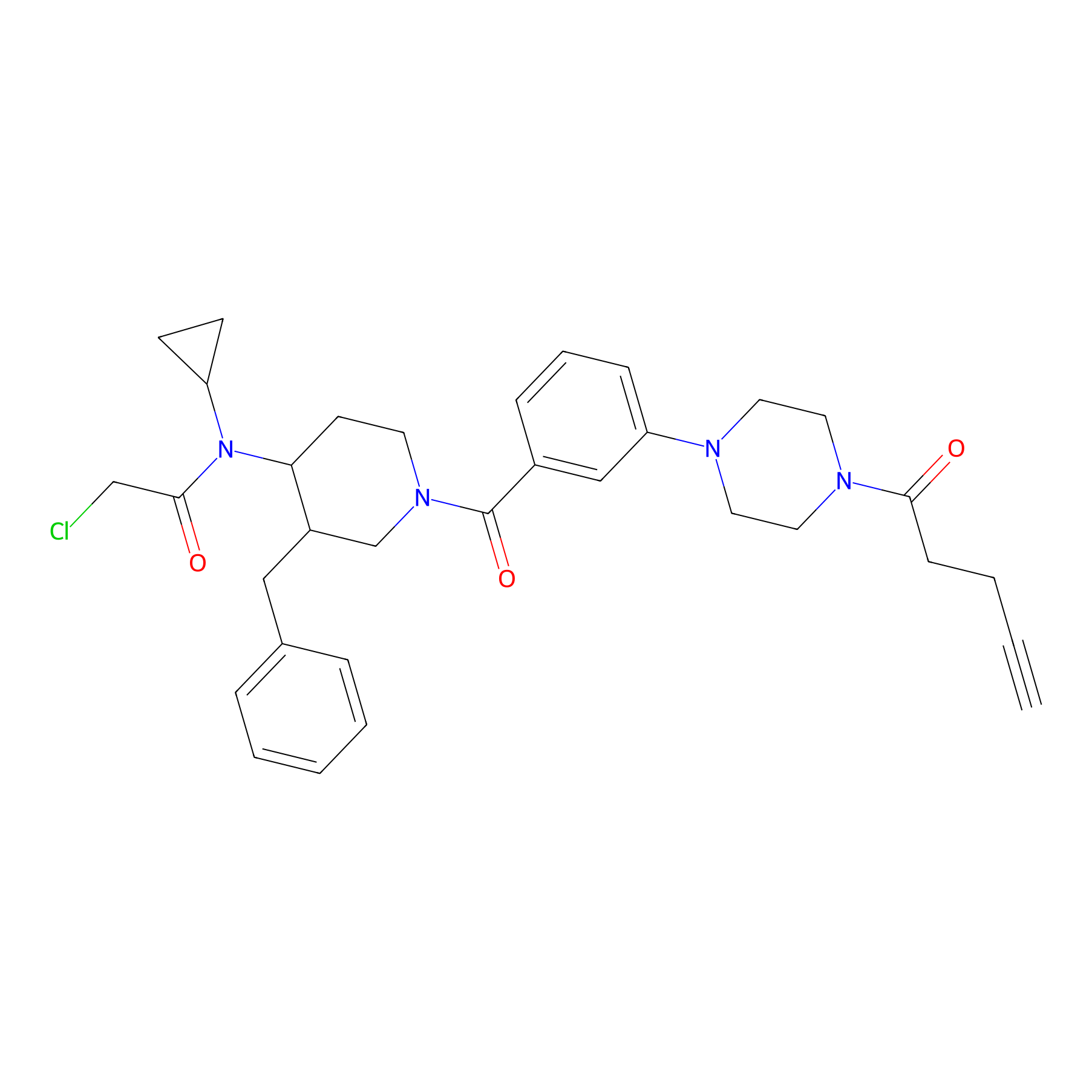 |
3.26 | LDD0400 | [5] | |
|
AHL-Pu-1 Probe Info |
 |
C518(2.65) | LDD0170 | [6] | |
|
DBIA Probe Info |
 |
C518(1.75) | LDD1095 | [7] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0162 | [8] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2224 | [9] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [10] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [11] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C091 Probe Info |
 |
13.64 | LDD1782 | [12] | |
|
C092 Probe Info |
 |
25.28 | LDD1783 | [12] | |
|
C094 Probe Info |
 |
43.11 | LDD1785 | [12] | |
|
C112 Probe Info |
 |
17.63 | LDD1799 | [12] | |
|
C429 Probe Info |
 |
11.88 | LDD2084 | [12] | |
|
C431 Probe Info |
 |
14.83 | LDD2086 | [12] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [13] | |
|
FFF probe13 Probe Info |
 |
18.55 | LDD0475 | [13] | |
|
FFF probe14 Probe Info |
 |
14.50 | LDD0477 | [13] | |
|
FFF probe3 Probe Info |
 |
12.25 | LDD0464 | [13] | |
|
JN0003 Probe Info |
 |
20.00 | LDD0469 | [13] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [14] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [15] | |
|
OEA-DA Probe Info |
 |
5.60 | LDD0046 | [16] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | DM93 | C518(2.65) | LDD0170 | [6] |
| LDCM0026 | 4SU-RNA+native RNA | DM93 | C518(8.43) | LDD0171 | [6] |
| LDCM0216 | AC100 | PaTu 8988t | C518(1.75) | LDD1095 | [7] |
| LDCM0217 | AC101 | PaTu 8988t | C518(2.01) | LDD1096 | [7] |
| LDCM0218 | AC102 | PaTu 8988t | C518(1.08) | LDD1097 | [7] |
| LDCM0219 | AC103 | PaTu 8988t | C518(1.06) | LDD1098 | [7] |
| LDCM0220 | AC104 | PaTu 8988t | C518(0.85) | LDD1099 | [7] |
| LDCM0221 | AC105 | PaTu 8988t | C518(1.31) | LDD1100 | [7] |
| LDCM0222 | AC106 | PaTu 8988t | C518(1.22) | LDD1101 | [7] |
| LDCM0223 | AC107 | PaTu 8988t | C518(1.32) | LDD1102 | [7] |
| LDCM0224 | AC108 | PaTu 8988t | C518(1.48) | LDD1103 | [7] |
| LDCM0225 | AC109 | PaTu 8988t | C518(1.95) | LDD1104 | [7] |
| LDCM0227 | AC110 | PaTu 8988t | C518(1.52) | LDD1106 | [7] |
| LDCM0228 | AC111 | PaTu 8988t | C518(1.17) | LDD1107 | [7] |
| LDCM0229 | AC112 | PaTu 8988t | C518(1.91) | LDD1108 | [7] |
| LDCM0230 | AC113 | PaTu 8988t | C518(0.73) | LDD1109 | [7] |
| LDCM0231 | AC114 | PaTu 8988t | C518(0.84) | LDD1110 | [7] |
| LDCM0232 | AC115 | PaTu 8988t | C518(0.76) | LDD1111 | [7] |
| LDCM0233 | AC116 | PaTu 8988t | C518(0.99) | LDD1112 | [7] |
| LDCM0234 | AC117 | PaTu 8988t | C518(0.91) | LDD1113 | [7] |
| LDCM0235 | AC118 | PaTu 8988t | C518(1.23) | LDD1114 | [7] |
| LDCM0236 | AC119 | PaTu 8988t | C518(0.89) | LDD1115 | [7] |
| LDCM0238 | AC120 | PaTu 8988t | C518(0.94) | LDD1117 | [7] |
| LDCM0239 | AC121 | PaTu 8988t | C518(0.86) | LDD1118 | [7] |
| LDCM0240 | AC122 | PaTu 8988t | C518(0.80) | LDD1119 | [7] |
| LDCM0241 | AC123 | PaTu 8988t | C518(1.19) | LDD1120 | [7] |
| LDCM0242 | AC124 | PaTu 8988t | C518(1.38) | LDD1121 | [7] |
| LDCM0243 | AC125 | PaTu 8988t | C518(1.53) | LDD1122 | [7] |
| LDCM0244 | AC126 | PaTu 8988t | C518(1.32) | LDD1123 | [7] |
| LDCM0245 | AC127 | PaTu 8988t | C518(1.20) | LDD1124 | [7] |
| LDCM0246 | AC128 | PaTu 8988t | C518(0.57) | LDD1125 | [7] |
| LDCM0247 | AC129 | PaTu 8988t | C518(0.54) | LDD1126 | [7] |
| LDCM0249 | AC130 | PaTu 8988t | C518(0.52) | LDD1128 | [7] |
| LDCM0250 | AC131 | PaTu 8988t | C518(1.77) | LDD1129 | [7] |
| LDCM0251 | AC132 | PaTu 8988t | C518(0.76) | LDD1130 | [7] |
| LDCM0252 | AC133 | PaTu 8988t | C518(0.83) | LDD1131 | [7] |
| LDCM0253 | AC134 | PaTu 8988t | C518(0.92) | LDD1132 | [7] |
| LDCM0254 | AC135 | PaTu 8988t | C518(0.86) | LDD1133 | [7] |
| LDCM0255 | AC136 | PaTu 8988t | C518(0.49) | LDD1134 | [7] |
| LDCM0256 | AC137 | PaTu 8988t | C518(0.57) | LDD1135 | [7] |
| LDCM0257 | AC138 | PaTu 8988t | C518(0.48) | LDD1136 | [7] |
| LDCM0258 | AC139 | PaTu 8988t | C518(0.48) | LDD1137 | [7] |
| LDCM0260 | AC140 | PaTu 8988t | C518(0.72) | LDD1139 | [7] |
| LDCM0261 | AC141 | PaTu 8988t | C518(0.69) | LDD1140 | [7] |
| LDCM0262 | AC142 | PaTu 8988t | C518(0.66) | LDD1141 | [7] |
| LDCM0285 | AC25 | PaTu 8988t | C518(0.66) | LDD1164 | [7] |
| LDCM0286 | AC26 | PaTu 8988t | C518(0.72) | LDD1165 | [7] |
| LDCM0287 | AC27 | PaTu 8988t | C518(0.95) | LDD1166 | [7] |
| LDCM0288 | AC28 | PaTu 8988t | C518(1.27) | LDD1167 | [7] |
| LDCM0289 | AC29 | PaTu 8988t | C518(0.65) | LDD1168 | [7] |
| LDCM0291 | AC30 | PaTu 8988t | C518(0.65) | LDD1170 | [7] |
| LDCM0292 | AC31 | PaTu 8988t | C518(0.97) | LDD1171 | [7] |
| LDCM0293 | AC32 | PaTu 8988t | C518(1.02) | LDD1172 | [7] |
| LDCM0294 | AC33 | PaTu 8988t | C518(1.87) | LDD1173 | [7] |
| LDCM0295 | AC34 | PaTu 8988t | C518(0.69) | LDD1174 | [7] |
| LDCM0308 | AC46 | PaTu 8988t | C518(1.20) | LDD1187 | [7] |
| LDCM0309 | AC47 | PaTu 8988t | C518(1.02) | LDD1188 | [7] |
| LDCM0310 | AC48 | PaTu 8988t | C518(1.01) | LDD1189 | [7] |
| LDCM0311 | AC49 | PaTu 8988t | C518(1.39) | LDD1190 | [7] |
| LDCM0313 | AC50 | PaTu 8988t | C518(1.63) | LDD1192 | [7] |
| LDCM0314 | AC51 | PaTu 8988t | C518(1.47) | LDD1193 | [7] |
| LDCM0315 | AC52 | PaTu 8988t | C518(1.59) | LDD1194 | [7] |
| LDCM0316 | AC53 | PaTu 8988t | C518(1.57) | LDD1195 | [7] |
| LDCM0317 | AC54 | PaTu 8988t | C518(1.84) | LDD1196 | [7] |
| LDCM0318 | AC55 | PaTu 8988t | C518(1.01) | LDD1197 | [7] |
| LDCM0319 | AC56 | PaTu 8988t | C518(2.33) | LDD1198 | [7] |
| LDCM0349 | AC83 | PaTu 8988t | C518(1.16) | LDD1228 | [7] |
| LDCM0350 | AC84 | PaTu 8988t | C518(1.14) | LDD1229 | [7] |
| LDCM0351 | AC85 | PaTu 8988t | C518(1.08) | LDD1230 | [7] |
| LDCM0352 | AC86 | PaTu 8988t | C518(1.21) | LDD1231 | [7] |
| LDCM0353 | AC87 | PaTu 8988t | C518(0.94) | LDD1232 | [7] |
| LDCM0354 | AC88 | PaTu 8988t | C518(1.15) | LDD1233 | [7] |
| LDCM0355 | AC89 | PaTu 8988t | C518(1.44) | LDD1234 | [7] |
| LDCM0357 | AC90 | PaTu 8988t | C518(1.02) | LDD1236 | [7] |
| LDCM0358 | AC91 | PaTu 8988t | C518(0.98) | LDD1237 | [7] |
| LDCM0359 | AC92 | PaTu 8988t | C518(1.12) | LDD1238 | [7] |
| LDCM0360 | AC93 | PaTu 8988t | C518(1.16) | LDD1239 | [7] |
| LDCM0361 | AC94 | PaTu 8988t | C518(1.11) | LDD1240 | [7] |
| LDCM0362 | AC95 | PaTu 8988t | C518(1.53) | LDD1241 | [7] |
| LDCM0363 | AC96 | PaTu 8988t | C518(0.76) | LDD1242 | [7] |
| LDCM0364 | AC97 | PaTu 8988t | C518(1.82) | LDD1243 | [7] |
| LDCM0365 | AC98 | PaTu 8988t | C518(1.81) | LDD1244 | [7] |
| LDCM0366 | AC99 | PaTu 8988t | C518(1.28) | LDD1245 | [7] |
| LDCM0382 | CL112 | PaTu 8988t | C518(1.38) | LDD1261 | [7] |
| LDCM0383 | CL113 | PaTu 8988t | C518(0.78) | LDD1262 | [7] |
| LDCM0384 | CL114 | PaTu 8988t | C518(2.62) | LDD1263 | [7] |
| LDCM0385 | CL115 | PaTu 8988t | C518(1.15) | LDD1264 | [7] |
| LDCM0386 | CL116 | PaTu 8988t | C518(1.00) | LDD1265 | [7] |
| LDCM0392 | CL121 | PaTu 8988t | C518(1.19) | LDD1271 | [7] |
| LDCM0393 | CL122 | PaTu 8988t | C518(4.45) | LDD1272 | [7] |
| LDCM0394 | CL123 | PaTu 8988t | C518(10.40) | LDD1273 | [7] |
| LDCM0395 | CL124 | PaTu 8988t | C518(3.34) | LDD1274 | [7] |
| LDCM0469 | CL76 | PaTu 8988t | C518(1.00) | LDD1348 | [7] |
| LDCM0470 | CL77 | PaTu 8988t | C518(4.01) | LDD1349 | [7] |
| LDCM0471 | CL78 | PaTu 8988t | C518(1.32) | LDD1350 | [7] |
| LDCM0472 | CL79 | PaTu 8988t | C518(1.29) | LDD1351 | [7] |
| LDCM0474 | CL80 | PaTu 8988t | C518(1.65) | LDD1353 | [7] |
| LDCM0475 | CL81 | PaTu 8988t | C518(1.83) | LDD1354 | [7] |
| LDCM0476 | CL82 | PaTu 8988t | C518(0.98) | LDD1355 | [7] |
| LDCM0477 | CL83 | PaTu 8988t | C518(1.59) | LDD1356 | [7] |
| LDCM0478 | CL84 | PaTu 8988t | C518(1.17) | LDD1357 | [7] |
| LDCM0479 | CL85 | PaTu 8988t | C518(1.89) | LDD1358 | [7] |
| LDCM0480 | CL86 | PaTu 8988t | C518(1.00) | LDD1359 | [7] |
| LDCM0481 | CL87 | PaTu 8988t | C518(1.89) | LDD1360 | [7] |
| LDCM0482 | CL88 | PaTu 8988t | C518(1.47) | LDD1361 | [7] |
| LDCM0483 | CL89 | PaTu 8988t | C518(1.57) | LDD1362 | [7] |
| LDCM0485 | CL90 | PaTu 8988t | C518(1.85) | LDD1364 | [7] |
| LDCM0191 | Compound 21 | HEK-293T | 6.62 | LDD0508 | [13] |
| LDCM0192 | Compound 35 | HEK-293T | 3.52 | LDD0509 | [13] |
| LDCM0193 | Compound 36 | HEK-293T | 5.89 | LDD0511 | [13] |
| LDCM0615 | Fragment63-R | Jurkat | _(20.00) | LDD1487 | [4] |
| LDCM0154 | YY4 | T cell | 3.26 | LDD0400 | [5] |
The Interaction Atlas With This Target
References
