Details of the Target
General Information of Target
| Target ID | LDTP13241 | |||||
|---|---|---|---|---|---|---|
| Target Name | LIM domain-containing protein 1 (LIMD1) | |||||
| Gene Name | LIMD1 | |||||
| Gene ID | 8994 | |||||
| Synonyms |
LIM domain-containing protein 1 |
|||||
| 3D Structure | ||||||
| Sequence |
MAARRRRSTGGGRARALNESKRVNNGNTAPEDSSPAKKTRRCQRQESKKMPVAGGKANKD
RTEDKQDGMPGRSWASKRVSESVKALLLKGKAPVDPECTAKVGKAHVYCEGNDVYDVMLN QTNLQFNNNKYYLIQLLEDDAQRNFSVWMRWGRVGKMGQHSLVACSGNLNKAKEIFQKKF LDKTKNNWEDREKFEKVPGKYDMLQMDYATNTQDEEETKKEESLKSPLKPESQLDLRVQE LIKLICNVQAMEEMMMEMKYNTKKAPLGKLTVAQIKAGYQSLKKIEDCIRAGQHGRALME ACNEFYTRIPHDFGLRTPPLIRTQKELSEKIQLLEALGDIEIAIKLVKTELQSPEHPLDQ HYRNLHCALRPLDHESYEFKVISQYLQSTHAPTHSDYTMTLLDLFEVEKDGEKEAFREDL HNRMLLWHGSRMSNWVGILSHGLRIAPPEAPITGYMFGKGIYFADMSSKSANYCFASRLK NTGLLLLSEVALGQCNELLEANPKAEGLLQGKHSTKGLGKMAPSSAHFVTLNGSTVPLGP ASDTGILNPDGYTLNYNEYIVYNPNQVRMRYLLKVQFNFLQLW |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Zyxin/ajuba family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Adapter or scaffold protein which participates in the assembly of numerous protein complexes and is involved in several cellular processes such as cell fate determination, cytoskeletal organization, repression of gene transcription, cell-cell adhesion, cell differentiation, proliferation and migration. Positively regulates microRNA (miRNA)-mediated gene silencing and is essential for P-body formation and integrity. Acts as a hypoxic regulator by bridging an association between the prolyl hydroxylases and VHL enabling efficient degradation of HIF1A. Acts as a transcriptional corepressor for SNAI1- and SNAI2/SLUG-dependent repression of E-cadherin transcription. Negatively regulates the Hippo signaling pathway and antagonizes phosphorylation of YAP1. Inhibits E2F-mediated transcription, and suppresses the expression of the majority of genes with E2F1-responsive elements. Regulates osteoblast development, function, differentiation and stress osteoclastogenesis. Enhances the ability of TRAF6 to activate adapter protein complex 1 (AP-1) and negatively regulates the canonical Wnt receptor signaling pathway in osteoblasts. May act as a tumor suppressor by inhibiting cell proliferation.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
DA-P2 Probe Info |
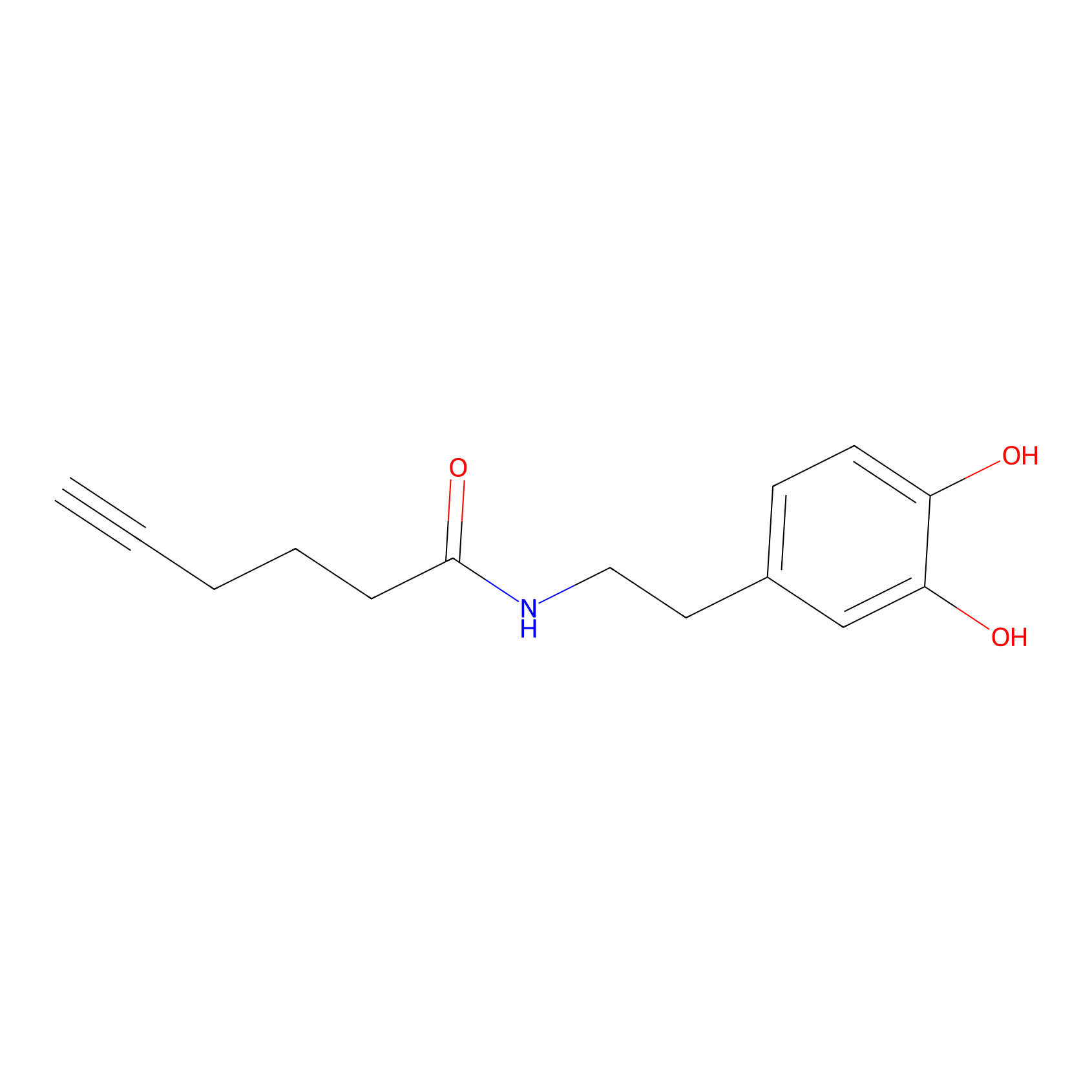 |
9.56 | LDD0348 | [2] | |
|
HRP Probe Info |
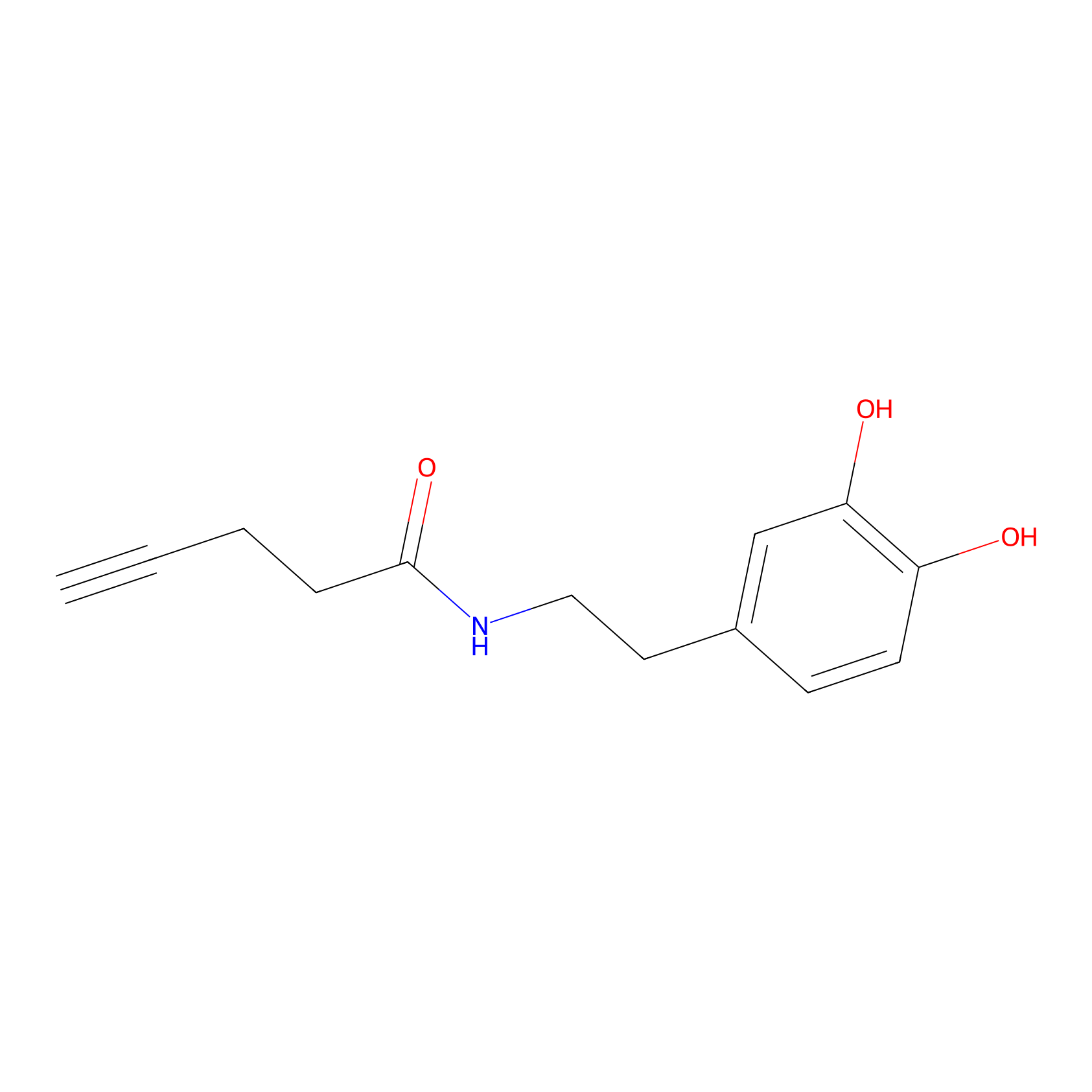 |
5.38 | LDD0347 | [2] | |
|
BTD Probe Info |
 |
C305(1.14) | LDD2090 | [3] | |
|
DA-P3 Probe Info |
 |
19.45 | LDD0179 | [2] | |
|
AHL-Pu-1 Probe Info |
 |
C401(20.00); C305(5.75); C148(20.00) | LDD0169 | [4] | |
|
DBIA Probe Info |
 |
C305(0.95) | LDD0078 | [5] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C472(0.00); C148(0.00); C560(0.00); C431(0.00) | LDD0038 | [6] | |
|
IA-alkyne Probe Info |
 |
C472(0.00); C148(0.00); C560(0.00) | LDD0036 | [6] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [6] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [7] | |
|
NAIA_4 Probe Info |
 |
C305(0.00); C374(0.00); C401(0.00); C431(0.00) | LDD2226 | [8] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [7] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [9] | |
|
aHNE Probe Info |
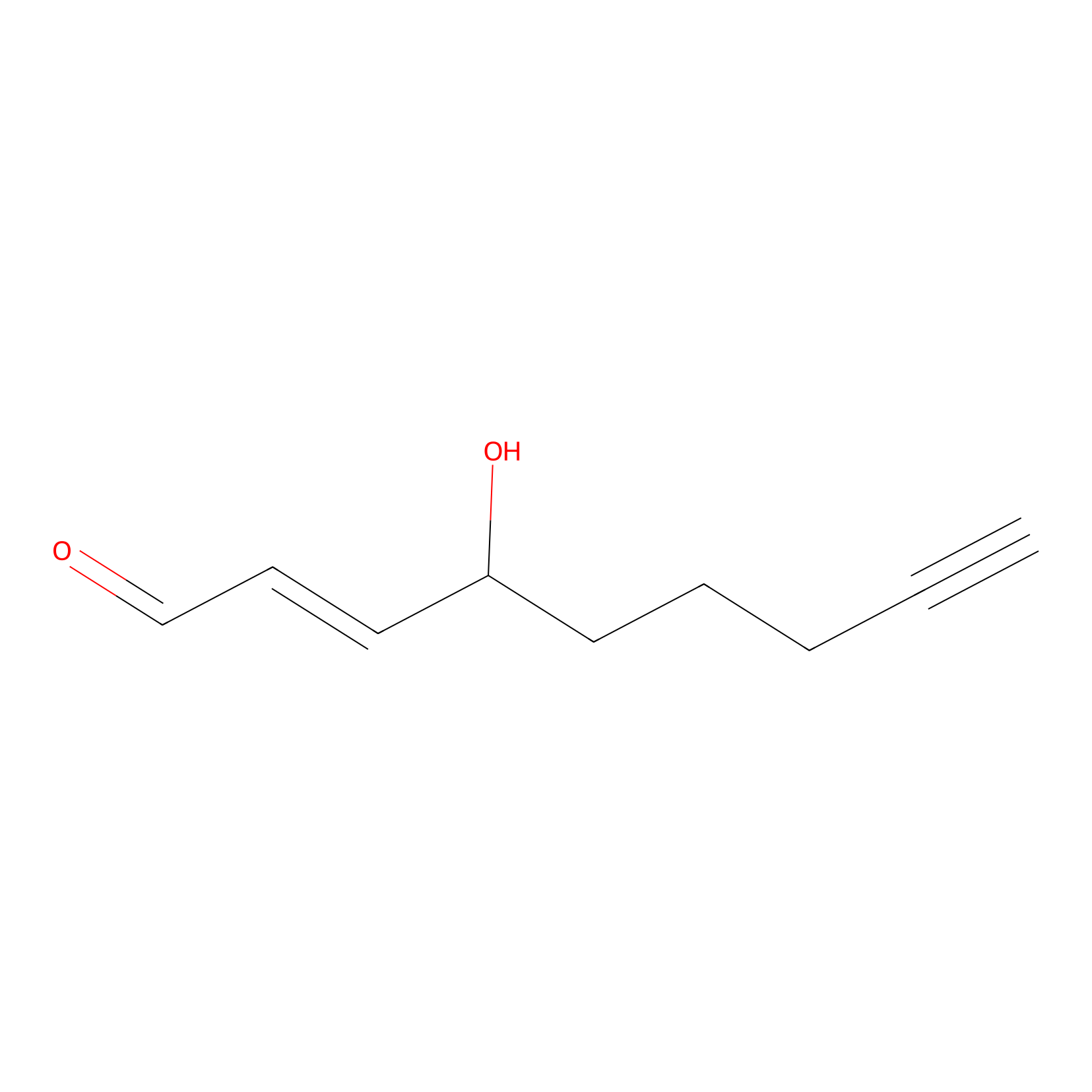 |
N.A. | LDD0001 | [9] | |
|
Compound 10 Probe Info |
 |
C305(0.00); C431(0.00) | LDD2216 | [10] | |
|
Compound 11 Probe Info |
 |
C305(0.00); C431(0.00) | LDD2213 | [10] | |
|
IPM Probe Info |
 |
C431(0.00); C401(0.00); C305(0.00) | LDD0005 | [9] | |
|
VSF Probe Info |
 |
C148(0.00); C374(0.00); C305(0.00); C401(0.00) | LDD0007 | [9] | |
|
Phosphinate-6 Probe Info |
 |
C305(0.00); C431(0.00); C374(0.00); C401(0.00) | LDD0018 | [11] | |
|
Methacrolein Probe Info |
 |
C222(0.00); C148(0.00) | LDD0218 | [12] | |
|
W1 Probe Info |
 |
C305(0.00); C148(0.00) | LDD0236 | [13] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [14] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [8] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C401(20.00); C305(5.75); C148(20.00) | LDD0169 | [4] |
| LDCM0214 | AC1 | HCT 116 | C222(1.13) | LDD0531 | [5] |
| LDCM0215 | AC10 | HCT 116 | C222(0.72) | LDD0532 | [5] |
| LDCM0216 | AC100 | HCT 116 | C222(0.96) | LDD0533 | [5] |
| LDCM0217 | AC101 | HCT 116 | C222(1.09) | LDD0534 | [5] |
| LDCM0218 | AC102 | HCT 116 | C222(0.75) | LDD0535 | [5] |
| LDCM0219 | AC103 | HCT 116 | C222(1.29) | LDD0536 | [5] |
| LDCM0220 | AC104 | HCT 116 | C222(1.03) | LDD0537 | [5] |
| LDCM0221 | AC105 | HCT 116 | C222(1.44) | LDD0538 | [5] |
| LDCM0222 | AC106 | HCT 116 | C222(0.98) | LDD0539 | [5] |
| LDCM0223 | AC107 | HCT 116 | C222(1.40) | LDD0540 | [5] |
| LDCM0224 | AC108 | HCT 116 | C222(0.88) | LDD0541 | [5] |
| LDCM0225 | AC109 | HCT 116 | C222(0.78) | LDD0542 | [5] |
| LDCM0226 | AC11 | HCT 116 | C222(1.14) | LDD0543 | [5] |
| LDCM0227 | AC110 | HCT 116 | C222(0.88) | LDD0544 | [5] |
| LDCM0228 | AC111 | HCT 116 | C222(0.81) | LDD0545 | [5] |
| LDCM0229 | AC112 | HCT 116 | C222(1.08) | LDD0546 | [5] |
| LDCM0230 | AC113 | HCT 116 | C222(0.97) | LDD0547 | [5] |
| LDCM0231 | AC114 | HCT 116 | C222(0.95) | LDD0548 | [5] |
| LDCM0232 | AC115 | HCT 116 | C222(1.09) | LDD0549 | [5] |
| LDCM0233 | AC116 | HCT 116 | C222(0.90) | LDD0550 | [5] |
| LDCM0234 | AC117 | HCT 116 | C222(1.63) | LDD0551 | [5] |
| LDCM0235 | AC118 | HCT 116 | C222(1.05) | LDD0552 | [5] |
| LDCM0236 | AC119 | HCT 116 | C222(1.05) | LDD0553 | [5] |
| LDCM0237 | AC12 | HCT 116 | C222(0.82) | LDD0554 | [5] |
| LDCM0238 | AC120 | HCT 116 | C222(0.75) | LDD0555 | [5] |
| LDCM0239 | AC121 | HCT 116 | C222(1.72) | LDD0556 | [5] |
| LDCM0240 | AC122 | HCT 116 | C222(1.00) | LDD0557 | [5] |
| LDCM0241 | AC123 | HCT 116 | C222(2.54) | LDD0558 | [5] |
| LDCM0242 | AC124 | HCT 116 | C222(1.28) | LDD0559 | [5] |
| LDCM0243 | AC125 | HCT 116 | C222(0.76) | LDD0560 | [5] |
| LDCM0244 | AC126 | HCT 116 | C222(0.87) | LDD0561 | [5] |
| LDCM0245 | AC127 | HCT 116 | C222(1.02) | LDD0562 | [5] |
| LDCM0246 | AC128 | HEK-293T | C597(1.52); C222(1.09) | LDD0844 | [5] |
| LDCM0247 | AC129 | HEK-293T | C597(0.61); C222(1.09) | LDD0845 | [5] |
| LDCM0249 | AC130 | HEK-293T | C597(0.75); C222(1.24) | LDD0847 | [5] |
| LDCM0250 | AC131 | HEK-293T | C597(0.73); C222(0.96) | LDD0848 | [5] |
| LDCM0251 | AC132 | HEK-293T | C597(0.75); C222(1.33) | LDD0849 | [5] |
| LDCM0252 | AC133 | HEK-293T | C597(0.48); C222(0.73) | LDD0850 | [5] |
| LDCM0253 | AC134 | HEK-293T | C597(0.54); C222(0.98) | LDD0851 | [5] |
| LDCM0254 | AC135 | HEK-293T | C597(0.54); C222(0.82) | LDD0852 | [5] |
| LDCM0255 | AC136 | HEK-293T | C597(0.70); C222(1.26) | LDD0853 | [5] |
| LDCM0256 | AC137 | HEK-293T | C597(0.45); C222(1.13) | LDD0854 | [5] |
| LDCM0257 | AC138 | HEK-293T | C597(0.44); C222(1.13) | LDD0855 | [5] |
| LDCM0258 | AC139 | HEK-293T | C597(0.43); C222(1.27) | LDD0856 | [5] |
| LDCM0259 | AC14 | HCT 116 | C222(0.68) | LDD0576 | [5] |
| LDCM0260 | AC140 | HEK-293T | C597(0.54); C222(0.97) | LDD0858 | [5] |
| LDCM0261 | AC141 | HEK-293T | C597(0.41); C222(1.22) | LDD0859 | [5] |
| LDCM0262 | AC142 | HEK-293T | C597(0.53); C222(1.19) | LDD0860 | [5] |
| LDCM0263 | AC143 | HCT 116 | C222(0.91) | LDD0580 | [5] |
| LDCM0264 | AC144 | HCT 116 | C222(1.10) | LDD0581 | [5] |
| LDCM0265 | AC145 | HCT 116 | C222(1.03) | LDD0582 | [5] |
| LDCM0266 | AC146 | HCT 116 | C222(0.90) | LDD0583 | [5] |
| LDCM0267 | AC147 | HCT 116 | C222(0.90) | LDD0584 | [5] |
| LDCM0268 | AC148 | HCT 116 | C222(0.94) | LDD0585 | [5] |
| LDCM0269 | AC149 | HCT 116 | C222(1.04) | LDD0586 | [5] |
| LDCM0270 | AC15 | HCT 116 | C222(1.01) | LDD0587 | [5] |
| LDCM0271 | AC150 | HCT 116 | C222(0.92) | LDD0588 | [5] |
| LDCM0272 | AC151 | HCT 116 | C222(1.07) | LDD0589 | [5] |
| LDCM0273 | AC152 | HCT 116 | C222(0.80) | LDD0590 | [5] |
| LDCM0274 | AC153 | HCT 116 | C222(0.90) | LDD0591 | [5] |
| LDCM0621 | AC154 | HCT 116 | C222(0.86) | LDD2158 | [5] |
| LDCM0622 | AC155 | HCT 116 | C222(1.04) | LDD2159 | [5] |
| LDCM0623 | AC156 | HCT 116 | C222(0.94) | LDD2160 | [5] |
| LDCM0624 | AC157 | HCT 116 | C222(0.85) | LDD2161 | [5] |
| LDCM0276 | AC17 | HCT 116 | C222(1.18) | LDD0593 | [5] |
| LDCM0277 | AC18 | HCT 116 | C222(0.68) | LDD0594 | [5] |
| LDCM0278 | AC19 | HCT 116 | C222(1.11) | LDD0595 | [5] |
| LDCM0279 | AC2 | HCT 116 | C222(1.06) | LDD0596 | [5] |
| LDCM0280 | AC20 | HCT 116 | C222(1.22) | LDD0597 | [5] |
| LDCM0281 | AC21 | HCT 116 | C222(1.10) | LDD0598 | [5] |
| LDCM0282 | AC22 | HCT 116 | C222(1.01) | LDD0599 | [5] |
| LDCM0283 | AC23 | HCT 116 | C222(1.14) | LDD0600 | [5] |
| LDCM0284 | AC24 | HCT 116 | C222(1.19) | LDD0601 | [5] |
| LDCM0285 | AC25 | HEK-293T | C597(1.07); C600(0.84) | LDD0883 | [5] |
| LDCM0286 | AC26 | HEK-293T | C597(1.05); C600(0.99) | LDD0884 | [5] |
| LDCM0287 | AC27 | HEK-293T | C597(0.95); C600(1.11) | LDD0885 | [5] |
| LDCM0288 | AC28 | HEK-293T | C597(0.80); C600(1.11) | LDD0886 | [5] |
| LDCM0289 | AC29 | HEK-293T | C597(0.79); C600(1.27) | LDD0887 | [5] |
| LDCM0290 | AC3 | HCT 116 | C222(0.99) | LDD0607 | [5] |
| LDCM0291 | AC30 | HEK-293T | C597(1.06); C600(0.75) | LDD0889 | [5] |
| LDCM0292 | AC31 | HEK-293T | C597(0.85); C600(0.99) | LDD0890 | [5] |
| LDCM0293 | AC32 | HEK-293T | C597(0.89); C600(1.15) | LDD0891 | [5] |
| LDCM0294 | AC33 | HEK-293T | C597(0.94); C600(0.97) | LDD0892 | [5] |
| LDCM0295 | AC34 | HEK-293T | C597(0.96); C600(1.16) | LDD0893 | [5] |
| LDCM0296 | AC35 | HCT 116 | C222(1.11) | LDD0613 | [5] |
| LDCM0297 | AC36 | HCT 116 | C222(1.28) | LDD0614 | [5] |
| LDCM0298 | AC37 | HCT 116 | C222(1.41) | LDD0615 | [5] |
| LDCM0299 | AC38 | HCT 116 | C222(1.08) | LDD0616 | [5] |
| LDCM0300 | AC39 | HCT 116 | C222(1.21) | LDD0617 | [5] |
| LDCM0301 | AC4 | HCT 116 | C222(1.21) | LDD0618 | [5] |
| LDCM0302 | AC40 | HCT 116 | C222(1.46) | LDD0619 | [5] |
| LDCM0303 | AC41 | HCT 116 | C222(1.52) | LDD0620 | [5] |
| LDCM0304 | AC42 | HCT 116 | C222(1.36) | LDD0621 | [5] |
| LDCM0305 | AC43 | HCT 116 | C222(1.50) | LDD0622 | [5] |
| LDCM0306 | AC44 | HCT 116 | C222(1.34) | LDD0623 | [5] |
| LDCM0307 | AC45 | HCT 116 | C222(1.52) | LDD0624 | [5] |
| LDCM0308 | AC46 | HCT 116 | C222(1.36) | LDD0625 | [5] |
| LDCM0309 | AC47 | HCT 116 | C222(1.45) | LDD0626 | [5] |
| LDCM0310 | AC48 | HCT 116 | C222(1.97) | LDD0627 | [5] |
| LDCM0311 | AC49 | HCT 116 | C222(1.78) | LDD0628 | [5] |
| LDCM0312 | AC5 | HCT 116 | C222(0.91) | LDD0629 | [5] |
| LDCM0313 | AC50 | HCT 116 | C222(1.37) | LDD0630 | [5] |
| LDCM0314 | AC51 | HCT 116 | C222(1.04) | LDD0631 | [5] |
| LDCM0315 | AC52 | HCT 116 | C222(1.35) | LDD0632 | [5] |
| LDCM0316 | AC53 | HCT 116 | C222(1.32) | LDD0633 | [5] |
| LDCM0317 | AC54 | HCT 116 | C222(1.58) | LDD0634 | [5] |
| LDCM0318 | AC55 | HCT 116 | C222(1.69) | LDD0635 | [5] |
| LDCM0319 | AC56 | HCT 116 | C222(2.11) | LDD0636 | [5] |
| LDCM0320 | AC57 | HCT 116 | C222(1.38) | LDD0637 | [5] |
| LDCM0321 | AC58 | HCT 116 | C222(1.48) | LDD0638 | [5] |
| LDCM0322 | AC59 | HCT 116 | C222(1.48) | LDD0639 | [5] |
| LDCM0323 | AC6 | HCT 116 | C222(1.05) | LDD0640 | [5] |
| LDCM0324 | AC60 | HCT 116 | C222(1.71) | LDD0641 | [5] |
| LDCM0325 | AC61 | HCT 116 | C222(1.54) | LDD0642 | [5] |
| LDCM0326 | AC62 | HCT 116 | C222(2.25) | LDD0643 | [5] |
| LDCM0327 | AC63 | HCT 116 | C222(1.09) | LDD0644 | [5] |
| LDCM0328 | AC64 | HCT 116 | C222(1.60) | LDD0645 | [5] |
| LDCM0329 | AC65 | HCT 116 | C222(0.56) | LDD0646 | [5] |
| LDCM0330 | AC66 | HCT 116 | C222(0.61) | LDD0647 | [5] |
| LDCM0331 | AC67 | HCT 116 | C222(0.63) | LDD0648 | [5] |
| LDCM0332 | AC68 | HEK-293T | C597(1.39); C222(0.86) | LDD0930 | [5] |
| LDCM0333 | AC69 | HEK-293T | C597(1.65); C222(0.99) | LDD0931 | [5] |
| LDCM0334 | AC7 | HCT 116 | C222(0.99) | LDD0651 | [5] |
| LDCM0335 | AC70 | HEK-293T | C597(1.02); C222(0.81) | LDD0933 | [5] |
| LDCM0336 | AC71 | HEK-293T | C597(1.10); C222(0.83) | LDD0934 | [5] |
| LDCM0337 | AC72 | HEK-293T | C597(2.57); C222(0.91) | LDD0935 | [5] |
| LDCM0338 | AC73 | HEK-293T | C597(0.81); C222(0.62) | LDD0936 | [5] |
| LDCM0339 | AC74 | HEK-293T | C597(1.29); C222(0.79) | LDD0937 | [5] |
| LDCM0340 | AC75 | HEK-293T | C597(1.96); C222(0.83) | LDD0938 | [5] |
| LDCM0341 | AC76 | HEK-293T | C597(1.58); C222(1.20) | LDD0939 | [5] |
| LDCM0342 | AC77 | HEK-293T | C597(1.43); C222(0.99) | LDD0940 | [5] |
| LDCM0343 | AC78 | HEK-293T | C597(1.61); C222(0.98) | LDD0941 | [5] |
| LDCM0344 | AC79 | HEK-293T | C597(1.33); C222(0.99) | LDD0942 | [5] |
| LDCM0345 | AC8 | HCT 116 | C222(0.88) | LDD0662 | [5] |
| LDCM0346 | AC80 | HEK-293T | C597(1.23); C222(1.03) | LDD0944 | [5] |
| LDCM0347 | AC81 | HEK-293T | C597(1.22); C222(1.18) | LDD0945 | [5] |
| LDCM0348 | AC82 | HEK-293T | C597(1.06); C222(1.20) | LDD0946 | [5] |
| LDCM0349 | AC83 | HCT 116 | C222(1.29) | LDD0666 | [5] |
| LDCM0350 | AC84 | HCT 116 | C222(1.12) | LDD0667 | [5] |
| LDCM0351 | AC85 | HCT 116 | C222(0.92) | LDD0668 | [5] |
| LDCM0352 | AC86 | HCT 116 | C222(0.97) | LDD0669 | [5] |
| LDCM0353 | AC87 | HCT 116 | C222(0.99) | LDD0670 | [5] |
| LDCM0354 | AC88 | HCT 116 | C222(1.12) | LDD0671 | [5] |
| LDCM0355 | AC89 | HCT 116 | C222(1.05) | LDD0672 | [5] |
| LDCM0357 | AC90 | HCT 116 | C222(1.20) | LDD0674 | [5] |
| LDCM0358 | AC91 | HCT 116 | C222(0.94) | LDD0675 | [5] |
| LDCM0359 | AC92 | HCT 116 | C222(1.14) | LDD0676 | [5] |
| LDCM0360 | AC93 | HCT 116 | C222(1.03) | LDD0677 | [5] |
| LDCM0361 | AC94 | HCT 116 | C222(0.65) | LDD0678 | [5] |
| LDCM0362 | AC95 | HCT 116 | C222(1.29) | LDD0679 | [5] |
| LDCM0363 | AC96 | HCT 116 | C222(0.85) | LDD0680 | [5] |
| LDCM0364 | AC97 | HCT 116 | C222(1.12) | LDD0681 | [5] |
| LDCM0365 | AC98 | HCT 116 | C222(1.95) | LDD0682 | [5] |
| LDCM0366 | AC99 | HCT 116 | C222(0.94) | LDD0683 | [5] |
| LDCM0248 | AKOS034007472 | HCT 116 | C222(0.70) | LDD0565 | [5] |
| LDCM0356 | AKOS034007680 | HCT 116 | C222(1.19) | LDD0673 | [5] |
| LDCM0275 | AKOS034007705 | HCT 116 | C222(0.80) | LDD0592 | [5] |
| LDCM0020 | ARS-1620 | HCC44 | C305(0.95) | LDD0078 | [5] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C431(0.57) | LDD2091 | [3] |
| LDCM0087 | Capsaicin | HEK-293T | 6.42 | LDD0185 | [2] |
| LDCM0632 | CL-Sc | Hep-G2 | C431(1.32); C431(0.79); C401(0.44) | LDD2227 | [8] |
| LDCM0367 | CL1 | HEK-293T | C222(1.10) | LDD0965 | [5] |
| LDCM0368 | CL10 | HEK-293T | C222(0.88) | LDD0966 | [5] |
| LDCM0369 | CL100 | HCT 116 | C222(1.09) | LDD0686 | [5] |
| LDCM0370 | CL101 | HCT 116 | C222(0.79) | LDD0687 | [5] |
| LDCM0371 | CL102 | HCT 116 | C222(0.81) | LDD0688 | [5] |
| LDCM0372 | CL103 | HCT 116 | C222(0.69) | LDD0689 | [5] |
| LDCM0373 | CL104 | HCT 116 | C222(0.69) | LDD0690 | [5] |
| LDCM0374 | CL105 | HCT 116 | C222(1.41) | LDD0691 | [5] |
| LDCM0375 | CL106 | HCT 116 | C222(1.44) | LDD0692 | [5] |
| LDCM0376 | CL107 | HCT 116 | C222(1.37) | LDD0693 | [5] |
| LDCM0377 | CL108 | HCT 116 | C222(1.31) | LDD0694 | [5] |
| LDCM0378 | CL109 | HCT 116 | C222(1.46) | LDD0695 | [5] |
| LDCM0379 | CL11 | HEK-293T | C222(0.85) | LDD0977 | [5] |
| LDCM0380 | CL110 | HCT 116 | C222(1.69) | LDD0697 | [5] |
| LDCM0381 | CL111 | HCT 116 | C222(1.23) | LDD0698 | [5] |
| LDCM0382 | CL112 | HEK-293T | C597(1.02); C600(1.34) | LDD0980 | [5] |
| LDCM0383 | CL113 | HEK-293T | C597(0.90); C600(1.09) | LDD0981 | [5] |
| LDCM0384 | CL114 | HEK-293T | C597(0.73); C600(1.11) | LDD0982 | [5] |
| LDCM0385 | CL115 | HEK-293T | C597(0.92); C600(1.56) | LDD0983 | [5] |
| LDCM0386 | CL116 | HEK-293T | C597(0.91); C600(1.04) | LDD0984 | [5] |
| LDCM0387 | CL117 | HCT 116 | C222(1.13) | LDD0704 | [5] |
| LDCM0388 | CL118 | HCT 116 | C222(1.15) | LDD0705 | [5] |
| LDCM0389 | CL119 | HCT 116 | C222(1.35) | LDD0706 | [5] |
| LDCM0390 | CL12 | HEK-293T | C222(0.90) | LDD0988 | [5] |
| LDCM0391 | CL120 | HCT 116 | C222(1.51) | LDD0708 | [5] |
| LDCM0392 | CL121 | HCT 116 | C222(1.00) | LDD0709 | [5] |
| LDCM0393 | CL122 | HCT 116 | C222(1.17) | LDD0710 | [5] |
| LDCM0394 | CL123 | HCT 116 | C222(1.30) | LDD0711 | [5] |
| LDCM0395 | CL124 | HCT 116 | C222(1.42) | LDD0712 | [5] |
| LDCM0396 | CL125 | HCT 116 | C222(1.05) | LDD0713 | [5] |
| LDCM0397 | CL126 | HCT 116 | C222(1.00) | LDD0714 | [5] |
| LDCM0398 | CL127 | HCT 116 | C222(0.85) | LDD0715 | [5] |
| LDCM0399 | CL128 | HCT 116 | C222(1.27) | LDD0716 | [5] |
| LDCM0400 | CL13 | HEK-293T | C222(0.84) | LDD0998 | [5] |
| LDCM0401 | CL14 | HEK-293T | C222(1.00) | LDD0999 | [5] |
| LDCM0402 | CL15 | HEK-293T | C222(0.89) | LDD1000 | [5] |
| LDCM0403 | CL16 | HCT 116 | C222(1.25) | LDD0720 | [5] |
| LDCM0404 | CL17 | HCT 116 | C222(1.06) | LDD0721 | [5] |
| LDCM0405 | CL18 | HCT 116 | C222(1.08) | LDD0722 | [5] |
| LDCM0406 | CL19 | HCT 116 | C222(1.07) | LDD0723 | [5] |
| LDCM0407 | CL2 | HEK-293T | C222(0.98) | LDD1005 | [5] |
| LDCM0408 | CL20 | HCT 116 | C222(1.18) | LDD0725 | [5] |
| LDCM0409 | CL21 | HCT 116 | C222(1.44) | LDD0726 | [5] |
| LDCM0410 | CL22 | HCT 116 | C222(1.28) | LDD0727 | [5] |
| LDCM0411 | CL23 | HCT 116 | C222(1.38) | LDD0728 | [5] |
| LDCM0412 | CL24 | HCT 116 | C222(1.06) | LDD0729 | [5] |
| LDCM0413 | CL25 | HCT 116 | C222(1.03) | LDD0730 | [5] |
| LDCM0414 | CL26 | HCT 116 | C222(1.66) | LDD0731 | [5] |
| LDCM0415 | CL27 | HCT 116 | C222(1.08) | LDD0732 | [5] |
| LDCM0416 | CL28 | HCT 116 | C222(1.32) | LDD0733 | [5] |
| LDCM0417 | CL29 | HCT 116 | C222(1.16) | LDD0734 | [5] |
| LDCM0418 | CL3 | HEK-293T | C222(0.80) | LDD1016 | [5] |
| LDCM0419 | CL30 | HCT 116 | C222(1.70) | LDD0736 | [5] |
| LDCM0420 | CL31 | HEK-293T | C597(2.03) | LDD1018 | [5] |
| LDCM0421 | CL32 | HEK-293T | C597(1.01) | LDD1019 | [5] |
| LDCM0422 | CL33 | HEK-293T | C597(0.91) | LDD1020 | [5] |
| LDCM0423 | CL34 | HEK-293T | C597(0.87) | LDD1021 | [5] |
| LDCM0424 | CL35 | HEK-293T | C597(1.13) | LDD1022 | [5] |
| LDCM0425 | CL36 | HEK-293T | C597(0.69) | LDD1023 | [5] |
| LDCM0426 | CL37 | HEK-293T | C597(0.67) | LDD1024 | [5] |
| LDCM0428 | CL39 | HEK-293T | C597(0.49) | LDD1026 | [5] |
| LDCM0429 | CL4 | HEK-293T | C222(1.19) | LDD1027 | [5] |
| LDCM0430 | CL40 | HEK-293T | C597(0.97) | LDD1028 | [5] |
| LDCM0431 | CL41 | HEK-293T | C597(0.64) | LDD1029 | [5] |
| LDCM0432 | CL42 | HEK-293T | C597(0.68) | LDD1030 | [5] |
| LDCM0433 | CL43 | HEK-293T | C597(0.64) | LDD1031 | [5] |
| LDCM0434 | CL44 | HEK-293T | C597(0.93) | LDD1032 | [5] |
| LDCM0435 | CL45 | HEK-293T | C597(0.70) | LDD1033 | [5] |
| LDCM0436 | CL46 | HCT 116 | C222(1.11) | LDD0753 | [5] |
| LDCM0437 | CL47 | HCT 116 | C222(0.76) | LDD0754 | [5] |
| LDCM0438 | CL48 | HCT 116 | C222(0.85) | LDD0755 | [5] |
| LDCM0439 | CL49 | HCT 116 | C222(1.03) | LDD0756 | [5] |
| LDCM0440 | CL5 | HEK-293T | C222(0.96) | LDD1038 | [5] |
| LDCM0441 | CL50 | HCT 116 | C222(0.84) | LDD0758 | [5] |
| LDCM0442 | CL51 | HCT 116 | C222(1.01) | LDD0759 | [5] |
| LDCM0443 | CL52 | HCT 116 | C222(1.04) | LDD0760 | [5] |
| LDCM0444 | CL53 | HCT 116 | C222(1.25) | LDD0761 | [5] |
| LDCM0445 | CL54 | HCT 116 | C222(0.98) | LDD0762 | [5] |
| LDCM0446 | CL55 | HCT 116 | C222(0.83) | LDD0763 | [5] |
| LDCM0447 | CL56 | HCT 116 | C222(0.78) | LDD0764 | [5] |
| LDCM0448 | CL57 | HCT 116 | C222(0.89) | LDD0765 | [5] |
| LDCM0449 | CL58 | HCT 116 | C222(0.91) | LDD0766 | [5] |
| LDCM0450 | CL59 | HCT 116 | C222(1.10) | LDD0767 | [5] |
| LDCM0451 | CL6 | HEK-293T | C222(0.90) | LDD1049 | [5] |
| LDCM0452 | CL60 | HCT 116 | C222(0.94) | LDD0769 | [5] |
| LDCM0453 | CL61 | HCT 116 | C222(1.01) | LDD0770 | [5] |
| LDCM0454 | CL62 | HCT 116 | C222(1.06) | LDD0771 | [5] |
| LDCM0455 | CL63 | HCT 116 | C222(0.85) | LDD0772 | [5] |
| LDCM0456 | CL64 | HCT 116 | C222(0.52) | LDD0773 | [5] |
| LDCM0457 | CL65 | HCT 116 | C222(0.72) | LDD0774 | [5] |
| LDCM0458 | CL66 | HCT 116 | C222(0.77) | LDD0775 | [5] |
| LDCM0459 | CL67 | HCT 116 | C222(0.67) | LDD0776 | [5] |
| LDCM0460 | CL68 | HCT 116 | C222(0.82) | LDD0777 | [5] |
| LDCM0461 | CL69 | HCT 116 | C222(0.83) | LDD0778 | [5] |
| LDCM0462 | CL7 | HEK-293T | C222(0.93) | LDD1060 | [5] |
| LDCM0463 | CL70 | HCT 116 | C222(0.90) | LDD0780 | [5] |
| LDCM0464 | CL71 | HCT 116 | C222(0.67) | LDD0781 | [5] |
| LDCM0465 | CL72 | HCT 116 | C222(1.08) | LDD0782 | [5] |
| LDCM0466 | CL73 | HCT 116 | C222(0.71) | LDD0783 | [5] |
| LDCM0467 | CL74 | HCT 116 | C222(0.91) | LDD0784 | [5] |
| LDCM0469 | CL76 | HCT 116 | C222(1.00) | LDD0786 | [5] |
| LDCM0470 | CL77 | HCT 116 | C222(0.89) | LDD0787 | [5] |
| LDCM0471 | CL78 | HCT 116 | C222(0.96) | LDD0788 | [5] |
| LDCM0472 | CL79 | HCT 116 | C222(0.86) | LDD0789 | [5] |
| LDCM0473 | CL8 | HEK-293T | C222(1.71) | LDD1071 | [5] |
| LDCM0474 | CL80 | HCT 116 | C222(0.83) | LDD0791 | [5] |
| LDCM0475 | CL81 | HCT 116 | C222(0.88) | LDD0792 | [5] |
| LDCM0476 | CL82 | HCT 116 | C222(0.76) | LDD0793 | [5] |
| LDCM0477 | CL83 | HCT 116 | C222(0.90) | LDD0794 | [5] |
| LDCM0478 | CL84 | HCT 116 | C222(0.92) | LDD0795 | [5] |
| LDCM0479 | CL85 | HCT 116 | C222(0.91) | LDD0796 | [5] |
| LDCM0480 | CL86 | HCT 116 | C222(0.98) | LDD0797 | [5] |
| LDCM0481 | CL87 | HCT 116 | C222(0.84) | LDD0798 | [5] |
| LDCM0482 | CL88 | HCT 116 | C222(0.81) | LDD0799 | [5] |
| LDCM0483 | CL89 | HCT 116 | C222(0.98) | LDD0800 | [5] |
| LDCM0484 | CL9 | HEK-293T | C222(0.75) | LDD1082 | [5] |
| LDCM0485 | CL90 | HCT 116 | C222(0.87) | LDD0802 | [5] |
| LDCM0486 | CL91 | HCT 116 | C222(1.18) | LDD0803 | [5] |
| LDCM0487 | CL92 | HCT 116 | C222(1.05) | LDD0804 | [5] |
| LDCM0488 | CL93 | HCT 116 | C222(1.08) | LDD0805 | [5] |
| LDCM0489 | CL94 | HCT 116 | C222(1.41) | LDD0806 | [5] |
| LDCM0490 | CL95 | HCT 116 | C222(1.45) | LDD0807 | [5] |
| LDCM0491 | CL96 | HCT 116 | C222(1.05) | LDD0808 | [5] |
| LDCM0492 | CL97 | HCT 116 | C222(1.15) | LDD0809 | [5] |
| LDCM0493 | CL98 | HCT 116 | C222(1.22) | LDD0810 | [5] |
| LDCM0494 | CL99 | HCT 116 | C222(1.09) | LDD0811 | [5] |
| LDCM0027 | Dopamine | HEK-293T | 19.45 | LDD0179 | [2] |
| LDCM0495 | E2913 | HEK-293T | C148(1.15); C222(1.05); C374(1.29); C305(1.06) | LDD1698 | [15] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C222(1.22); C374(1.12); C431(1.06); C305(0.73) | LDD1702 | [3] |
| LDCM0031 | Epigallocatechin gallate | HEK-293T | 25.59 | LDD0183 | [2] |
| LDCM0468 | Fragment33 | HCT 116 | C222(0.71) | LDD0785 | [5] |
| LDCM0427 | Fragment51 | HEK-293T | C597(1.17) | LDD1025 | [5] |
| LDCM0022 | KB02 | HCT 116 | C305(2.34) | LDD0080 | [5] |
| LDCM0023 | KB03 | HCT 116 | C305(1.42) | LDD0081 | [5] |
| LDCM0024 | KB05 | HCT 116 | C305(1.74) | LDD0082 | [5] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C431(0.55) | LDD2121 | [3] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C305(1.14) | LDD2090 | [3] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C305(1.24) | LDD2096 | [3] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C374(1.05); C560(1.10) | LDD2099 | [3] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C431(1.12); C374(0.66) | LDD2107 | [3] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C374(0.98) | LDD2111 | [3] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C305(0.91) | LDD2118 | [3] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C431(0.92) | LDD2120 | [3] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C560(1.03) | LDD2123 | [3] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C305(1.21) | LDD2126 | [3] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C560(1.00) | LDD2137 | [3] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C374(0.74) | LDD2140 | [3] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C305(0.89) | LDD2143 | [3] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C431(0.41) | LDD2150 | [3] |
| LDCM0032 | Oleacein | HEK-293T | 5.76 | LDD0184 | [2] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C305(1.04) | LDD2207 | [16] |
| LDCM0029 | Quercetin | HEK-293T | 6.24 | LDD0181 | [2] |
| LDCM0131 | RA190 | MM1.R | C305(2.65); C374(1.87) | LDD0304 | [17] |
| LDCM0021 | THZ1 | HCT 116 | C305(1.07) | LDD2173 | [5] |
The Interaction Atlas With This Target
References
