Details of the Target
General Information of Target
| Target ID | LDTP09578 | |||||
|---|---|---|---|---|---|---|
| Target Name | Phosphatidylglycerophosphatase and protein-tyrosine phosphatase 1 (PTPMT1) | |||||
| Gene Name | PTPMT1 | |||||
| Gene ID | 114971 | |||||
| Synonyms |
MOSP; PLIP; Phosphatidylglycerophosphatase and protein-tyrosine phosphatase 1; EC 3.1.3.27; PTEN-like phosphatase; Phosphoinositide lipid phosphatase; Protein-tyrosine phosphatase mitochondrial 1; EC 3.1.3.16, EC 3.1.3.48
|
|||||
| 3D Structure | ||||||
| Sequence |
MAATALLEAGLARVLFYPTLLYTLFRGKVPGRAHRDWYHRIDPTVLLGALPLRSLTRQLV
QDENVRGVITMNEEYETRFLCNSSQEWKRLGVEQLRLSTVDMTGIPTLDNLQKGVQFALK YQSLGQCVYVHCKAGRSRSATMVAAYLIQVHKWSPEEAVRAIAKIRSYIHIRPGQLDVLK EFHKQITARATKDGTFVISKT |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Protein-tyrosine phosphatase family, Non-receptor class dual specificity subfamily
|
|||||
| Subcellular location |
Mitochondrion inner membrane
|
|||||
| Function |
Lipid phosphatase which dephosphorylates phosphatidylglycerophosphate (PGP) to phosphatidylglycerol (PG). PGP is an essential intermediate in the biosynthetic pathway of cardiolipin, a mitochondrial-specific phospholipid regulating the membrane integrity and activities of the organelle. Has also been shown to display phosphatase activity toward phosphoprotein substrates, specifically mediates dephosphorylation of mitochondrial proteins, thereby playing an essential role in ATP production. Has probably a preference for proteins phosphorylated on Ser and/or Thr residues compared to proteins phosphorylated on Tyr residues. Probably involved in regulation of insulin secretion in pancreatic beta cells. May prevent intrinsic apoptosis, probably by regulating mitochondrial membrane integrity.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
HDSF-alk Probe Info |
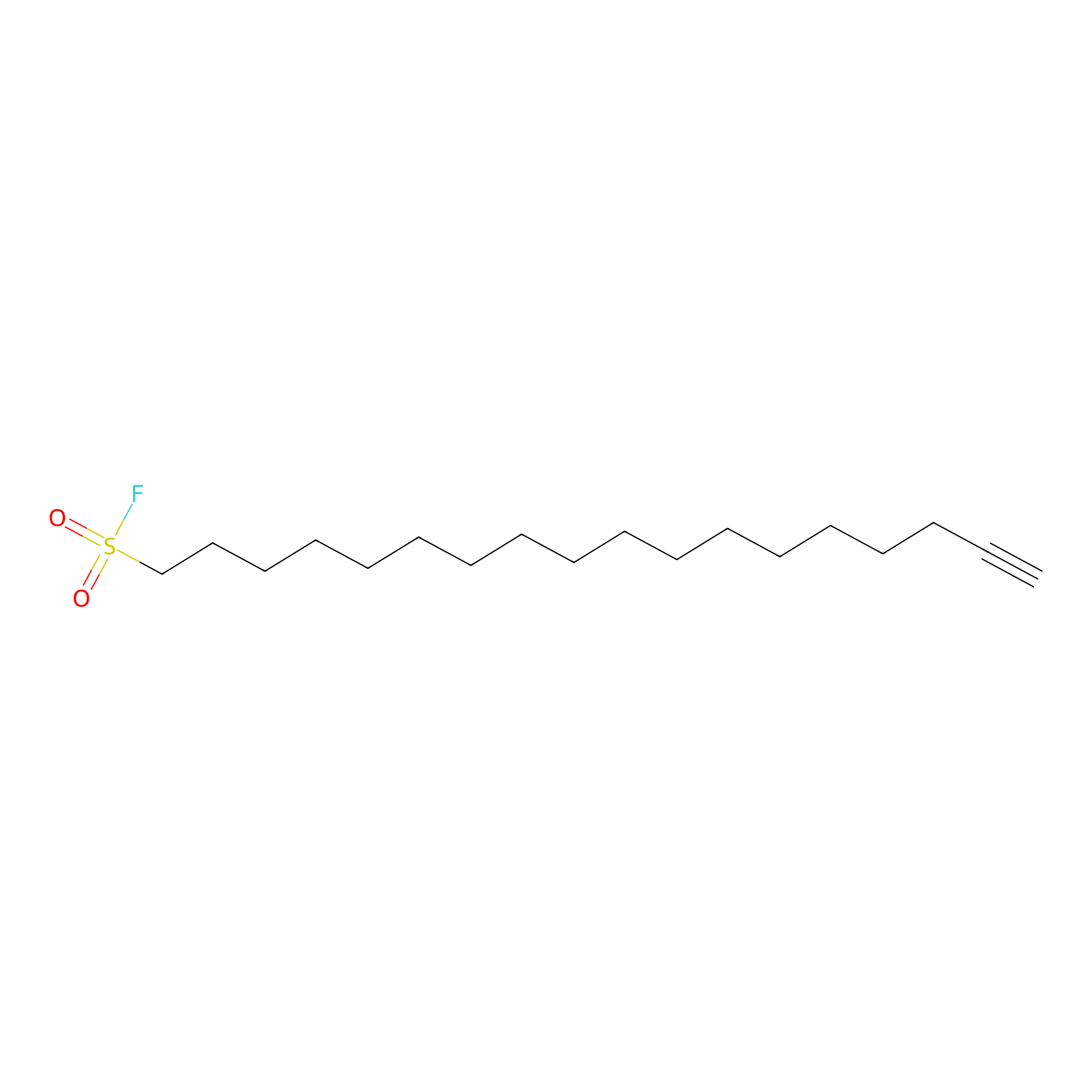 |
2.44 | LDD0197 | [2] | |
|
STPyne Probe Info |
 |
K28(3.28) | LDD0277 | [3] | |
|
IPM Probe Info |
 |
N.A. | LDD0241 | [4] | |
|
DBIA Probe Info |
 |
C81(1.77) | LDD3310 | [5] | |
|
IA-alkyne Probe Info |
 |
C81(1.37) | LDD2206 | [6] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [7] | |
|
AOyne Probe Info |
 |
14.10 | LDD0443 | [8] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [9] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [10] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C056 Probe Info |
 |
16.68 | LDD1753 | [11] | |
|
C091 Probe Info |
 |
9.99 | LDD1782 | [11] | |
|
C220 Probe Info |
 |
12.47 | LDD1894 | [11] | |
|
C228 Probe Info |
 |
17.63 | LDD1901 | [11] | |
|
C235 Probe Info |
 |
17.51 | LDD1908 | [11] | |
|
C285 Probe Info |
 |
21.11 | LDD1955 | [11] | |
|
C287 Probe Info |
 |
8.94 | LDD1957 | [11] | |
|
C288 Probe Info |
 |
6.45 | LDD1958 | [11] | |
|
C289 Probe Info |
 |
35.02 | LDD1959 | [11] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C81(0.99) | LDD1507 | [12] |
| LDCM0215 | AC10 | HEK-293T | C81(0.99); C127(0.96) | LDD1508 | [12] |
| LDCM0226 | AC11 | HEK-293T | C81(0.93) | LDD1509 | [12] |
| LDCM0237 | AC12 | HEK-293T | C81(0.90) | LDD1510 | [12] |
| LDCM0246 | AC128 | PaTu 8988t | C81(1.00) | LDD1125 | [13] |
| LDCM0247 | AC129 | PaTu 8988t | C81(0.95) | LDD1126 | [13] |
| LDCM0249 | AC130 | PaTu 8988t | C81(1.02) | LDD1128 | [13] |
| LDCM0250 | AC131 | PaTu 8988t | C81(0.98) | LDD1129 | [13] |
| LDCM0251 | AC132 | PaTu 8988t | C81(0.90) | LDD1130 | [13] |
| LDCM0252 | AC133 | PaTu 8988t | C81(0.90) | LDD1131 | [13] |
| LDCM0253 | AC134 | PaTu 8988t | C81(0.86) | LDD1132 | [13] |
| LDCM0254 | AC135 | PaTu 8988t | C81(0.91) | LDD1133 | [13] |
| LDCM0255 | AC136 | PaTu 8988t | C81(1.05) | LDD1134 | [13] |
| LDCM0256 | AC137 | PaTu 8988t | C81(0.96) | LDD1135 | [13] |
| LDCM0257 | AC138 | PaTu 8988t | C81(0.88) | LDD1136 | [13] |
| LDCM0258 | AC139 | PaTu 8988t | C81(1.12) | LDD1137 | [13] |
| LDCM0259 | AC14 | HEK-293T | C81(0.93); C127(0.91) | LDD1512 | [12] |
| LDCM0260 | AC140 | PaTu 8988t | C81(0.92) | LDD1139 | [13] |
| LDCM0261 | AC141 | PaTu 8988t | C81(1.04) | LDD1140 | [13] |
| LDCM0262 | AC142 | PaTu 8988t | C81(0.94) | LDD1141 | [13] |
| LDCM0263 | AC143 | PaTu 8988t | C81(0.85) | LDD1142 | [13] |
| LDCM0264 | AC144 | PaTu 8988t | C81(1.12) | LDD1143 | [13] |
| LDCM0265 | AC145 | PaTu 8988t | C81(1.03) | LDD1144 | [13] |
| LDCM0266 | AC146 | PaTu 8988t | C81(1.00) | LDD1145 | [13] |
| LDCM0267 | AC147 | PaTu 8988t | C81(0.99) | LDD1146 | [13] |
| LDCM0268 | AC148 | PaTu 8988t | C81(0.82) | LDD1147 | [13] |
| LDCM0269 | AC149 | PaTu 8988t | C81(0.96) | LDD1148 | [13] |
| LDCM0270 | AC15 | HEK-293T | C81(0.89) | LDD1513 | [12] |
| LDCM0271 | AC150 | PaTu 8988t | C81(1.24) | LDD1150 | [13] |
| LDCM0272 | AC151 | PaTu 8988t | C81(0.88) | LDD1151 | [13] |
| LDCM0273 | AC152 | PaTu 8988t | C81(0.92) | LDD1152 | [13] |
| LDCM0274 | AC153 | PaTu 8988t | C81(0.97) | LDD1153 | [13] |
| LDCM0621 | AC154 | PaTu 8988t | C81(0.98) | LDD2166 | [13] |
| LDCM0622 | AC155 | PaTu 8988t | C81(1.04) | LDD2167 | [13] |
| LDCM0623 | AC156 | PaTu 8988t | C81(0.97) | LDD2168 | [13] |
| LDCM0624 | AC157 | PaTu 8988t | C81(1.29) | LDD2169 | [13] |
| LDCM0276 | AC17 | HEK-293T | C81(0.97) | LDD1515 | [12] |
| LDCM0277 | AC18 | HEK-293T | C81(0.97); C127(0.90) | LDD1516 | [12] |
| LDCM0278 | AC19 | HEK-293T | C81(1.01) | LDD1517 | [12] |
| LDCM0279 | AC2 | HEK-293T | C81(1.01); C127(0.95) | LDD1518 | [12] |
| LDCM0280 | AC20 | HEK-293T | C81(0.91) | LDD1519 | [12] |
| LDCM0281 | AC21 | HEK-293T | C81(1.03); C127(0.77) | LDD1520 | [12] |
| LDCM0282 | AC22 | HEK-293T | C81(0.94); C127(0.94) | LDD1521 | [12] |
| LDCM0283 | AC23 | HEK-293T | C81(0.97) | LDD1522 | [12] |
| LDCM0284 | AC24 | HEK-293T | C81(0.94) | LDD1523 | [12] |
| LDCM0285 | AC25 | HEK-293T | C81(0.97) | LDD1524 | [12] |
| LDCM0286 | AC26 | HEK-293T | C81(1.01); C127(0.86) | LDD1525 | [12] |
| LDCM0287 | AC27 | HEK-293T | C81(0.96) | LDD1526 | [12] |
| LDCM0288 | AC28 | HEK-293T | C81(0.95) | LDD1527 | [12] |
| LDCM0289 | AC29 | HEK-293T | C81(0.97); C127(0.86) | LDD1528 | [12] |
| LDCM0290 | AC3 | HEK-293T | C81(1.00) | LDD1529 | [12] |
| LDCM0291 | AC30 | HEK-293T | C81(0.92); C127(0.96) | LDD1530 | [12] |
| LDCM0292 | AC31 | HEK-293T | C81(0.94) | LDD1531 | [12] |
| LDCM0293 | AC32 | HEK-293T | C81(1.01) | LDD1532 | [12] |
| LDCM0294 | AC33 | HEK-293T | C81(0.94) | LDD1533 | [12] |
| LDCM0295 | AC34 | HEK-293T | C81(0.92); C127(1.02) | LDD1534 | [12] |
| LDCM0296 | AC35 | HEK-293T | C81(1.00) | LDD1535 | [12] |
| LDCM0297 | AC36 | HEK-293T | C81(0.89) | LDD1536 | [12] |
| LDCM0298 | AC37 | HEK-293T | C81(0.95); C127(0.93) | LDD1537 | [12] |
| LDCM0299 | AC38 | HEK-293T | C81(0.84); C127(0.97) | LDD1538 | [12] |
| LDCM0300 | AC39 | HEK-293T | C81(0.94) | LDD1539 | [12] |
| LDCM0301 | AC4 | HEK-293T | C81(0.94) | LDD1540 | [12] |
| LDCM0302 | AC40 | HEK-293T | C81(0.97) | LDD1541 | [12] |
| LDCM0303 | AC41 | HEK-293T | C81(0.92) | LDD1542 | [12] |
| LDCM0304 | AC42 | HEK-293T | C81(1.01); C127(0.94) | LDD1543 | [12] |
| LDCM0305 | AC43 | HEK-293T | C81(0.93) | LDD1544 | [12] |
| LDCM0306 | AC44 | HEK-293T | C81(0.88) | LDD1545 | [12] |
| LDCM0307 | AC45 | HEK-293T | C81(0.96); C127(0.86) | LDD1546 | [12] |
| LDCM0308 | AC46 | PaTu 8988t | C81(1.07) | LDD1187 | [13] |
| LDCM0309 | AC47 | PaTu 8988t | C81(0.86) | LDD1188 | [13] |
| LDCM0310 | AC48 | PaTu 8988t | C81(0.66) | LDD1189 | [13] |
| LDCM0311 | AC49 | PaTu 8988t | C81(0.78) | LDD1190 | [13] |
| LDCM0312 | AC5 | HEK-293T | C81(0.98); C127(0.80) | LDD1551 | [12] |
| LDCM0313 | AC50 | PaTu 8988t | C81(0.85) | LDD1192 | [13] |
| LDCM0314 | AC51 | PaTu 8988t | C81(0.57) | LDD1193 | [13] |
| LDCM0315 | AC52 | PaTu 8988t | C81(0.63) | LDD1194 | [13] |
| LDCM0316 | AC53 | PaTu 8988t | C81(0.59) | LDD1195 | [13] |
| LDCM0317 | AC54 | PaTu 8988t | C81(0.63) | LDD1196 | [13] |
| LDCM0318 | AC55 | PaTu 8988t | C81(1.04) | LDD1197 | [13] |
| LDCM0319 | AC56 | PaTu 8988t | C81(0.70) | LDD1198 | [13] |
| LDCM0320 | AC57 | PaTu 8988t | C81(0.87) | LDD1199 | [13] |
| LDCM0321 | AC58 | PaTu 8988t | C81(0.84) | LDD1200 | [13] |
| LDCM0322 | AC59 | PaTu 8988t | C81(1.06) | LDD1201 | [13] |
| LDCM0323 | AC6 | HEK-293T | C81(0.96); C127(1.04) | LDD1562 | [12] |
| LDCM0324 | AC60 | PaTu 8988t | C81(0.85) | LDD1203 | [13] |
| LDCM0325 | AC61 | PaTu 8988t | C81(1.09) | LDD1204 | [13] |
| LDCM0326 | AC62 | PaTu 8988t | C81(1.12) | LDD1205 | [13] |
| LDCM0327 | AC63 | PaTu 8988t | C81(1.00) | LDD1206 | [13] |
| LDCM0328 | AC64 | PaTu 8988t | C81(1.31) | LDD1207 | [13] |
| LDCM0329 | AC65 | PaTu 8988t | C81(1.15) | LDD1208 | [13] |
| LDCM0330 | AC66 | PaTu 8988t | C81(1.06) | LDD1209 | [13] |
| LDCM0331 | AC67 | PaTu 8988t | C81(1.19) | LDD1210 | [13] |
| LDCM0334 | AC7 | HEK-293T | C81(0.97) | LDD1568 | [12] |
| LDCM0345 | AC8 | HEK-293T | C81(0.96) | LDD1569 | [12] |
| LDCM0248 | AKOS034007472 | HEK-293T | C81(0.98); C127(0.84) | LDD1511 | [12] |
| LDCM0356 | AKOS034007680 | HEK-293T | C81(0.94) | LDD1570 | [12] |
| LDCM0275 | AKOS034007705 | HEK-293T | C81(0.94) | LDD1514 | [12] |
| LDCM0156 | Aniline | NCI-H1299 | 15.00 | LDD0403 | [1] |
| LDCM0367 | CL1 | HEK-293T | C81(1.02) | LDD1571 | [12] |
| LDCM0368 | CL10 | HEK-293T | C81(1.78); C127(1.39) | LDD1572 | [12] |
| LDCM0369 | CL100 | HEK-293T | C81(1.01); C127(1.06) | LDD1573 | [12] |
| LDCM0370 | CL101 | HEK-293T | C81(0.95) | LDD1574 | [12] |
| LDCM0371 | CL102 | HEK-293T | C81(1.12); C127(1.07) | LDD1575 | [12] |
| LDCM0372 | CL103 | HEK-293T | C81(0.93); C127(0.84) | LDD1576 | [12] |
| LDCM0373 | CL104 | HEK-293T | C81(0.96); C127(0.99) | LDD1577 | [12] |
| LDCM0374 | CL105 | HEK-293T | C81(0.98) | LDD1578 | [12] |
| LDCM0375 | CL106 | HEK-293T | C81(0.99); C127(0.88) | LDD1579 | [12] |
| LDCM0376 | CL107 | HEK-293T | C81(0.94); C127(0.77) | LDD1580 | [12] |
| LDCM0377 | CL108 | HEK-293T | C81(1.00); C127(0.92) | LDD1581 | [12] |
| LDCM0378 | CL109 | HEK-293T | C81(1.01) | LDD1582 | [12] |
| LDCM0379 | CL11 | HEK-293T | C81(1.15) | LDD1583 | [12] |
| LDCM0380 | CL110 | HEK-293T | C81(1.11); C127(0.93) | LDD1584 | [12] |
| LDCM0381 | CL111 | HEK-293T | C81(0.99); C127(0.92) | LDD1585 | [12] |
| LDCM0382 | CL112 | HEK-293T | C81(1.01); C127(0.91) | LDD1586 | [12] |
| LDCM0383 | CL113 | HEK-293T | C81(0.93) | LDD1587 | [12] |
| LDCM0384 | CL114 | HEK-293T | C81(1.24); C127(1.08) | LDD1588 | [12] |
| LDCM0385 | CL115 | HEK-293T | C81(0.89); C127(0.94) | LDD1589 | [12] |
| LDCM0386 | CL116 | HEK-293T | C81(0.99); C127(1.05) | LDD1590 | [12] |
| LDCM0387 | CL117 | HEK-293T | C81(0.90) | LDD1591 | [12] |
| LDCM0388 | CL118 | HEK-293T | C81(0.90); C127(0.89) | LDD1592 | [12] |
| LDCM0389 | CL119 | HEK-293T | C81(0.95); C127(0.83) | LDD1593 | [12] |
| LDCM0390 | CL12 | HEK-293T | C81(1.02) | LDD1594 | [12] |
| LDCM0391 | CL120 | HEK-293T | C81(0.93); C127(0.99) | LDD1595 | [12] |
| LDCM0392 | CL121 | PaTu 8988t | C81(0.99) | LDD1271 | [13] |
| LDCM0393 | CL122 | PaTu 8988t | C81(0.70) | LDD1272 | [13] |
| LDCM0394 | CL123 | PaTu 8988t | C81(0.76) | LDD1273 | [13] |
| LDCM0395 | CL124 | PaTu 8988t | C81(1.07) | LDD1274 | [13] |
| LDCM0396 | CL125 | PaTu 8988t | C81(1.00) | LDD1275 | [13] |
| LDCM0397 | CL126 | PaTu 8988t | C81(1.10) | LDD1276 | [13] |
| LDCM0398 | CL127 | PaTu 8988t | C81(1.08) | LDD1277 | [13] |
| LDCM0399 | CL128 | PaTu 8988t | C81(0.99) | LDD1278 | [13] |
| LDCM0400 | CL13 | HEK-293T | C81(1.01) | LDD1604 | [12] |
| LDCM0401 | CL14 | HEK-293T | C81(1.01); C127(0.88) | LDD1605 | [12] |
| LDCM0402 | CL15 | HEK-293T | C81(1.15); C127(1.17) | LDD1606 | [12] |
| LDCM0403 | CL16 | HEK-293T | C81(0.98); C127(1.08) | LDD1607 | [12] |
| LDCM0404 | CL17 | HEK-293T | C81(1.26) | LDD1608 | [12] |
| LDCM0405 | CL18 | HEK-293T | C81(0.94); C127(1.02) | LDD1609 | [12] |
| LDCM0406 | CL19 | HEK-293T | C81(0.97) | LDD1610 | [12] |
| LDCM0407 | CL2 | HEK-293T | C81(1.02); C127(0.87) | LDD1611 | [12] |
| LDCM0408 | CL20 | HEK-293T | C81(0.97) | LDD1612 | [12] |
| LDCM0409 | CL21 | HEK-293T | C81(1.27); C127(1.11) | LDD1613 | [12] |
| LDCM0410 | CL22 | HEK-293T | C81(1.21); C127(1.09) | LDD1614 | [12] |
| LDCM0411 | CL23 | HEK-293T | C81(1.07) | LDD1615 | [12] |
| LDCM0412 | CL24 | HEK-293T | C81(0.96) | LDD1616 | [12] |
| LDCM0413 | CL25 | HEK-293T | C81(1.04) | LDD1617 | [12] |
| LDCM0414 | CL26 | HEK-293T | C81(1.11); C127(0.85) | LDD1618 | [12] |
| LDCM0415 | CL27 | HEK-293T | C81(0.91); C127(0.95) | LDD1619 | [12] |
| LDCM0416 | CL28 | HEK-293T | C81(1.06); C127(1.02) | LDD1620 | [12] |
| LDCM0417 | CL29 | HEK-293T | C81(0.88) | LDD1621 | [12] |
| LDCM0418 | CL3 | HEK-293T | C81(0.98); C127(0.94) | LDD1622 | [12] |
| LDCM0419 | CL30 | HEK-293T | C81(1.02); C127(1.00) | LDD1623 | [12] |
| LDCM0420 | CL31 | HEK-293T | C81(1.05) | LDD1624 | [12] |
| LDCM0421 | CL32 | HEK-293T | C81(0.97) | LDD1625 | [12] |
| LDCM0422 | CL33 | HEK-293T | C81(1.57); C127(1.05) | LDD1626 | [12] |
| LDCM0423 | CL34 | HEK-293T | C81(1.20); C127(1.13) | LDD1627 | [12] |
| LDCM0424 | CL35 | HEK-293T | C81(1.08) | LDD1628 | [12] |
| LDCM0425 | CL36 | HEK-293T | C81(0.98) | LDD1629 | [12] |
| LDCM0426 | CL37 | HEK-293T | C81(0.91) | LDD1630 | [12] |
| LDCM0428 | CL39 | HEK-293T | C81(1.00); C127(0.79) | LDD1632 | [12] |
| LDCM0429 | CL4 | HEK-293T | C81(1.09); C127(1.16) | LDD1633 | [12] |
| LDCM0430 | CL40 | HEK-293T | C81(1.01); C127(1.00) | LDD1634 | [12] |
| LDCM0431 | CL41 | HEK-293T | C81(1.06) | LDD1635 | [12] |
| LDCM0432 | CL42 | HEK-293T | C81(0.97); C127(1.12) | LDD1636 | [12] |
| LDCM0433 | CL43 | HEK-293T | C81(0.93) | LDD1637 | [12] |
| LDCM0434 | CL44 | HEK-293T | C81(1.01) | LDD1638 | [12] |
| LDCM0435 | CL45 | HEK-293T | C81(1.21); C127(1.15) | LDD1639 | [12] |
| LDCM0436 | CL46 | PaTu 8988t | C81(1.03) | LDD1315 | [13] |
| LDCM0437 | CL47 | PaTu 8988t | C81(1.15) | LDD1316 | [13] |
| LDCM0438 | CL48 | PaTu 8988t | C81(1.19) | LDD1317 | [13] |
| LDCM0439 | CL49 | PaTu 8988t | C81(1.30) | LDD1318 | [13] |
| LDCM0440 | CL5 | HEK-293T | C81(1.05) | LDD1644 | [12] |
| LDCM0441 | CL50 | PaTu 8988t | C81(1.11) | LDD1320 | [13] |
| LDCM0442 | CL51 | PaTu 8988t | C81(1.38) | LDD1321 | [13] |
| LDCM0443 | CL52 | PaTu 8988t | C81(1.42) | LDD1322 | [13] |
| LDCM0444 | CL53 | PaTu 8988t | C81(1.35) | LDD1323 | [13] |
| LDCM0445 | CL54 | PaTu 8988t | C81(1.59) | LDD1324 | [13] |
| LDCM0446 | CL55 | PaTu 8988t | C81(1.94) | LDD1325 | [13] |
| LDCM0447 | CL56 | PaTu 8988t | C81(0.35) | LDD1326 | [13] |
| LDCM0448 | CL57 | PaTu 8988t | C81(1.40) | LDD1327 | [13] |
| LDCM0449 | CL58 | PaTu 8988t | C81(1.48) | LDD1328 | [13] |
| LDCM0450 | CL59 | PaTu 8988t | C81(1.33) | LDD1329 | [13] |
| LDCM0451 | CL6 | HEK-293T | C81(1.26); C127(1.19) | LDD1654 | [12] |
| LDCM0452 | CL60 | PaTu 8988t | C81(1.40) | LDD1331 | [13] |
| LDCM0453 | CL61 | HEK-293T | C81(0.93) | LDD1656 | [12] |
| LDCM0454 | CL62 | HEK-293T | C81(0.97); C127(0.72) | LDD1657 | [12] |
| LDCM0455 | CL63 | HEK-293T | C81(0.88); C127(0.86) | LDD1658 | [12] |
| LDCM0456 | CL64 | HEK-293T | C81(1.13); C127(1.10) | LDD1659 | [12] |
| LDCM0457 | CL65 | HEK-293T | C81(0.97) | LDD1660 | [12] |
| LDCM0458 | CL66 | HEK-293T | C81(1.02); C127(0.95) | LDD1661 | [12] |
| LDCM0459 | CL67 | HEK-293T | C81(0.92) | LDD1662 | [12] |
| LDCM0460 | CL68 | HEK-293T | C81(1.04) | LDD1663 | [12] |
| LDCM0461 | CL69 | HEK-293T | C81(1.07); C127(0.84) | LDD1664 | [12] |
| LDCM0462 | CL7 | HEK-293T | C81(1.03) | LDD1665 | [12] |
| LDCM0463 | CL70 | HEK-293T | C81(1.08); C127(1.11) | LDD1666 | [12] |
| LDCM0464 | CL71 | HEK-293T | C81(1.07) | LDD1667 | [12] |
| LDCM0465 | CL72 | HEK-293T | C81(0.94) | LDD1668 | [12] |
| LDCM0466 | CL73 | HEK-293T | C81(0.99) | LDD1669 | [12] |
| LDCM0467 | CL74 | HEK-293T | C81(0.92); C127(0.92) | LDD1670 | [12] |
| LDCM0469 | CL76 | HEK-293T | C81(0.93); C127(0.97) | LDD1672 | [12] |
| LDCM0470 | CL77 | HEK-293T | C81(1.22) | LDD1673 | [12] |
| LDCM0471 | CL78 | HEK-293T | C81(0.99); C127(1.11) | LDD1674 | [12] |
| LDCM0472 | CL79 | HEK-293T | C81(0.89) | LDD1675 | [12] |
| LDCM0473 | CL8 | HEK-293T | C81(1.83) | LDD1676 | [12] |
| LDCM0474 | CL80 | HEK-293T | C81(0.94) | LDD1677 | [12] |
| LDCM0475 | CL81 | HEK-293T | C81(1.00); C127(0.94) | LDD1678 | [12] |
| LDCM0476 | CL82 | HEK-293T | C81(1.00); C127(1.13) | LDD1679 | [12] |
| LDCM0477 | CL83 | HEK-293T | C81(1.06) | LDD1680 | [12] |
| LDCM0478 | CL84 | HEK-293T | C81(1.16) | LDD1681 | [12] |
| LDCM0479 | CL85 | HEK-293T | C81(0.87) | LDD1682 | [12] |
| LDCM0480 | CL86 | HEK-293T | C81(0.99); C127(0.85) | LDD1683 | [12] |
| LDCM0481 | CL87 | HEK-293T | C81(0.77); C127(0.78) | LDD1684 | [12] |
| LDCM0482 | CL88 | HEK-293T | C81(0.96); C127(0.95) | LDD1685 | [12] |
| LDCM0483 | CL89 | HEK-293T | C81(0.90) | LDD1686 | [12] |
| LDCM0484 | CL9 | HEK-293T | C81(1.07); C127(1.03) | LDD1687 | [12] |
| LDCM0485 | CL90 | HEK-293T | C81(1.63); C127(1.19) | LDD1688 | [12] |
| LDCM0486 | CL91 | HEK-293T | C81(0.93) | LDD1689 | [12] |
| LDCM0487 | CL92 | HEK-293T | C81(1.02) | LDD1690 | [12] |
| LDCM0488 | CL93 | HEK-293T | C81(1.14); C127(1.06) | LDD1691 | [12] |
| LDCM0489 | CL94 | HEK-293T | C81(1.05); C127(1.06) | LDD1692 | [12] |
| LDCM0490 | CL95 | HEK-293T | C81(1.46) | LDD1693 | [12] |
| LDCM0491 | CL96 | HEK-293T | C81(1.17) | LDD1694 | [12] |
| LDCM0492 | CL97 | HEK-293T | C81(1.03) | LDD1695 | [12] |
| LDCM0493 | CL98 | HEK-293T | C81(0.95); C127(0.84) | LDD1696 | [12] |
| LDCM0494 | CL99 | HEK-293T | C81(0.93); C127(0.84) | LDD1697 | [12] |
| LDCM0495 | E2913 | HEK-293T | C81(0.92); C127(0.74) | LDD1698 | [12] |
| LDCM0468 | Fragment33 | HEK-293T | C81(0.91); C127(0.81) | LDD1671 | [12] |
| LDCM0427 | Fragment51 | HEK-293T | C81(0.93); C127(0.84) | LDD1631 | [12] |
| LDCM0022 | KB02 | HEK-293T | C81(0.91) | LDD1492 | [12] |
| LDCM0023 | KB03 | HEK-293T | C81(1.00) | LDD1497 | [12] |
| LDCM0024 | KB05 | COLO792 | C81(1.77) | LDD3310 | [5] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C81(1.37) | LDD2206 | [6] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C81(1.29) | LDD2207 | [6] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| E3 ubiquitin-protein ligase SIAH1 (SIAH1) | SINA (Seven in absentia) family | Q8IUQ4 | |||
| Zinc finger protein RFP (TRIM27) | TRIM/RBCC family | P14373 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| MyoD family inhibitor (MDFI) | MDFI family | Q99750 | |||
| Transmembrane protein 14B (TMEM14B) | TMEM14 family | Q9NUH8 | |||
Other
References
