Details of the Target
General Information of Target
| Target ID | LDTP06176 | |||||
|---|---|---|---|---|---|---|
| Target Name | Mediator of DNA damage checkpoint protein 1 (MDC1) | |||||
| Gene Name | MDC1 | |||||
| Gene ID | 9656 | |||||
| Synonyms |
KIAA0170; NFBD1; Mediator of DNA damage checkpoint protein 1; Nuclear factor with BRCT domains 1 |
|||||
| 3D Structure | ||||||
| Sequence |
MEDTQAIDWDVEEEEETEQSSESLRCNVEPVGRLHIFSGAHGPEKDFPLHLGKNVVGRMP
DCSVALPFPSISKQHAEIEILAWDKAPILRDCGSLNGTQILRPPKVLSPGVSHRLRDQEL ILFADLLCQYHRLDVSLPFVSRGPLTVEETPRVQGETQPQRLLLAEDSEEEVDFLSERRM VKKSRTTSSSVIVPESDEEGHSPVLGGLGPPFAFNLNSDTDVEEGQQPATEEASSAARRG ATVEAKQSEAEVVTEIQLEKDQPLVKERDNDTKVKRGAGNGVVPAGVILERSQPPGEDSD TDVDDDSRPPGRPAEVHLERAQPFGFIDSDTDAEEERIPATPVVIPMKKRKIFHGVGTRG PGAPGLAHLQESQAGSDTDVEEGKAPQAVPLEKSQASMVINSDTDDEEEVSAALTLAHLK ESQPAIWNRDAEEDMPQRVVLLQRSQTTTERDSDTDVEEEELPVENREAVLKDHTKIRAL VRAHSEKDQPPFGDSDDSVEADKSSPGIHLERSQASTTVDINTQVEKEVPPGSAIIHIKK HQVSVEGTNQTDVKAVGGPAKLLVVSLEEAWPLHGDCETDAEEGTSLTASVVADVRKSQL PAEGDAGAEWAAAVLKQERAHEVGAQGGPPVAQVEQDLPISRENLTDLVVDTDTLGESTQ PQREGAQVPTGREREQHVGGTKDSEDNYGDSEDLDLQATQCFLENQGLEAVQSMEDEPTQ AFMLTPPQELGPSHCSFQTTGTLDEPWEVLATQPFCLRESEDSETQPFDTHLEAYGPCLS PPRAIPGDQHPESPVHTEPMGIQGRGRQTVDKVMGIPKETAERVGPERGPLERETEKLLP ERQTDVTGEEELTKGKQDREQKQLLARDTQRQESDKNGESASPERDRESLKVEIETSEEI QEKQVQKQTLPSKAFEREVERPVANRECDPAELEEKVPKVILERDTQRGEPEGGSQDQKG QASSPTPEPGVGAGDLPGPTSAPVPSGSQSGGRGSPVSPRRHQKGLLNCKMPPAEKASRI RAAEKVSRGDQESPDACLPPTVPEAPAPPQKPLNSQSQKHLAPPPLLSPLLPSIKPTVRK TRQDGSQEAPEAPLSSELEPFHPKPKIRTRKSSRMTPFPATSAAPEPHPSTSTAQPVTPK PTSQATRSRTNRSSVKTPEPVVPTAPELQPSTSTDQPVTSEPTSQVTRGRKSRSSVKTPE TVVPTALELQPSTSTDRPVTSEPTSQATRGRKNRSSVKTPEPVVPTAPELQPSTSTDQPV TSEPTYQATRGRKNRSSVKTPEPVVPTAPELRPSTSTDRPVTPKPTSRTTRSRTNMSSVK TPETVVPTAPELQISTSTDQPVTPKPTSRTTRSRTNMSSVKNPESTVPIAPELPPSTSTE QPVTPEPTSRATRGRKNRSSGKTPETLVPTAPKLEPSTSTDQPVTPEPTSQATRGRTNRS SVKTPETVVPTAPELQPSTSTDQPVTPEPTSQATRGRTDRSSVKTPETVVPTAPELQASA STDQPVTSEPTSRTTRGRKNRSSVKTPETVVPAAPELQPSTSTDQPVTPEPTSRATRGRT NRSSVKTPESIVPIAPELQPSTSRNQLVTPEPTSRATRCRTNRSSVKTPEPVVPTAPEPH PTTSTDQPVTPKLTSRATRRKTNRSSVKTPKPVEPAASDLEPFTPTDQSVTPEAIAQGGQ SKTLRSSTVRAMPVPTTPEFQSPVTTDQPISPEPITQPSCIKRQRAAGNPGSLAAPIDHK PCSAPLEPKSQASRNQRWGAVRAAESLTAIPEPASPQLLETPIHASQIQKVEPAGRSRFT PELQPKASQSRKRSLATMDSPPHQKQPQRGEVSQKTVIIKEEEEDTAEKPGKEEDVVTPK PGKRKRDQAEEEPNRIPSRSLRRTKLNQESTAPKVLFTGVVDARGERAVLALGGSLAGSA AEASHLVTDRIRRTVKFLCALGRGIPILSLDWLHQSRKAGFFLPPDEYVVTDPEQEKNFG FSLQDALSRARERRLLEGYEIYVTPGVQPPPPQMGEIISCCGGTYLPSMPRSYKPQRVVI TCPQDFPHCSIPLRVGLPLLSPEFLLTGVLKQEAKPEAFVLSPLEMSST |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Histone reader protein required for checkpoint-mediated cell cycle arrest in response to DNA damage within both the S phase and G2/M phases of the cell cycle. Specifically recognizes and binds histone H2AX phosphorylated at 'Ser-139', a marker of DNA damage, serving as a scaffold for the recruitment of DNA repair and signal transduction proteins to discrete foci of DNA damage sites. Also required for downstream events subsequent to the recruitment of these proteins. These include phosphorylation and activation of the ATM, CHEK1 and CHEK2 kinases, and stabilization of TP53/p53 and apoptosis. ATM and CHEK2 may also be activated independently by a parallel pathway mediated by TP53BP1. Required for chromosomal stability during mitosis by promoting recruitment of TOPBP1 to DNA double strand breaks (DSBs): TOPBP1 forms filamentous assemblies that bridge MDC1 and tether broken chromosomes during mitosis.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| AN3CA | SNV: p.I2051T | DBIA Probe Info | |||
| CAL148 | SNV: p.D502N | DBIA Probe Info | |||
| CAMA1 | SNV: p.S1583C | DBIA Probe Info | |||
| CCK81 | Deletion: p.E468del | DBIA Probe Info | |||
| CMK | SNV: p.R161G | DBIA Probe Info | |||
| DETROIT562 | Deletion: p.N644PfsTer41 | DBIA Probe Info | |||
| DOTC24510 | SNV: p.Q1185H | DBIA Probe Info | |||
| HCC44 | SNV: p.S763F | DBIA Probe Info | |||
| HSC3 | SNV: p.R308M | DBIA Probe Info | |||
| HT115 | SNV: p.S1594Y | DBIA Probe Info | |||
| IGR1 | Insertion: p.S1915KfsTer15 | DBIA Probe Info | |||
| IGROV1 | SNV: p.T1109A | DBIA Probe Info | |||
| Ishikawa (Heraklio) 02 ER | SNV: p.V31A | DBIA Probe Info | |||
| JHH7 | SNV: p.G42E | . | |||
| JURKAT | Insertion: p.G680WfsTer6 SNV: p.E841K;p.M1692I;p.A1794V |
Compound 10 Probe Info | |||
| KMCH1 | SNV: p.K183Ter; p.Q843R; p.T1589N | DBIA Probe Info | |||
| MDST8 | SNV: p.P1052S | DBIA Probe Info | |||
| MFE319 | Deletion: p.N877MfsTer14 | DBIA Probe Info | |||
| MOLT4 | SNV: p.S23F; p.S402N; p.D2045N | IA-alkyne Probe Info | |||
| P31FUJ | SNV: p.E2D | DBIA Probe Info | |||
| SAOS2 | SNV: p.R359G | DBIA Probe Info | |||
| SHP77 | SNV: p.E14Ter | DBIA Probe Info | |||
| SKMEL30 | SNV: p.P1105R | DBIA Probe Info | |||
| TE4 | SNV: p.T1183I | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
13.51 | LDD0402 | [1] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [2] | |
|
STPyne Probe Info |
 |
K476(1.35) | LDD0277 | [3] | |
|
BTD Probe Info |
 |
C1939(0.54) | LDD2109 | [4] | |
|
MCL-4 Probe Info |
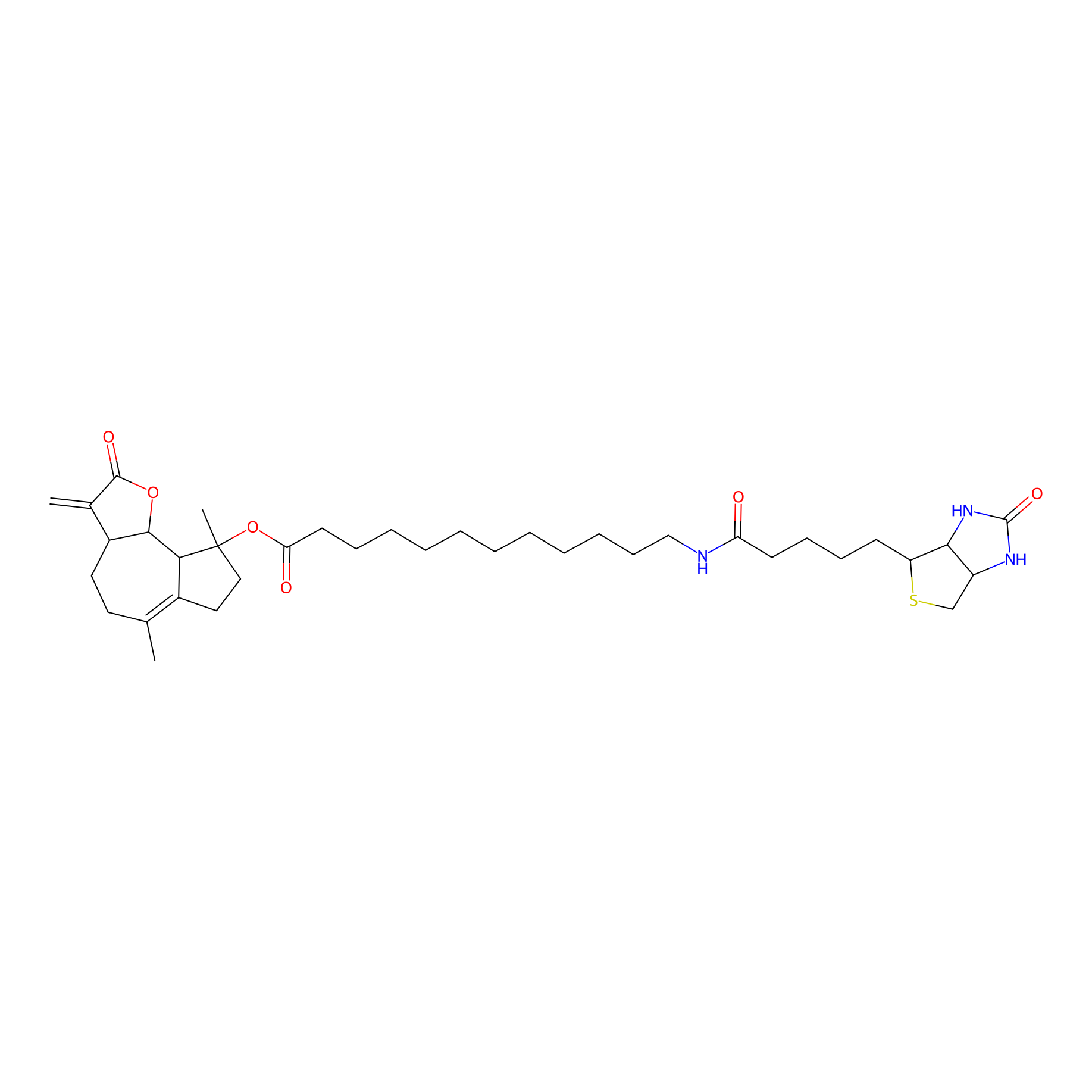 |
1.50 | LDD0049 | [5] | |
|
AHL-Pu-1 Probe Info |
 |
C928(2.11); C26(3.11); C1742(2.15); C1037(2.20) | LDD0169 | [6] | |
|
DBIA Probe Info |
 |
C26(1.03) | LDD0078 | [7] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C26(0.00); C92(0.00); C128(0.00) | LDD0038 | [8] | |
|
IA-alkyne Probe Info |
 |
C1742(0.00); C2042(0.00) | LDD0032 | [9] | |
|
Lodoacetamide azide Probe Info |
 |
C26(0.00); C128(0.00) | LDD0037 | [8] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [10] | |
|
TFBX Probe Info |
 |
C26(0.00); C1742(0.00) | LDD0027 | [10] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [11] | |
|
aHNE Probe Info |
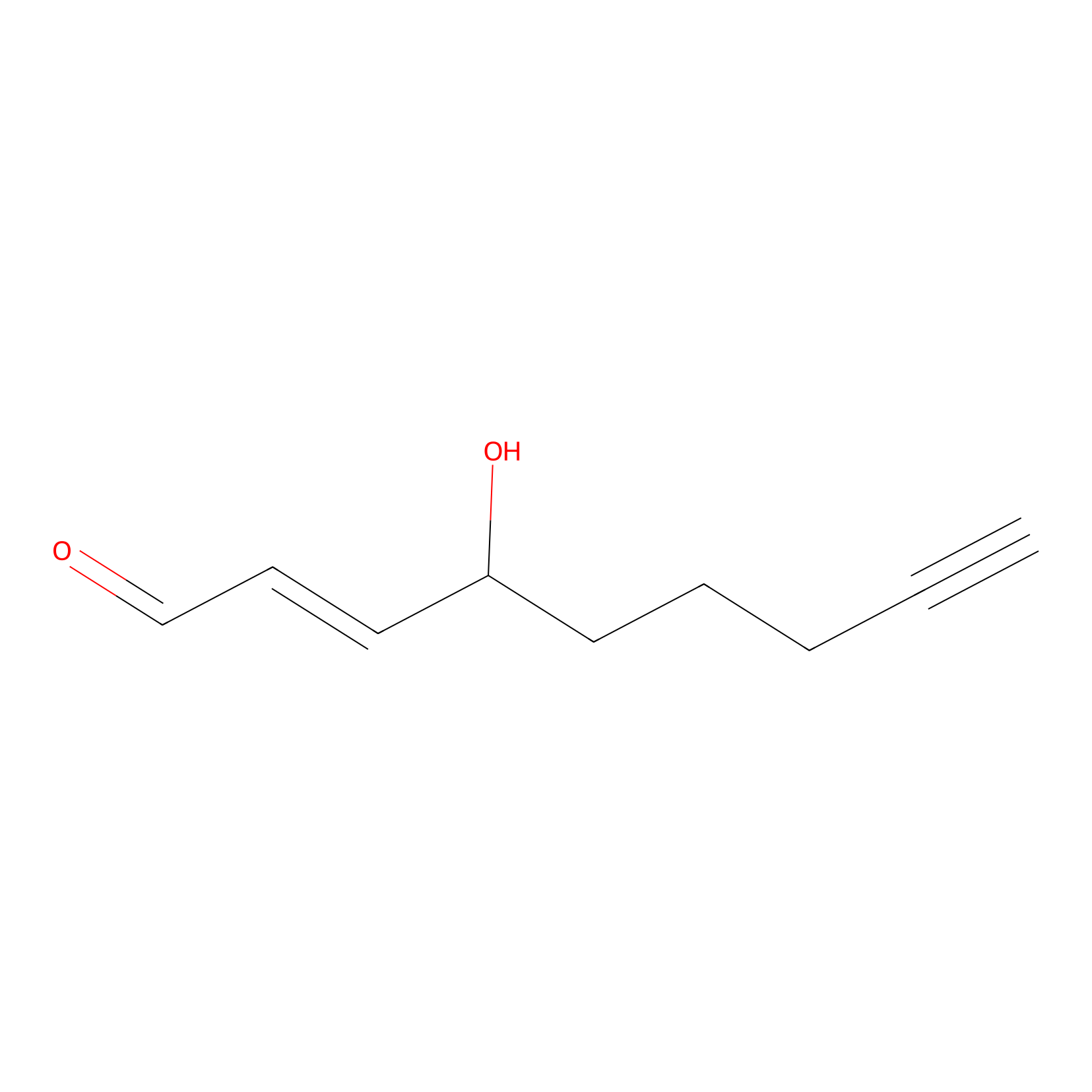 |
N.A. | LDD0001 | [11] | |
|
Compound 10 Probe Info |
 |
C1742(0.00); C26(0.00) | LDD2216 | [12] | |
|
Compound 11 Probe Info |
 |
C1742(0.00); C26(0.00) | LDD2213 | [12] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [11] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [11] | |
|
NPM Probe Info |
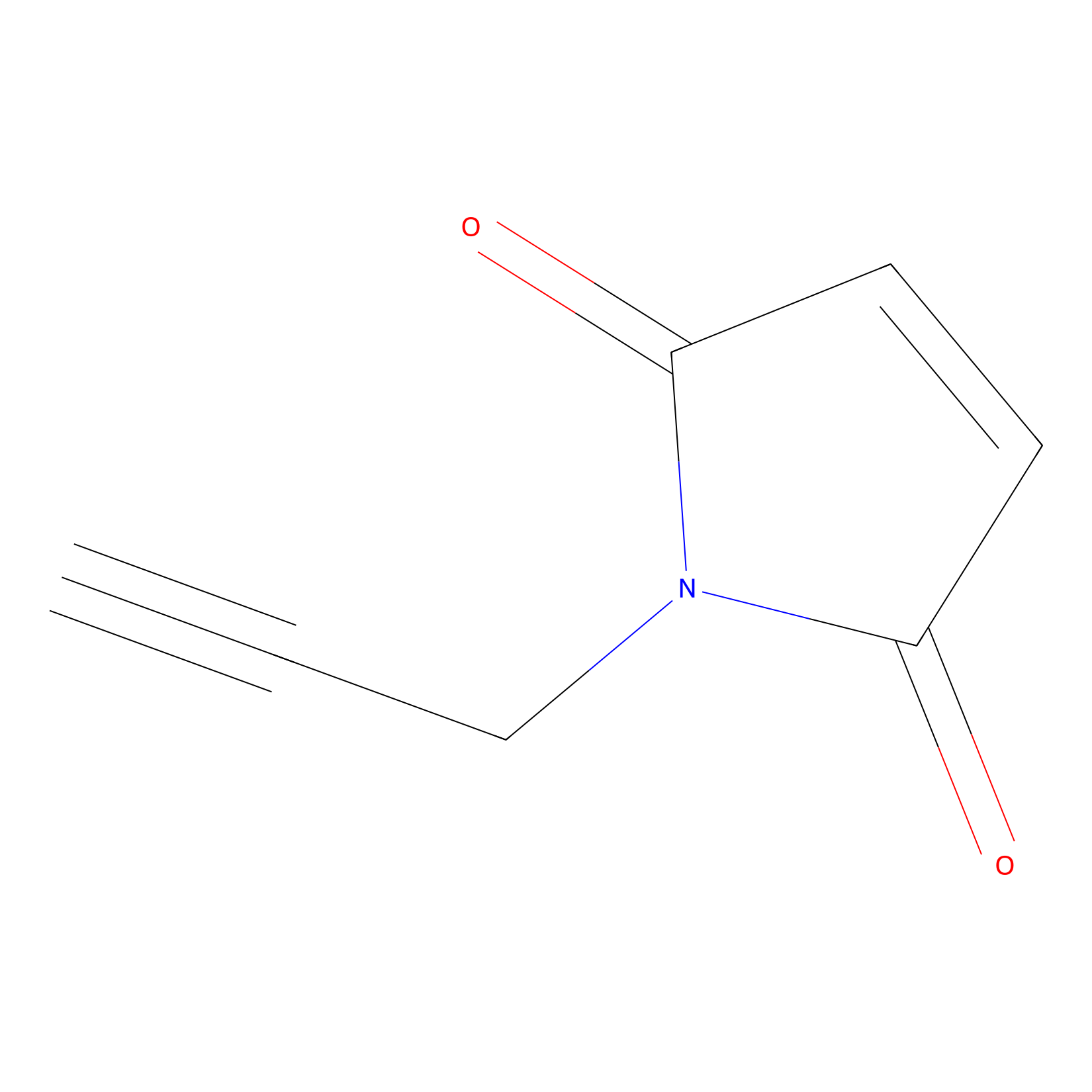 |
N.A. | LDD0016 | [11] | |
|
VSF Probe Info |
 |
C26(0.00); C1720(0.00); C1037(0.00); C1742(0.00) | LDD0007 | [11] | |
|
Phosphinate-6 Probe Info |
 |
C1009(0.00); C1742(0.00); C1037(0.00); C26(0.00) | LDD0018 | [13] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [14] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [15] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [15] | |
|
NAIA_5 Probe Info |
 |
C92(0.00); C62(0.00) | LDD2223 | [16] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe3 Probe Info |
 |
13.26 | LDD0464 | [17] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C1939(0.50) | LDD2117 | [4] |
| LDCM0025 | 4SU-RNA | DM93 | C928(2.48); C1742(8.55) | LDD0170 | [6] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C928(2.11); C26(3.11); C1742(2.15); C1037(2.20) | LDD0169 | [6] |
| LDCM0561 | Abegg_cp(-)-10 | HeLa | C2042(2.03) | LDD0312 | [10] |
| LDCM0214 | AC1 | HCT 116 | C1939(1.23); C26(0.80) | LDD0531 | [7] |
| LDCM0215 | AC10 | HCT 116 | C1939(1.24) | LDD0532 | [7] |
| LDCM0216 | AC100 | HCT 116 | C1939(0.90); C26(1.18) | LDD0533 | [7] |
| LDCM0217 | AC101 | HCT 116 | C1939(0.98); C26(1.07) | LDD0534 | [7] |
| LDCM0218 | AC102 | HCT 116 | C1939(0.88); C26(1.00) | LDD0535 | [7] |
| LDCM0219 | AC103 | HCT 116 | C1939(0.90); C26(1.09) | LDD0536 | [7] |
| LDCM0220 | AC104 | HCT 116 | C1939(0.79); C26(0.98) | LDD0537 | [7] |
| LDCM0221 | AC105 | HCT 116 | C1939(0.74); C26(1.04) | LDD0538 | [7] |
| LDCM0222 | AC106 | HCT 116 | C1939(0.90); C26(1.05) | LDD0539 | [7] |
| LDCM0223 | AC107 | HCT 116 | C1939(0.93); C26(1.00) | LDD0540 | [7] |
| LDCM0224 | AC108 | HCT 116 | C1939(0.98); C26(1.02) | LDD0541 | [7] |
| LDCM0225 | AC109 | HCT 116 | C1939(0.84); C26(0.87) | LDD0542 | [7] |
| LDCM0226 | AC11 | HCT 116 | C1939(1.27) | LDD0543 | [7] |
| LDCM0227 | AC110 | HCT 116 | C1939(0.85); C26(0.99) | LDD0544 | [7] |
| LDCM0228 | AC111 | HCT 116 | C1939(0.87); C26(1.21) | LDD0545 | [7] |
| LDCM0229 | AC112 | HCT 116 | C1939(0.80); C26(1.00) | LDD0546 | [7] |
| LDCM0230 | AC113 | HCT 116 | C1939(1.03); C2020(0.94); C2021(0.94); C26(1.16) | LDD0547 | [7] |
| LDCM0231 | AC114 | HCT 116 | C1939(1.03); C2020(1.37); C2021(1.37); C26(1.04) | LDD0548 | [7] |
| LDCM0232 | AC115 | HCT 116 | C1939(1.05); C2020(1.04); C2021(1.04); C26(1.00) | LDD0549 | [7] |
| LDCM0233 | AC116 | HCT 116 | C1939(1.00); C2020(0.93); C2021(0.93); C26(1.31) | LDD0550 | [7] |
| LDCM0234 | AC117 | HCT 116 | C1939(0.99); C2020(1.45); C2021(1.45); C26(1.13) | LDD0551 | [7] |
| LDCM0235 | AC118 | HCT 116 | C1939(0.95); C2020(1.29); C2021(1.29); C26(0.87) | LDD0552 | [7] |
| LDCM0236 | AC119 | HCT 116 | C1939(0.96); C2020(1.10); C2021(1.10); C26(1.10) | LDD0553 | [7] |
| LDCM0237 | AC12 | HCT 116 | C1939(1.18) | LDD0554 | [7] |
| LDCM0238 | AC120 | HCT 116 | C1939(1.51); C2020(1.28); C2021(1.28); C26(0.92) | LDD0555 | [7] |
| LDCM0239 | AC121 | HCT 116 | C1939(0.95); C2020(1.20); C2021(1.20); C26(0.90) | LDD0556 | [7] |
| LDCM0240 | AC122 | HCT 116 | C1939(0.77); C2020(0.80); C2021(0.80); C26(0.96) | LDD0557 | [7] |
| LDCM0241 | AC123 | HCT 116 | C1939(0.86); C2020(0.98); C2021(0.98); C26(0.87) | LDD0558 | [7] |
| LDCM0242 | AC124 | HCT 116 | C1939(1.13); C2020(1.07); C2021(1.07); C26(0.82) | LDD0559 | [7] |
| LDCM0243 | AC125 | HCT 116 | C1939(1.02); C2020(1.03); C2021(1.03); C26(0.85) | LDD0560 | [7] |
| LDCM0244 | AC126 | HCT 116 | C1939(0.97); C2020(1.16); C2021(1.16); C26(0.77) | LDD0561 | [7] |
| LDCM0245 | AC127 | HCT 116 | C1939(0.89); C2020(1.09); C2021(1.09); C26(0.85) | LDD0562 | [7] |
| LDCM0246 | AC128 | HCT 116 | C1939(1.43) | LDD0563 | [7] |
| LDCM0247 | AC129 | HCT 116 | C1939(1.27) | LDD0564 | [7] |
| LDCM0249 | AC130 | HCT 116 | C1939(1.25) | LDD0566 | [7] |
| LDCM0250 | AC131 | HCT 116 | C1939(1.17) | LDD0567 | [7] |
| LDCM0251 | AC132 | HCT 116 | C1939(1.40) | LDD0568 | [7] |
| LDCM0252 | AC133 | HCT 116 | C1939(1.19) | LDD0569 | [7] |
| LDCM0253 | AC134 | HCT 116 | C1939(1.18) | LDD0570 | [7] |
| LDCM0254 | AC135 | HCT 116 | C1939(1.19) | LDD0571 | [7] |
| LDCM0255 | AC136 | HCT 116 | C1939(1.27) | LDD0572 | [7] |
| LDCM0256 | AC137 | HCT 116 | C1939(1.09) | LDD0573 | [7] |
| LDCM0257 | AC138 | HCT 116 | C1939(1.13) | LDD0574 | [7] |
| LDCM0258 | AC139 | HCT 116 | C1939(1.16) | LDD0575 | [7] |
| LDCM0259 | AC14 | HCT 116 | C1939(1.17) | LDD0576 | [7] |
| LDCM0260 | AC140 | HCT 116 | C1939(1.15) | LDD0577 | [7] |
| LDCM0261 | AC141 | HCT 116 | C1939(1.29) | LDD0578 | [7] |
| LDCM0262 | AC142 | HCT 116 | C1939(1.19) | LDD0579 | [7] |
| LDCM0263 | AC143 | HCT 116 | C1939(0.68); C26(1.08) | LDD0580 | [7] |
| LDCM0264 | AC144 | HCT 116 | C1939(0.80); C26(0.98) | LDD0581 | [7] |
| LDCM0265 | AC145 | HCT 116 | C1939(0.98); C26(1.04) | LDD0582 | [7] |
| LDCM0266 | AC146 | HCT 116 | C1939(0.88); C26(1.10) | LDD0583 | [7] |
| LDCM0267 | AC147 | HCT 116 | C1939(0.82); C26(0.87) | LDD0584 | [7] |
| LDCM0268 | AC148 | HCT 116 | C1939(0.85); C26(1.12) | LDD0585 | [7] |
| LDCM0269 | AC149 | HCT 116 | C1939(0.70); C26(1.19) | LDD0586 | [7] |
| LDCM0270 | AC15 | HCT 116 | C1939(0.95) | LDD0587 | [7] |
| LDCM0271 | AC150 | HCT 116 | C1939(0.79); C26(0.93) | LDD0588 | [7] |
| LDCM0272 | AC151 | HCT 116 | C1939(0.84); C26(0.97) | LDD0589 | [7] |
| LDCM0273 | AC152 | HCT 116 | C1939(0.79); C26(1.02) | LDD0590 | [7] |
| LDCM0274 | AC153 | HCT 116 | C1939(0.74); C26(1.50) | LDD0591 | [7] |
| LDCM0621 | AC154 | HCT 116 | C1939(0.90); C26(0.99) | LDD2158 | [7] |
| LDCM0622 | AC155 | HCT 116 | C1939(0.75); C26(1.20) | LDD2159 | [7] |
| LDCM0623 | AC156 | HCT 116 | C1939(0.77); C26(0.99) | LDD2160 | [7] |
| LDCM0624 | AC157 | HCT 116 | C1939(0.84); C26(1.25) | LDD2161 | [7] |
| LDCM0276 | AC17 | HCT 116 | C1939(0.98) | LDD0593 | [7] |
| LDCM0277 | AC18 | HCT 116 | C1939(1.06) | LDD0594 | [7] |
| LDCM0278 | AC19 | HCT 116 | C1939(0.81) | LDD0595 | [7] |
| LDCM0279 | AC2 | HCT 116 | C1939(1.14); C26(1.17) | LDD0596 | [7] |
| LDCM0280 | AC20 | HCT 116 | C1939(0.74) | LDD0597 | [7] |
| LDCM0281 | AC21 | HCT 116 | C1939(0.83) | LDD0598 | [7] |
| LDCM0282 | AC22 | HCT 116 | C1939(0.99) | LDD0599 | [7] |
| LDCM0283 | AC23 | HCT 116 | C1939(0.87) | LDD0600 | [7] |
| LDCM0284 | AC24 | HCT 116 | C1939(0.95) | LDD0601 | [7] |
| LDCM0285 | AC25 | HCT 116 | C26(1.25); C1939(1.33) | LDD0602 | [7] |
| LDCM0286 | AC26 | HCT 116 | C1939(1.14); C26(1.28) | LDD0603 | [7] |
| LDCM0287 | AC27 | HCT 116 | C26(1.14); C1939(1.16) | LDD0604 | [7] |
| LDCM0288 | AC28 | HCT 116 | C26(1.09); C1939(1.23) | LDD0605 | [7] |
| LDCM0289 | AC29 | HCT 116 | C1939(1.39); C26(1.50) | LDD0606 | [7] |
| LDCM0290 | AC3 | HCT 116 | C26(1.07); C1939(1.32) | LDD0607 | [7] |
| LDCM0291 | AC30 | HCT 116 | C1939(0.95); C26(1.50) | LDD0608 | [7] |
| LDCM0292 | AC31 | HCT 116 | C1939(1.26); C26(1.51) | LDD0609 | [7] |
| LDCM0293 | AC32 | HCT 116 | C26(1.47); C1939(1.55) | LDD0610 | [7] |
| LDCM0294 | AC33 | HCT 116 | C26(1.18); C1939(1.20) | LDD0611 | [7] |
| LDCM0295 | AC34 | HCT 116 | C1939(1.14); C26(1.27) | LDD0612 | [7] |
| LDCM0296 | AC35 | HCT 116 | C26(0.78); C1939(0.82) | LDD0613 | [7] |
| LDCM0297 | AC36 | HCT 116 | C1939(0.71); C26(0.97) | LDD0614 | [7] |
| LDCM0298 | AC37 | HCT 116 | C1939(0.78); C26(1.21) | LDD0615 | [7] |
| LDCM0299 | AC38 | HCT 116 | C1939(0.85); C26(1.34) | LDD0616 | [7] |
| LDCM0300 | AC39 | HCT 116 | C1939(0.78); C26(1.26) | LDD0617 | [7] |
| LDCM0301 | AC4 | HCT 116 | C26(0.94); C1939(1.46) | LDD0618 | [7] |
| LDCM0302 | AC40 | HCT 116 | C1939(0.83); C26(1.28) | LDD0619 | [7] |
| LDCM0303 | AC41 | HCT 116 | C1939(0.89); C26(1.04) | LDD0620 | [7] |
| LDCM0304 | AC42 | HCT 116 | C1939(0.83); C26(1.46) | LDD0621 | [7] |
| LDCM0305 | AC43 | HCT 116 | C1939(0.78); C26(1.23) | LDD0622 | [7] |
| LDCM0306 | AC44 | HCT 116 | C1939(0.74); C26(1.23) | LDD0623 | [7] |
| LDCM0307 | AC45 | HCT 116 | C1939(0.72); C26(1.34) | LDD0624 | [7] |
| LDCM0308 | AC46 | HCT 116 | C1939(0.82); C26(1.04) | LDD0625 | [7] |
| LDCM0309 | AC47 | HCT 116 | C1939(0.94); C26(1.11) | LDD0626 | [7] |
| LDCM0310 | AC48 | HCT 116 | C1939(1.02); C26(1.68) | LDD0627 | [7] |
| LDCM0311 | AC49 | HCT 116 | C1939(0.81); C26(1.83) | LDD0628 | [7] |
| LDCM0312 | AC5 | HCT 116 | C26(1.12); C1939(1.31) | LDD0629 | [7] |
| LDCM0313 | AC50 | HCT 116 | C1939(0.95); C26(1.65) | LDD0630 | [7] |
| LDCM0314 | AC51 | HCT 116 | C1939(0.79); C26(0.91) | LDD0631 | [7] |
| LDCM0315 | AC52 | HCT 116 | C1939(0.82); C26(1.18) | LDD0632 | [7] |
| LDCM0316 | AC53 | HCT 116 | C1939(0.71); C26(1.38) | LDD0633 | [7] |
| LDCM0317 | AC54 | HCT 116 | C1939(0.86); C26(1.32) | LDD0634 | [7] |
| LDCM0318 | AC55 | HCT 116 | C1939(0.93); C26(1.94) | LDD0635 | [7] |
| LDCM0319 | AC56 | HCT 116 | C1939(0.86); C26(2.73) | LDD0636 | [7] |
| LDCM0320 | AC57 | HCT 116 | C1939(1.10); C26(1.18) | LDD0637 | [7] |
| LDCM0321 | AC58 | HCT 116 | C26(0.92); C1939(1.18) | LDD0638 | [7] |
| LDCM0322 | AC59 | HCT 116 | C1939(1.15); C26(1.29) | LDD0639 | [7] |
| LDCM0323 | AC6 | HCT 116 | C1939(1.07) | LDD0640 | [7] |
| LDCM0324 | AC60 | HCT 116 | C26(1.31); C1939(1.32) | LDD0641 | [7] |
| LDCM0325 | AC61 | HCT 116 | C1939(1.03); C26(1.14) | LDD0642 | [7] |
| LDCM0326 | AC62 | HCT 116 | C1939(1.15); C26(1.26) | LDD0643 | [7] |
| LDCM0327 | AC63 | HCT 116 | C1939(0.94); C26(1.24) | LDD0644 | [7] |
| LDCM0328 | AC64 | HCT 116 | C1939(1.20); C26(1.24) | LDD0645 | [7] |
| LDCM0329 | AC65 | HCT 116 | C26(1.09); C1939(1.17) | LDD0646 | [7] |
| LDCM0330 | AC66 | HCT 116 | C1939(1.01); C26(1.13) | LDD0647 | [7] |
| LDCM0331 | AC67 | HCT 116 | C1939(1.14); C26(1.30) | LDD0648 | [7] |
| LDCM0332 | AC68 | HCT 116 | C26(0.79); C1939(1.16) | LDD0649 | [7] |
| LDCM0333 | AC69 | HCT 116 | C26(0.96); C1939(1.02) | LDD0650 | [7] |
| LDCM0334 | AC7 | HCT 116 | C1939(0.96) | LDD0651 | [7] |
| LDCM0335 | AC70 | HCT 116 | C1939(0.93); C26(1.11) | LDD0652 | [7] |
| LDCM0336 | AC71 | HCT 116 | C26(1.02); C1939(1.20) | LDD0653 | [7] |
| LDCM0337 | AC72 | HCT 116 | C26(1.00); C1939(1.48) | LDD0654 | [7] |
| LDCM0338 | AC73 | HCT 116 | C26(0.95); C1939(1.31) | LDD0655 | [7] |
| LDCM0339 | AC74 | HCT 116 | C26(1.12); C1939(1.39) | LDD0656 | [7] |
| LDCM0340 | AC75 | HCT 116 | C1939(0.93); C26(1.34) | LDD0657 | [7] |
| LDCM0341 | AC76 | HCT 116 | C26(1.05); C1939(1.19) | LDD0658 | [7] |
| LDCM0342 | AC77 | HCT 116 | C26(0.86); C1939(0.97) | LDD0659 | [7] |
| LDCM0343 | AC78 | HCT 116 | C1939(1.01); C26(1.19) | LDD0660 | [7] |
| LDCM0344 | AC79 | HCT 116 | C26(0.99); C1939(1.03) | LDD0661 | [7] |
| LDCM0345 | AC8 | HCT 116 | C1939(1.10) | LDD0662 | [7] |
| LDCM0346 | AC80 | HCT 116 | C26(1.19); C1939(1.28) | LDD0663 | [7] |
| LDCM0347 | AC81 | HCT 116 | C1939(1.03); C26(1.06) | LDD0664 | [7] |
| LDCM0348 | AC82 | HCT 116 | C26(1.17); C1939(1.25) | LDD0665 | [7] |
| LDCM0349 | AC83 | HCT 116 | C1939(1.13); C26(1.59) | LDD0666 | [7] |
| LDCM0350 | AC84 | HCT 116 | C1939(1.10); C26(1.31) | LDD0667 | [7] |
| LDCM0351 | AC85 | HCT 116 | C1939(1.09); C26(1.40) | LDD0668 | [7] |
| LDCM0352 | AC86 | HCT 116 | C1939(1.15); C26(1.26) | LDD0669 | [7] |
| LDCM0353 | AC87 | HCT 116 | C26(0.98); C1939(1.11) | LDD0670 | [7] |
| LDCM0354 | AC88 | HCT 116 | C1939(0.89); C26(0.99) | LDD0671 | [7] |
| LDCM0355 | AC89 | HCT 116 | C1939(1.09); C26(1.33) | LDD0672 | [7] |
| LDCM0357 | AC90 | HCT 116 | C1939(1.07); C26(1.13) | LDD0674 | [7] |
| LDCM0358 | AC91 | HCT 116 | C1939(1.27); C26(1.58) | LDD0675 | [7] |
| LDCM0359 | AC92 | HCT 116 | C1939(1.21); C26(1.55) | LDD0676 | [7] |
| LDCM0360 | AC93 | HCT 116 | C1939(0.97); C26(1.03) | LDD0677 | [7] |
| LDCM0361 | AC94 | HCT 116 | C1939(0.98); C26(1.29) | LDD0678 | [7] |
| LDCM0362 | AC95 | HCT 116 | C1939(1.11); C26(1.12) | LDD0679 | [7] |
| LDCM0363 | AC96 | HCT 116 | C1939(0.97); C26(1.49) | LDD0680 | [7] |
| LDCM0364 | AC97 | HCT 116 | C1939(1.02); C26(1.30) | LDD0681 | [7] |
| LDCM0365 | AC98 | HCT 116 | C1939(0.90); C26(0.95) | LDD0682 | [7] |
| LDCM0366 | AC99 | HCT 116 | C1939(0.80); C26(0.94) | LDD0683 | [7] |
| LDCM0248 | AKOS034007472 | HCT 116 | C1939(0.96) | LDD0565 | [7] |
| LDCM0356 | AKOS034007680 | HCT 116 | C1939(1.03) | LDD0673 | [7] |
| LDCM0275 | AKOS034007705 | HCT 116 | C1939(1.05) | LDD0592 | [7] |
| LDCM0156 | Aniline | NCI-H1299 | 12.00 | LDD0403 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C26(1.03) | LDD0078 | [7] |
| LDCM0108 | Chloroacetamide | HeLa | C1939(0.00); H1925(0.00); C2049(0.00) | LDD0222 | [15] |
| LDCM0367 | CL1 | HCT 116 | C2020(0.97); C2021(0.97); C1939(1.04); C26(1.04) | LDD0684 | [7] |
| LDCM0368 | CL10 | HCT 116 | C2020(0.83); C2021(0.83); C26(0.93); C1939(1.86) | LDD0685 | [7] |
| LDCM0369 | CL100 | HCT 116 | C26(0.89); C1939(1.17) | LDD0686 | [7] |
| LDCM0370 | CL101 | HCT 116 | C1939(0.91) | LDD0687 | [7] |
| LDCM0371 | CL102 | HCT 116 | C1939(1.11) | LDD0688 | [7] |
| LDCM0372 | CL103 | HCT 116 | C1939(1.11) | LDD0689 | [7] |
| LDCM0373 | CL104 | HCT 116 | C1939(1.05) | LDD0690 | [7] |
| LDCM0374 | CL105 | HCT 116 | C1939(0.96) | LDD0691 | [7] |
| LDCM0375 | CL106 | HCT 116 | C1939(1.12) | LDD0692 | [7] |
| LDCM0376 | CL107 | HCT 116 | C1939(0.91) | LDD0693 | [7] |
| LDCM0377 | CL108 | HCT 116 | C1939(1.00) | LDD0694 | [7] |
| LDCM0378 | CL109 | HCT 116 | C1939(0.96) | LDD0695 | [7] |
| LDCM0379 | CL11 | HCT 116 | C2020(0.85); C2021(0.85); C26(1.06); C1939(1.13) | LDD0696 | [7] |
| LDCM0380 | CL110 | HCT 116 | C1939(1.06) | LDD0697 | [7] |
| LDCM0381 | CL111 | HCT 116 | C1939(1.00) | LDD0698 | [7] |
| LDCM0382 | CL112 | HCT 116 | C26(1.08); C1939(1.16) | LDD0699 | [7] |
| LDCM0383 | CL113 | HCT 116 | C1939(0.85); C26(1.37) | LDD0700 | [7] |
| LDCM0384 | CL114 | HCT 116 | C1939(1.04); C26(1.56) | LDD0701 | [7] |
| LDCM0385 | CL115 | HCT 116 | C1939(1.31); C26(1.32) | LDD0702 | [7] |
| LDCM0386 | CL116 | HCT 116 | C1939(1.23); C26(1.44) | LDD0703 | [7] |
| LDCM0387 | CL117 | HCT 116 | C1939(0.70); C26(1.55) | LDD0704 | [7] |
| LDCM0388 | CL118 | HCT 116 | C1939(0.79); C26(1.20) | LDD0705 | [7] |
| LDCM0389 | CL119 | HCT 116 | C1939(0.90); C26(1.16) | LDD0706 | [7] |
| LDCM0390 | CL12 | HCT 116 | C2020(0.88); C2021(0.88); C26(1.05); C1939(2.53) | LDD0707 | [7] |
| LDCM0391 | CL120 | HCT 116 | C1939(0.85); C26(1.09) | LDD0708 | [7] |
| LDCM0392 | CL121 | HCT 116 | C1939(0.87); C26(1.05) | LDD0709 | [7] |
| LDCM0393 | CL122 | HCT 116 | C1939(0.81); C26(1.47) | LDD0710 | [7] |
| LDCM0394 | CL123 | HCT 116 | C1939(0.88); C26(1.58) | LDD0711 | [7] |
| LDCM0395 | CL124 | HCT 116 | C1939(1.02); C26(1.91) | LDD0712 | [7] |
| LDCM0396 | CL125 | HCT 116 | C1939(1.01); C26(1.17) | LDD0713 | [7] |
| LDCM0397 | CL126 | HCT 116 | C26(1.07); C1939(1.19) | LDD0714 | [7] |
| LDCM0398 | CL127 | HCT 116 | C1939(1.16); C26(1.19) | LDD0715 | [7] |
| LDCM0399 | CL128 | HCT 116 | C1939(1.05); C26(1.06) | LDD0716 | [7] |
| LDCM0400 | CL13 | HCT 116 | C2020(0.82); C2021(0.82); C26(1.04); C1939(3.11) | LDD0717 | [7] |
| LDCM0401 | CL14 | HCT 116 | C2020(0.93); C2021(0.93); C26(0.96); C1939(2.82) | LDD0718 | [7] |
| LDCM0402 | CL15 | HCT 116 | C2020(0.83); C2021(0.83); C26(1.03); C1939(3.67) | LDD0719 | [7] |
| LDCM0403 | CL16 | HCT 116 | C1939(0.85); C2020(1.27); C2021(1.27) | LDD0720 | [7] |
| LDCM0404 | CL17 | HCT 116 | C1939(0.81); C2020(0.89); C2021(0.89) | LDD0721 | [7] |
| LDCM0405 | CL18 | HCT 116 | C2020(0.68); C2021(0.68); C1939(0.85) | LDD0722 | [7] |
| LDCM0406 | CL19 | HCT 116 | C1939(0.83); C2020(0.92); C2021(0.92) | LDD0723 | [7] |
| LDCM0407 | CL2 | HCT 116 | C2020(0.80); C2021(0.80); C26(1.20); C1939(1.23) | LDD0724 | [7] |
| LDCM0408 | CL20 | HCT 116 | C1939(0.88); C2020(0.89); C2021(0.89) | LDD0725 | [7] |
| LDCM0409 | CL21 | HCT 116 | C1939(0.82); C2020(0.88); C2021(0.88) | LDD0726 | [7] |
| LDCM0410 | CL22 | HCT 116 | C2020(0.96); C2021(0.96); C1939(1.06) | LDD0727 | [7] |
| LDCM0411 | CL23 | HCT 116 | C2020(0.83); C2021(0.83); C1939(0.94) | LDD0728 | [7] |
| LDCM0412 | CL24 | HCT 116 | C2020(0.76); C2021(0.76); C1939(1.03) | LDD0729 | [7] |
| LDCM0413 | CL25 | HCT 116 | C1939(1.09); C2020(0.98); C2021(0.98) | LDD0730 | [7] |
| LDCM0414 | CL26 | HCT 116 | C1939(0.88); C2020(0.81); C2021(0.81) | LDD0731 | [7] |
| LDCM0415 | CL27 | HCT 116 | C1939(0.97); C2020(0.86); C2021(0.86) | LDD0732 | [7] |
| LDCM0416 | CL28 | HCT 116 | C1939(0.90); C2020(0.88); C2021(0.88) | LDD0733 | [7] |
| LDCM0417 | CL29 | HCT 116 | C1939(0.82); C2020(0.79); C2021(0.79) | LDD0734 | [7] |
| LDCM0418 | CL3 | HCT 116 | C1939(1.23); C2020(0.78); C2021(0.78); C26(1.01) | LDD0735 | [7] |
| LDCM0419 | CL30 | HCT 116 | C1939(0.80); C2020(0.74); C2021(0.74) | LDD0736 | [7] |
| LDCM0420 | CL31 | HCT 116 | C1939(1.05); C26(1.02) | LDD0737 | [7] |
| LDCM0421 | CL32 | HCT 116 | C1939(1.15); C26(0.96) | LDD0738 | [7] |
| LDCM0422 | CL33 | HCT 116 | C1939(1.10); C26(1.16) | LDD0739 | [7] |
| LDCM0423 | CL34 | HCT 116 | C1939(1.17); C26(1.05) | LDD0740 | [7] |
| LDCM0424 | CL35 | HCT 116 | C1939(1.20); C26(1.11) | LDD0741 | [7] |
| LDCM0425 | CL36 | HCT 116 | C1939(1.08); C26(1.02) | LDD0742 | [7] |
| LDCM0426 | CL37 | HCT 116 | C1939(1.91); C26(0.84) | LDD0743 | [7] |
| LDCM0428 | CL39 | HCT 116 | C1939(1.08); C26(1.05) | LDD0745 | [7] |
| LDCM0429 | CL4 | HCT 116 | C1939(1.21); C2020(0.83); C2021(0.83); C26(1.05) | LDD0746 | [7] |
| LDCM0430 | CL40 | HCT 116 | C1939(0.98); C26(1.05) | LDD0747 | [7] |
| LDCM0431 | CL41 | HCT 116 | C1939(0.94); C26(0.94) | LDD0748 | [7] |
| LDCM0432 | CL42 | HCT 116 | C1939(1.09); C26(1.03) | LDD0749 | [7] |
| LDCM0433 | CL43 | HCT 116 | C1939(0.96); C26(1.07) | LDD0750 | [7] |
| LDCM0434 | CL44 | HCT 116 | C1939(1.11); C26(0.95) | LDD0751 | [7] |
| LDCM0435 | CL45 | HCT 116 | C1939(1.38); C26(0.92) | LDD0752 | [7] |
| LDCM0436 | CL46 | HCT 116 | C1939(0.93); C26(0.69) | LDD0753 | [7] |
| LDCM0437 | CL47 | HCT 116 | C1939(0.73); C26(0.70) | LDD0754 | [7] |
| LDCM0438 | CL48 | HCT 116 | C1939(0.73); C26(0.80) | LDD0755 | [7] |
| LDCM0439 | CL49 | HCT 116 | C1939(1.10); C26(0.81) | LDD0756 | [7] |
| LDCM0440 | CL5 | HCT 116 | C1939(1.87); C2020(0.83); C2021(0.83); C26(1.13) | LDD0757 | [7] |
| LDCM0441 | CL50 | HCT 116 | C1939(0.98); C26(1.02) | LDD0758 | [7] |
| LDCM0442 | CL51 | HCT 116 | C1939(0.84); C26(1.04) | LDD0759 | [7] |
| LDCM0443 | CL52 | HCT 116 | C1939(0.78); C26(1.39) | LDD0760 | [7] |
| LDCM0444 | CL53 | HCT 116 | C1939(0.68); C26(0.99) | LDD0761 | [7] |
| LDCM0445 | CL54 | HCT 116 | C1939(0.96); C26(0.76) | LDD0762 | [7] |
| LDCM0446 | CL55 | HCT 116 | C1939(0.82); C26(0.89) | LDD0763 | [7] |
| LDCM0447 | CL56 | HCT 116 | C1939(0.85); C26(1.11) | LDD0764 | [7] |
| LDCM0448 | CL57 | HCT 116 | C1939(0.75); C26(1.02) | LDD0765 | [7] |
| LDCM0449 | CL58 | HCT 116 | C1939(1.16); C26(0.66) | LDD0766 | [7] |
| LDCM0450 | CL59 | HCT 116 | C1939(0.91); C26(0.87) | LDD0767 | [7] |
| LDCM0451 | CL6 | HCT 116 | C1939(1.88); C2020(0.82); C2021(0.82); C26(0.99) | LDD0768 | [7] |
| LDCM0452 | CL60 | HCT 116 | C1939(0.97); C26(0.72) | LDD0769 | [7] |
| LDCM0453 | CL61 | HCT 116 | C1939(1.25); C2020(0.81); C2021(0.81); C26(1.04) | LDD0770 | [7] |
| LDCM0454 | CL62 | HCT 116 | C1939(1.43); C2020(1.06); C2021(1.06); C26(0.80) | LDD0771 | [7] |
| LDCM0455 | CL63 | HCT 116 | C1939(1.42); C2020(0.63); C2021(0.63); C26(0.72) | LDD0772 | [7] |
| LDCM0456 | CL64 | HCT 116 | C1939(1.28); C2020(0.79); C2021(0.79); C26(0.85) | LDD0773 | [7] |
| LDCM0457 | CL65 | HCT 116 | C1939(1.19); C2020(0.59); C2021(0.59); C26(0.67) | LDD0774 | [7] |
| LDCM0458 | CL66 | HCT 116 | C1939(1.33); C2020(0.95); C2021(0.95); C26(1.06) | LDD0775 | [7] |
| LDCM0459 | CL67 | HCT 116 | C1939(1.22); C2020(1.07); C2021(1.07); C26(0.66) | LDD0776 | [7] |
| LDCM0460 | CL68 | HCT 116 | C1939(1.16); C2020(0.80); C2021(0.80); C26(0.84) | LDD0777 | [7] |
| LDCM0461 | CL69 | HCT 116 | C1939(1.12); C2020(0.78); C2021(0.78); C26(1.11) | LDD0778 | [7] |
| LDCM0462 | CL7 | HCT 116 | C1939(3.05); C2020(0.97); C2021(0.97); C26(0.96) | LDD0779 | [7] |
| LDCM0463 | CL70 | HCT 116 | C1939(1.15); C2020(0.92); C2021(0.92); C26(0.76) | LDD0780 | [7] |
| LDCM0464 | CL71 | HCT 116 | C1939(1.13); C2020(0.57); C2021(0.57); C26(1.14) | LDD0781 | [7] |
| LDCM0465 | CL72 | HCT 116 | C1939(1.17); C2020(0.80); C2021(0.80); C26(0.89) | LDD0782 | [7] |
| LDCM0466 | CL73 | HCT 116 | C1939(1.23); C2020(0.78); C2021(0.78); C26(0.84) | LDD0783 | [7] |
| LDCM0467 | CL74 | HCT 116 | C1939(1.08); C2020(1.14); C2021(1.14); C26(0.87) | LDD0784 | [7] |
| LDCM0469 | CL76 | HCT 116 | C1939(0.99); C2020(0.51); C2021(0.51); C26(1.28) | LDD0786 | [7] |
| LDCM0470 | CL77 | HCT 116 | C1939(0.98); C2020(0.56); C2021(0.56); C26(1.33) | LDD0787 | [7] |
| LDCM0471 | CL78 | HCT 116 | C1939(1.03); C2020(0.54); C2021(0.54); C26(1.24) | LDD0788 | [7] |
| LDCM0472 | CL79 | HCT 116 | C1939(0.99); C2020(0.62); C2021(0.62); C26(0.96) | LDD0789 | [7] |
| LDCM0473 | CL8 | HCT 116 | C1939(0.75); C2020(1.00); C2021(1.00); C26(1.16) | LDD0790 | [7] |
| LDCM0474 | CL80 | HCT 116 | C1939(1.05); C2020(0.86); C2021(0.86); C26(1.26) | LDD0791 | [7] |
| LDCM0475 | CL81 | HCT 116 | C1939(1.05); C2020(0.82); C2021(0.82); C26(1.16) | LDD0792 | [7] |
| LDCM0476 | CL82 | HCT 116 | C1939(1.16); C2020(0.57); C2021(0.57); C26(1.07) | LDD0793 | [7] |
| LDCM0477 | CL83 | HCT 116 | C1939(1.08); C2020(0.79); C2021(0.79); C26(1.11) | LDD0794 | [7] |
| LDCM0478 | CL84 | HCT 116 | C1939(1.17); C2020(0.73); C2021(0.73); C26(0.99) | LDD0795 | [7] |
| LDCM0479 | CL85 | HCT 116 | C1939(0.97); C2020(0.53); C2021(0.53); C26(1.13) | LDD0796 | [7] |
| LDCM0480 | CL86 | HCT 116 | C1939(1.12); C2020(0.47); C2021(0.47); C26(1.11) | LDD0797 | [7] |
| LDCM0481 | CL87 | HCT 116 | C1939(1.12); C2020(0.56); C2021(0.56); C26(0.94) | LDD0798 | [7] |
| LDCM0482 | CL88 | HCT 116 | C1939(1.09); C2020(0.59); C2021(0.59); C26(1.03) | LDD0799 | [7] |
| LDCM0483 | CL89 | HCT 116 | C1939(1.25); C2020(0.89); C2021(0.89); C26(1.00) | LDD0800 | [7] |
| LDCM0484 | CL9 | HCT 116 | C1939(1.39); C2020(0.94); C2021(0.94); C26(1.25) | LDD0801 | [7] |
| LDCM0485 | CL90 | HCT 116 | C1939(1.05); C2020(0.56); C2021(0.56); C26(1.16) | LDD0802 | [7] |
| LDCM0486 | CL91 | HCT 116 | C1939(1.12); C26(1.45) | LDD0803 | [7] |
| LDCM0487 | CL92 | HCT 116 | C1939(1.12); C26(1.03) | LDD0804 | [7] |
| LDCM0488 | CL93 | HCT 116 | C1939(1.14); C26(0.91) | LDD0805 | [7] |
| LDCM0489 | CL94 | HCT 116 | C1939(1.13); C26(0.75) | LDD0806 | [7] |
| LDCM0490 | CL95 | HCT 116 | C1939(1.16); C26(1.25) | LDD0807 | [7] |
| LDCM0491 | CL96 | HCT 116 | C1939(1.21); C26(1.14) | LDD0808 | [7] |
| LDCM0492 | CL97 | HCT 116 | C1939(1.15); C26(1.08) | LDD0809 | [7] |
| LDCM0493 | CL98 | HCT 116 | C1939(1.17); C26(0.96) | LDD0810 | [7] |
| LDCM0494 | CL99 | HCT 116 | C1939(1.19); C26(0.92) | LDD0811 | [7] |
| LDCM0495 | E2913 | HEK-293T | C26(1.01); C1742(0.98); C1037(1.08); C1939(0.89) | LDD1698 | [18] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C1939(1.77); C1742(1.05); C26(1.01) | LDD1702 | [4] |
| LDCM0625 | F8 | Ramos | C26(0.98) | LDD2187 | [19] |
| LDCM0572 | Fragment10 | Ramos | C26(0.79) | LDD2189 | [19] |
| LDCM0574 | Fragment12 | Ramos | C26(0.39) | LDD2191 | [19] |
| LDCM0575 | Fragment13 | Ramos | C26(0.48) | LDD2192 | [19] |
| LDCM0576 | Fragment14 | Ramos | C26(0.73) | LDD2193 | [19] |
| LDCM0579 | Fragment20 | Ramos | C26(0.41) | LDD2194 | [19] |
| LDCM0580 | Fragment21 | Ramos | C26(0.48) | LDD2195 | [19] |
| LDCM0582 | Fragment23 | Ramos | C26(0.47) | LDD2196 | [19] |
| LDCM0578 | Fragment27 | Ramos | C26(0.38) | LDD2197 | [19] |
| LDCM0586 | Fragment28 | Ramos | C26(0.48) | LDD2198 | [19] |
| LDCM0588 | Fragment30 | Ramos | C26(0.44) | LDD2199 | [19] |
| LDCM0589 | Fragment31 | Ramos | C26(0.35) | LDD2200 | [19] |
| LDCM0590 | Fragment32 | Ramos | C26(0.57) | LDD2201 | [19] |
| LDCM0468 | Fragment33 | HCT 116 | C1939(1.01); C2020(0.70); C2021(0.70); C26(0.88) | LDD0785 | [7] |
| LDCM0596 | Fragment38 | Ramos | C26(0.36) | LDD2203 | [19] |
| LDCM0566 | Fragment4 | Ramos | C26(0.90) | LDD2184 | [19] |
| LDCM0427 | Fragment51 | HCT 116 | C1939(1.28); C26(1.10) | LDD0744 | [7] |
| LDCM0610 | Fragment52 | Ramos | C26(0.45) | LDD2204 | [19] |
| LDCM0614 | Fragment56 | Ramos | C26(0.57) | LDD2205 | [19] |
| LDCM0569 | Fragment7 | Ramos | C26(0.94) | LDD2186 | [19] |
| LDCM0571 | Fragment9 | Ramos | C26(0.42) | LDD2188 | [19] |
| LDCM0107 | IAA | HeLa | H1925(0.00); H2048(0.00) | LDD0221 | [15] |
| LDCM0022 | KB02 | HCT 116 | C26(1.71) | LDD0080 | [7] |
| LDCM0023 | KB03 | HCT 116 | C26(1.11) | LDD0081 | [7] |
| LDCM0024 | KB05 | HCT 116 | C26(1.44) | LDD0082 | [7] |
| LDCM0006 | Micheliolide | M9-ENL1 | 1.50 | LDD0049 | [5] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [15] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C1939(0.54) | LDD2109 | [4] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C1742(0.98); C1939(0.43) | LDD2123 | [4] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C1742(0.80) | LDD2127 | [4] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C1939(0.60) | LDD2137 | [4] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C1742(1.81) | LDD2147 | [4] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C1742(0.33) | LDD2150 | [4] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C26(0.93) | LDD2206 | [20] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C26(0.66) | LDD2207 | [20] |
| LDCM0021 | THZ1 | HCT 116 | C26(1.12); C1939(0.94); C2042(1.12); C2049(1.12) | LDD2173 | [7] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Histone H2AX (H2AX) | Histone H2A family | P16104 | |||
| DNA repair protein RAD51 homolog 1 (RAD51) | RecA family | Q06609 | |||
| Nibrin (NBN) | . | O60934 | |||
References
