Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P8 Probe Info |
 |
1.79 | LDD0451 | [1] | |
|
A-EBA Probe Info |
 |
4.17 | LDD0215 | [2] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [3] | |
|
C-Sul Probe Info |
 |
2.63 | LDD0066 | [4] | |
|
TH211 Probe Info |
 |
Y133(16.10) | LDD0257 | [5] | |
|
TH214 Probe Info |
 |
Y128(15.49); Y51(12.49) | LDD0258 | [5] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [6] | |
|
AZ-9 Probe Info |
 |
E164(1.37); E154(1.35) | LDD2208 | [7] | |
|
ONAyne Probe Info |
 |
K52(0.00); K178(0.00) | LDD0273 | [8] | |
|
OPA-S-S-alkyne Probe Info |
 |
K43(1.51); K52(1.66); K178(3.53) | LDD3494 | [9] | |
|
Probe 1 Probe Info |
 |
Y110(29.04); Y133(24.90); Y149(41.01); Y156(56.28) | LDD3495 | [10] | |
|
AF-1 Probe Info |
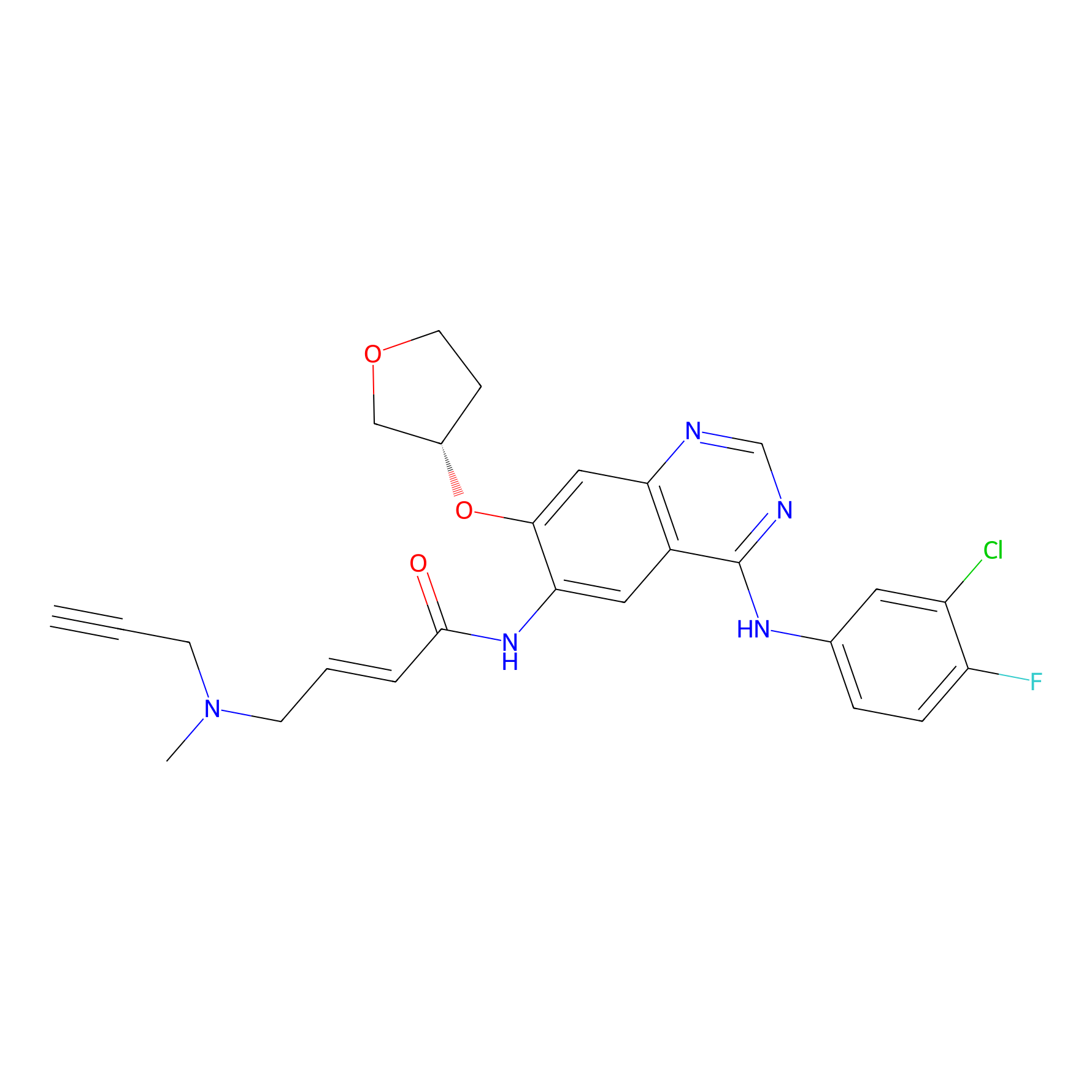 |
1.91 | LDD0421 | [11] | |
|
m-APA Probe Info |
 |
10.37 | LDD0403 | [12] | |
|
DBIA Probe Info |
 |
C79(1.01) | LDD3361 | [13] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [14] | |
|
THZ1-DTB Probe Info |
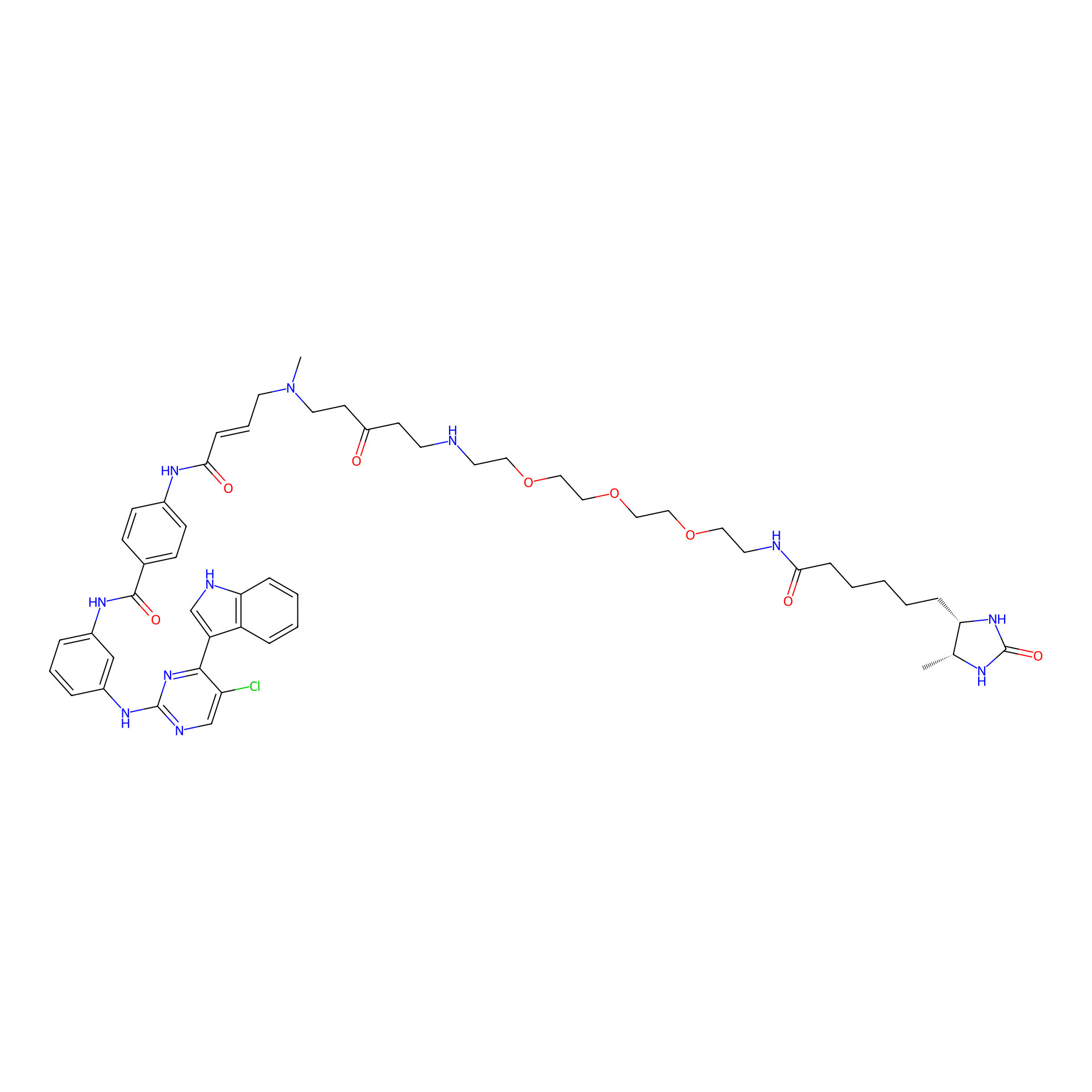 |
C79(1.15) | LDD0460 | [14] | |
|
DA-P3 Probe Info |
 |
11.08 | LDD0183 | [15] | |
|
HHS-482 Probe Info |
 |
Y128(1.00); Y133(1.01); Y144(1.27); Y149(0.73) | LDD0285 | [16] | |
|
HHS-475 Probe Info |
 |
Y144(0.80); Y133(0.92); Y51(1.32) | LDD0264 | [17] | |
|
HHS-465 Probe Info |
 |
Y133(9.35); Y144(3.57); Y156(5.71) | LDD2237 | [18] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0221 | [19] | |
|
ATP probe Probe Info |
 |
K178(0.00); K141(0.00); K138(0.00); K167(0.00) | LDD0199 | [20] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [21] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [21] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [22] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [21] | |
|
ATP probe Probe Info |
 |
K141(0.00); K167(0.00); K52(0.00); K127(0.00) | LDD0035 | [23] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [24] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [25] | |
|
SF Probe Info |
 |
Y133(0.00); K178(0.00); K135(0.00) | LDD0028 | [26] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [25] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [27] | |
|
1c-yne Probe Info |
 |
K43(0.00); K28(0.00) | LDD0228 | [28] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [19] | |
|
AOyne Probe Info |
 |
9.50 | LDD0443 | [29] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [24] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C282 Probe Info |
 |
17.03 | LDD1952 | [30] | |
|
Photocelecoxib Probe Info |
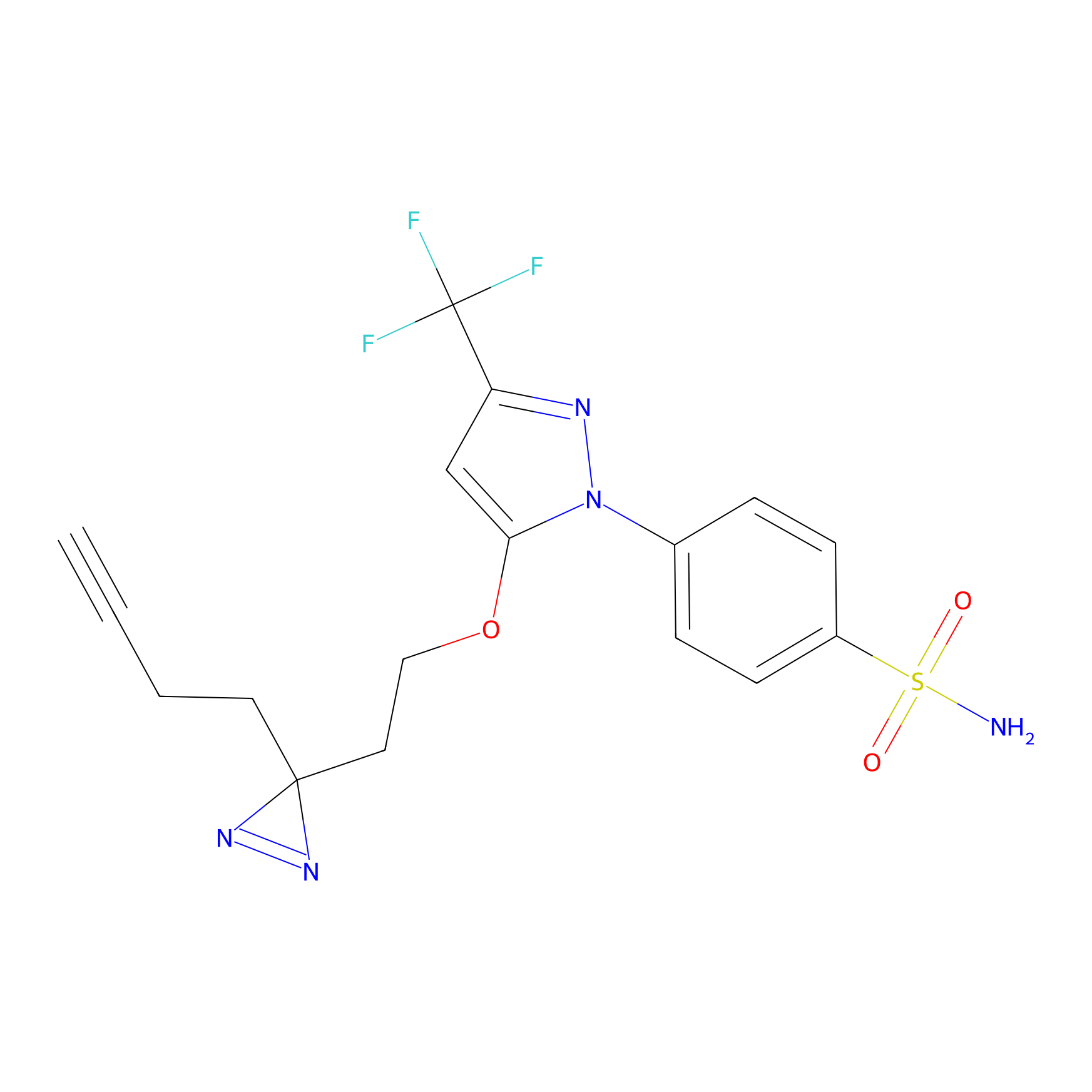 |
V162(0.00); E163(0.00) | LDD0019 | [31] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0070 | [32] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0068 | [33] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C79(0.91) | LDD1507 | [34] |
| LDCM0276 | AC17 | HEK-293T | C79(1.15) | LDD1515 | [34] |
| LDCM0285 | AC25 | HEK-293T | C79(1.08) | LDD1524 | [34] |
| LDCM0294 | AC33 | HEK-293T | C79(1.02) | LDD1533 | [34] |
| LDCM0303 | AC41 | HEK-293T | C79(0.85) | LDD1542 | [34] |
| LDCM0311 | AC49 | HEK-293T | C79(1.02) | LDD1550 | [34] |
| LDCM0320 | AC57 | HEK-293T | C79(0.75) | LDD1559 | [34] |
| LDCM0166 | Afatinib | A431 | 1.91 | LDD0421 | [11] |
| LDCM0356 | AKOS034007680 | HEK-293T | C79(1.06) | LDD1570 | [34] |
| LDCM0156 | Aniline | NCI-H1299 | 10.37 | LDD0403 | [12] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [19] |
| LDCM0404 | CL17 | HEK-293T | C79(1.14) | LDD1608 | [34] |
| LDCM0417 | CL29 | HEK-293T | C79(1.31) | LDD1621 | [34] |
| LDCM0431 | CL41 | HEK-293T | C79(1.17) | LDD1635 | [34] |
| LDCM0440 | CL5 | HEK-293T | C79(1.01) | LDD1644 | [34] |
| LDCM0444 | CL53 | HEK-293T | C79(1.00) | LDD1647 | [34] |
| LDCM0457 | CL65 | HEK-293T | C79(1.04) | LDD1660 | [34] |
| LDCM0470 | CL77 | HEK-293T | C79(1.03) | LDD1673 | [34] |
| LDCM0483 | CL89 | HEK-293T | C79(0.86) | LDD1686 | [34] |
| LDCM0031 | Epigallocatechin gallate | HEK-293T | 11.08 | LDD0183 | [15] |
| LDCM0116 | HHS-0101 | DM93 | Y144(0.80); Y133(0.92); Y51(1.32) | LDD0264 | [17] |
| LDCM0117 | HHS-0201 | DM93 | Y144(0.67); Y133(0.89); Y51(1.11) | LDD0265 | [17] |
| LDCM0118 | HHS-0301 | DM93 | Y133(0.87); Y51(0.99); Y144(1.38) | LDD0266 | [17] |
| LDCM0119 | HHS-0401 | DM93 | Y51(0.73); Y133(0.98); Y144(1.50) | LDD0267 | [17] |
| LDCM0120 | HHS-0701 | DM93 | Y144(0.82); Y133(0.89); Y51(1.04) | LDD0268 | [17] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [19] |
| LDCM0123 | JWB131 | DM93 | Y128(1.00); Y133(1.01); Y144(1.27); Y149(0.73) | LDD0285 | [16] |
| LDCM0124 | JWB142 | DM93 | Y128(0.50); Y133(1.09); Y144(1.57); Y149(0.97) | LDD0286 | [16] |
| LDCM0125 | JWB146 | DM93 | Y128(0.89); Y133(0.92); Y144(1.09); Y149(0.59) | LDD0287 | [16] |
| LDCM0126 | JWB150 | DM93 | Y128(5.61); Y133(3.59); Y144(0.12); Y149(1.76) | LDD0288 | [16] |
| LDCM0127 | JWB152 | DM93 | Y128(2.41); Y133(2.03); Y144(2.10); Y149(1.25) | LDD0289 | [16] |
| LDCM0128 | JWB198 | DM93 | Y128(0.75); Y133(1.81); Y144(0.60); Y149(0.13) | LDD0290 | [16] |
| LDCM0129 | JWB202 | DM93 | Y128(0.53); Y133(1.16); Y144(1.20); Y149(0.61) | LDD0291 | [16] |
| LDCM0130 | JWB211 | DM93 | Y128(0.94); Y133(1.30); Y144(0.67); Y149(0.40) | LDD0292 | [16] |
| LDCM0179 | JZ128 | PC-3 | N.A. | LDD0462 | [14] |
| LDCM0022 | KB02 | Calu-1 | C79(1.12) | LDD2292 | [13] |
| LDCM0023 | KB03 | Calu-1 | C79(1.04) | LDD2709 | [13] |
| LDCM0024 | KB05 | NCI-H358 | C79(1.01) | LDD3361 | [13] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [19] |
| LDCM0021 | THZ1 | HeLa S3 | C79(1.15) | LDD0460 | [14] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Cytokine and receptor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Tumor necrosis factor receptor superfamily member 19 (TNFRSF19) | . | Q9NS68 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| TERF1-interacting nuclear factor 2 (TINF2) | . | Q9BSI4 | |||
References
