Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P3 Probe Info |
 |
2.29 | LDD0450 | [1] | |
|
P8 Probe Info |
 |
2.66 | LDD0451 | [1] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [2] | |
|
Nap-2 Probe Info |
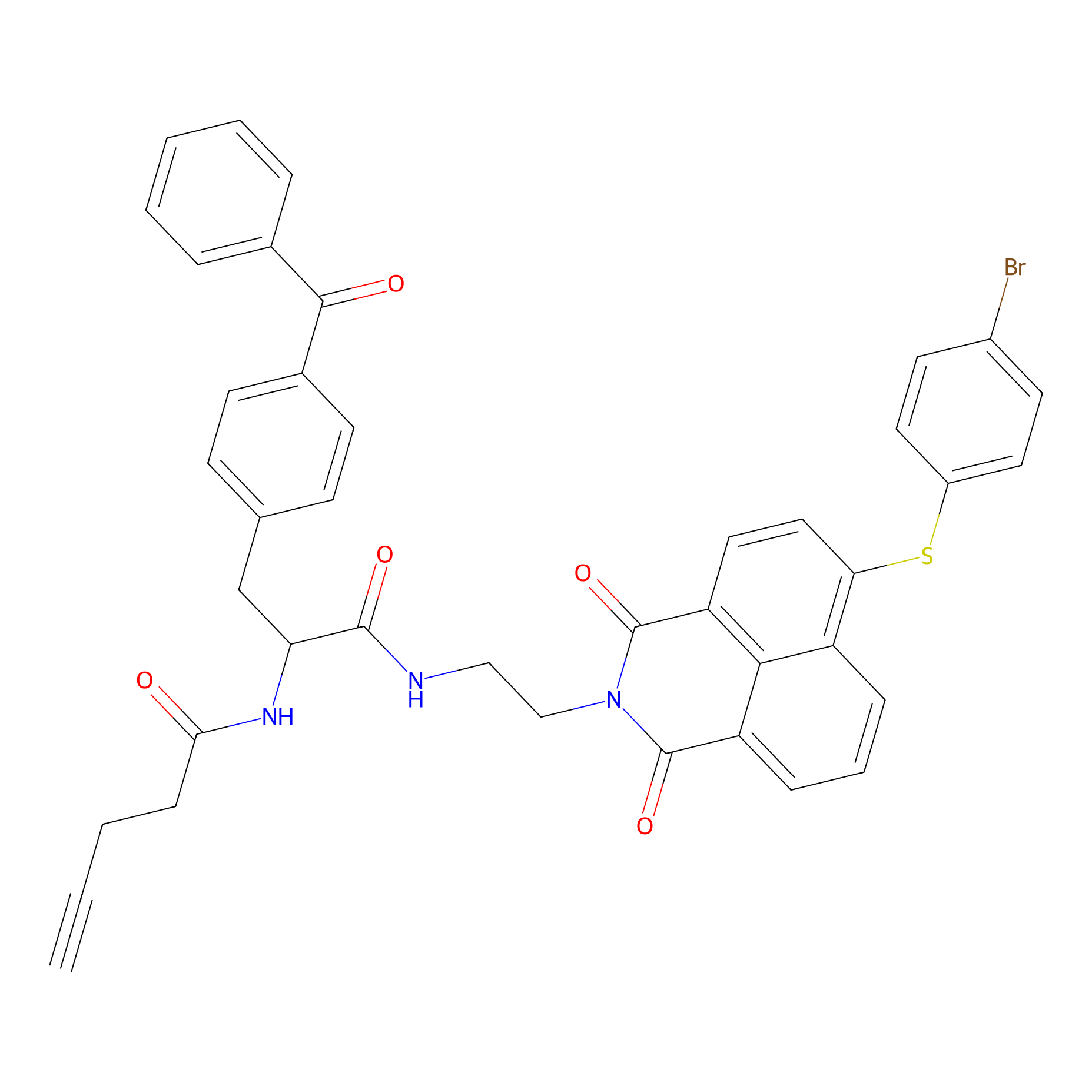 |
N.A. | LDD0435 | [3] | |
|
OPA-S-S-alkyne Probe Info |
 |
K656(0.97) | LDD3494 | [4] | |
|
DBIA Probe Info |
 |
C381(2.78); C573(1.53); C511(1.24); C341(1.31) | LDD3310 | [5] | |
|
BTD Probe Info |
 |
C609(1.08) | LDD2106 | [6] | |
|
Jackson_14 Probe Info |
 |
2.63 | LDD0123 | [7] | |
|
AHL-Pu-1 Probe Info |
 |
C511(2.00) | LDD0168 | [8] | |
|
5E-2FA Probe Info |
 |
H515(0.00); H129(0.00) | LDD2235 | [9] | |
|
m-APA Probe Info |
 |
H515(0.00); H129(0.00) | LDD2231 | [9] | |
|
N1 Probe Info |
 |
C563(0.00); E561(0.00); H568(0.00) | LDD0245 | [10] | |
|
IA-alkyne Probe Info |
 |
C372(0.00); C457(0.00) | LDD0032 | [11] | |
|
Lodoacetamide azide Probe Info |
 |
C372(0.00); C563(0.00) | LDD0037 | [12] | |
|
ATP probe Probe Info |
 |
K426(0.00); K124(0.00) | LDD0035 | [13] | |
|
IPM Probe Info |
 |
C49(0.00); C90(0.00); C511(0.00); C457(0.00) | LDD0025 | [14] | |
|
JW-RF-010 Probe Info |
 |
C511(0.00); C457(0.00) | LDD0026 | [14] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [15] | |
|
TFBX Probe Info |
 |
C341(0.00); C511(0.00); C90(0.00) | LDD0027 | [14] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [16] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [16] | |
|
DA-P3 Probe Info |
 |
N.A. | LDD0178 | [17] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0356 | [18] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [16] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [16] | |
|
PF-06672131 Probe Info |
 |
C511(0.00); C609(0.00) | LDD0017 | [19] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [20] | |
|
VSF Probe Info |
 |
C341(0.00); C90(0.00) | LDD0007 | [16] | |
|
Phosphinate-6 Probe Info |
 |
C511(0.00); C90(0.00); C341(0.00); C49(0.00) | LDD0018 | [21] | |
|
1c-yne Probe Info |
 |
K161(0.00); K124(0.00) | LDD0228 | [18] | |
|
Acrolein Probe Info |
 |
C511(0.00); C49(0.00); C33(0.00); H541(0.00) | LDD0217 | [22] | |
|
Crotonaldehyde Probe Info |
 |
C49(0.00); H360(0.00) | LDD0219 | [22] | |
|
Methacrolein Probe Info |
 |
C511(0.00); C33(0.00); C49(0.00); C457(0.00) | LDD0218 | [22] | |
|
W1 Probe Info |
 |
C372(0.00); C609(0.00); C457(0.00); C511(0.00) | LDD0236 | [23] | |
|
AOyne Probe Info |
 |
7.80 | LDD0443 | [24] | |
|
NAIA_5 Probe Info |
 |
C457(0.00); C204(0.00); C291(0.00); C49(0.00) | LDD2223 | [15] | |
|
HHS-465 Probe Info |
 |
N.A. | LDD2240 | [25] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe11 Probe Info |
 |
12.70 | LDD0471 | [26] | |
|
FFF probe12 Probe Info |
 |
8.54 | LDD0473 | [26] | |
|
FFF probe13 Probe Info |
 |
17.95 | LDD0475 | [26] | |
|
FFF probe15 Probe Info |
 |
5.88 | LDD0478 | [26] | |
|
FFF probe2 Probe Info |
 |
20.00 | LDD0479 | [26] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0464 | [26] | |
|
FFF probe6 Probe Info |
 |
12.44 | LDD0482 | [26] | |
|
FFF probe9 Probe Info |
 |
12.71 | LDD0470 | [26] | |
|
STS-1 Probe Info |
 |
1.09 | LDD0136 | [27] | |
|
STS-2 Probe Info |
 |
1.27 | LDD0138 | [27] | |
|
SAHA-CA-8PAP Probe Info |
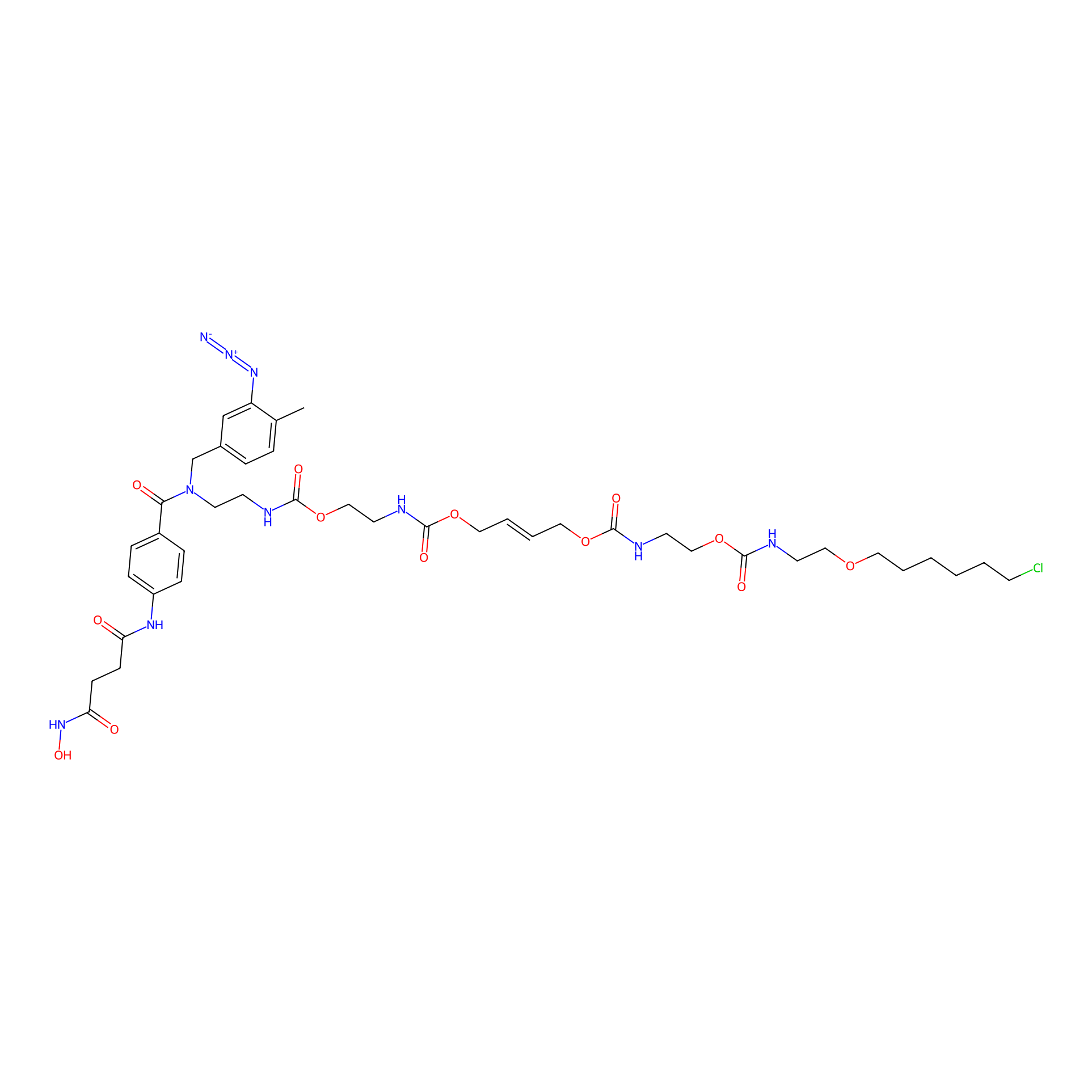 |
5.71 | LDD0362 | [28] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0071 | [29] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | HEK-293T | C511(2.00) | LDD0168 | [8] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C511(2.02); C33(2.56) | LDD0169 | [8] |
| LDCM0108 | Chloroacetamide | HeLa | C511(0.00); C49(0.00); H360(0.00); H541(0.00) | LDD0222 | [22] |
| LDCM0632 | CL-Sc | Hep-G2 | C372(1.70); C511(0.80); C204(0.43); C90(0.40) | LDD2227 | [15] |
| LDCM0189 | Compound 16 | HEK-293T | 71.43 | LDD0492 | [26] |
| LDCM0182 | Compound 18 | HEK-293T | 3.99 | LDD0501 | [26] |
| LDCM0191 | Compound 21 | HEK-293T | 8.58 | LDD0493 | [26] |
| LDCM0190 | Compound 34 | HEK-293T | 125.00 | LDD0497 | [26] |
| LDCM0192 | Compound 35 | HEK-293T | 7.66 | LDD0491 | [26] |
| LDCM0193 | Compound 36 | HEK-293T | 6.19 | LDD0511 | [26] |
| LDCM0195 | Compound 38 | HEK-293T | 14.71 | LDD0499 | [26] |
| LDCM0197 | Compound 40 | HEK-293T | 7.69 | LDD0495 | [26] |
| LDCM0634 | CY-0357 | Hep-G2 | C49(1.29); C511(0.57) | LDD2228 | [15] |
| LDCM0107 | IAA | HeLa | H360(0.00); H515(0.00); H129(0.00) | LDD0221 | [22] |
| LDCM0022 | KB02 | 22RV1 | C90(1.23); C49(1.65); C381(2.64); C573(1.61) | LDD2243 | [5] |
| LDCM0023 | KB03 | 22RV1 | C90(1.30); C49(1.82); C381(5.47); C573(2.73) | LDD2660 | [5] |
| LDCM0024 | KB05 | COLO792 | C381(2.78); C573(1.53); C511(1.24); C341(1.31) | LDD3310 | [5] |
| LDCM0109 | NEM | HeLa | H466(0.00); H541(0.00); H360(0.00); H440(0.00) | LDD0223 | [22] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C609(1.08) | LDD2106 | [6] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C457(0.93) | LDD2125 | [6] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C457(1.23); C511(0.73) | LDD2136 | [6] |
| LDCM0016 | Ranjitkar_cp1 | MDA-MB-231 | 2.63 | LDD0123 | [7] |
| LDCM0096 | SAHA | K562 | 5.71 | LDD0362 | [28] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Platelet endothelial cell adhesion molecule (PECAM1) | . | P16284 | |||
Other
References
