Details of the Target
General Information of Target
| Target ID | LDTP02435 | |||||
|---|---|---|---|---|---|---|
| Target Name | Heat shock 70 kDa protein 1B (HSPA1B) | |||||
| Gene Name | HSPA1B | |||||
| Gene ID | 3303 | |||||
| Synonyms |
HSP72; Heat shock 70 kDa protein 1B; Heat shock 70 kDa protein 2; HSP70-2; HSP70.2 |
|||||
| 3D Structure | ||||||
| Sequence |
MAKAAAIGIDLGTTYSCVGVFQHGKVEIIANDQGNRTTPSYVAFTDTERLIGDAAKNQVA
LNPQNTVFDAKRLIGRKFGDPVVQSDMKHWPFQVINDGDKPKVQVSYKGETKAFYPEEIS SMVLTKMKEIAEAYLGYPVTNAVITVPAYFNDSQRQATKDAGVIAGLNVLRIINEPTAAA IAYGLDRTGKGERNVLIFDLGGGTFDVSILTIDDGIFEVKATAGDTHLGGEDFDNRLVNH FVEEFKRKHKKDISQNKRAVRRLRTACERAKRTLSSSTQASLEIDSLFEGIDFYTSITRA RFEELCSDLFRSTLEPVEKALRDAKLDKAQIHDLVLVGGSTRIPKVQKLLQDFFNGRDLN KSINPDEAVAYGAAVQAAILMGDKSENVQDLLLLDVAPLSLGLETAGGVMTALIKRNSTI PTKQTQIFTTYSDNQPGVLIQVYEGERAMTKDNNLLGRFELSGIPPAPRGVPQIEVTFDI DANGILNVTATDKSTGKANKITITNDKGRLSKEEIERMVQEAEKYKAEDEVQRERVSAKN ALESYAFNMKSAVEDEGLKGKISEADKKKVLDKCQEVISWLDANTLAEKDEFEHKRKELE QVCNPIISGLYQGAGGPGPGGFGAQGPKGGSGSGPTIEEVD |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Heat shock protein 70 family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Molecular chaperone implicated in a wide variety of cellular processes, including protection of the proteome from stress, folding and transport of newly synthesized polypeptides, activation of proteolysis of misfolded proteins and the formation and dissociation of protein complexes. Plays a pivotal role in the protein quality control system, ensuring the correct folding of proteins, the re-folding of misfolded proteins and controlling the targeting of proteins for subsequent degradation. This is achieved through cycles of ATP binding, ATP hydrolysis and ADP release, mediated by co-chaperones. The co-chaperones have been shown to not only regulate different steps of the ATPase cycle, but they also have an individual specificity such that one co-chaperone may promote folding of a substrate while another may promote degradation. The affinity for polypeptides is regulated by its nucleotide bound state. In the ATP-bound form, it has a low affinity for substrate proteins. However, upon hydrolysis of the ATP to ADP, it undergoes a conformational change that increases its affinity for substrate proteins. It goes through repeated cycles of ATP hydrolysis and nucleotide exchange, which permits cycles of substrate binding and release. The co-chaperones are of three types: J-domain co-chaperones such as HSP40s (stimulate ATPase hydrolysis by HSP70), the nucleotide exchange factors (NEF) such as BAG1/2/3 (facilitate conversion of HSP70 from the ADP-bound to the ATP-bound state thereby promoting substrate release), and the TPR domain chaperones such as HOPX and STUB1. Maintains protein homeostasis during cellular stress through two opposing mechanisms: protein refolding and degradation. Its acetylation/deacetylation state determines whether it functions in protein refolding or protein degradation by controlling the competitive binding of co-chaperones HOPX and STUB1. During the early stress response, the acetylated form binds to HOPX which assists in chaperone-mediated protein refolding, thereafter, it is deacetylated and binds to ubiquitin ligase STUB1 that promotes ubiquitin-mediated protein degradation. Regulates centrosome integrity during mitosis, and is required for the maintenance of a functional mitotic centrosome that supports the assembly of a bipolar mitotic spindle. Enhances STUB1-mediated SMAD3 ubiquitination and degradation and facilitates STUB1-mediated inhibition of TGF-beta signaling. Essential for STUB1-mediated ubiquitination and degradation of FOXP3 in regulatory T-cells (Treg) during inflammation.; (Microbial infection) In case of rotavirus A infection, serves as a post-attachment receptor for the virus to facilitate entry into the cell.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P1 Probe Info |
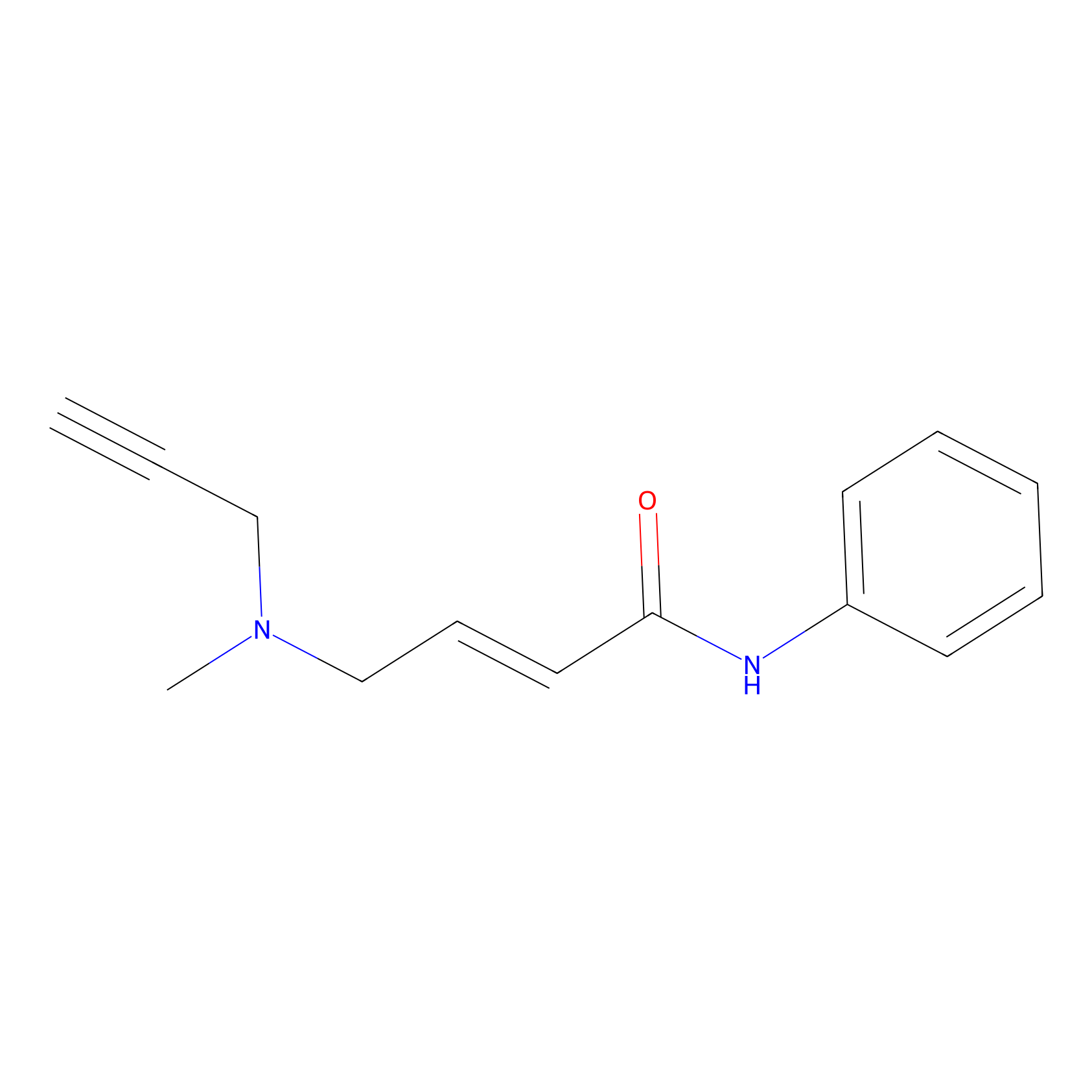 |
7.09 | LDD0448 | [1] | |
|
P2 Probe Info |
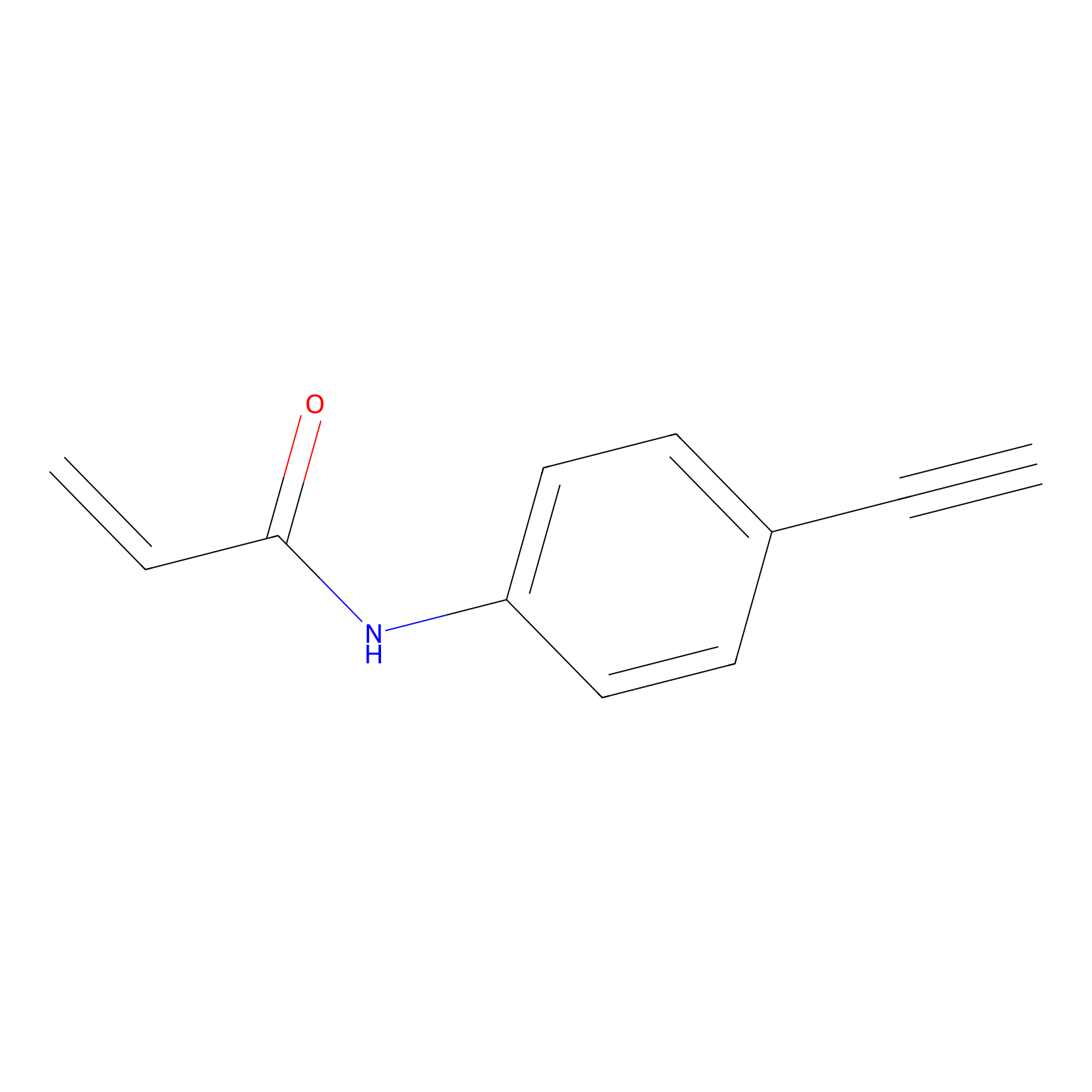 |
10.00 | LDD0449 | [1] | |
|
P3 Probe Info |
 |
10.00 | LDD0450 | [1] | |
|
P8 Probe Info |
 |
5.86 | LDD0451 | [1] | |
|
A-EBA Probe Info |
 |
4.46 | LDD0215 | [2] | |
|
11RK72 Probe Info |
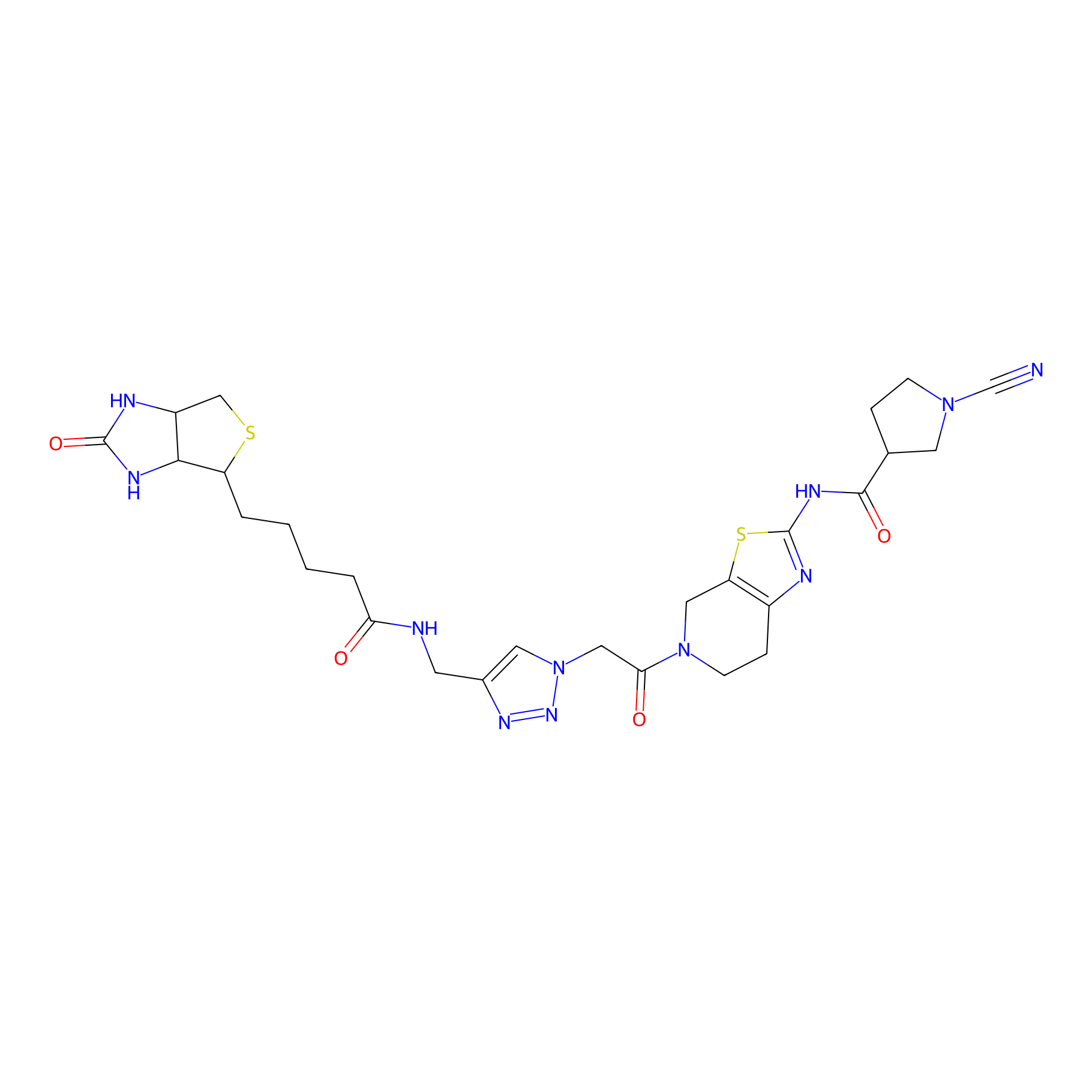 |
3.40 | LDD0327 | [3] | |
|
11RK73 Probe Info |
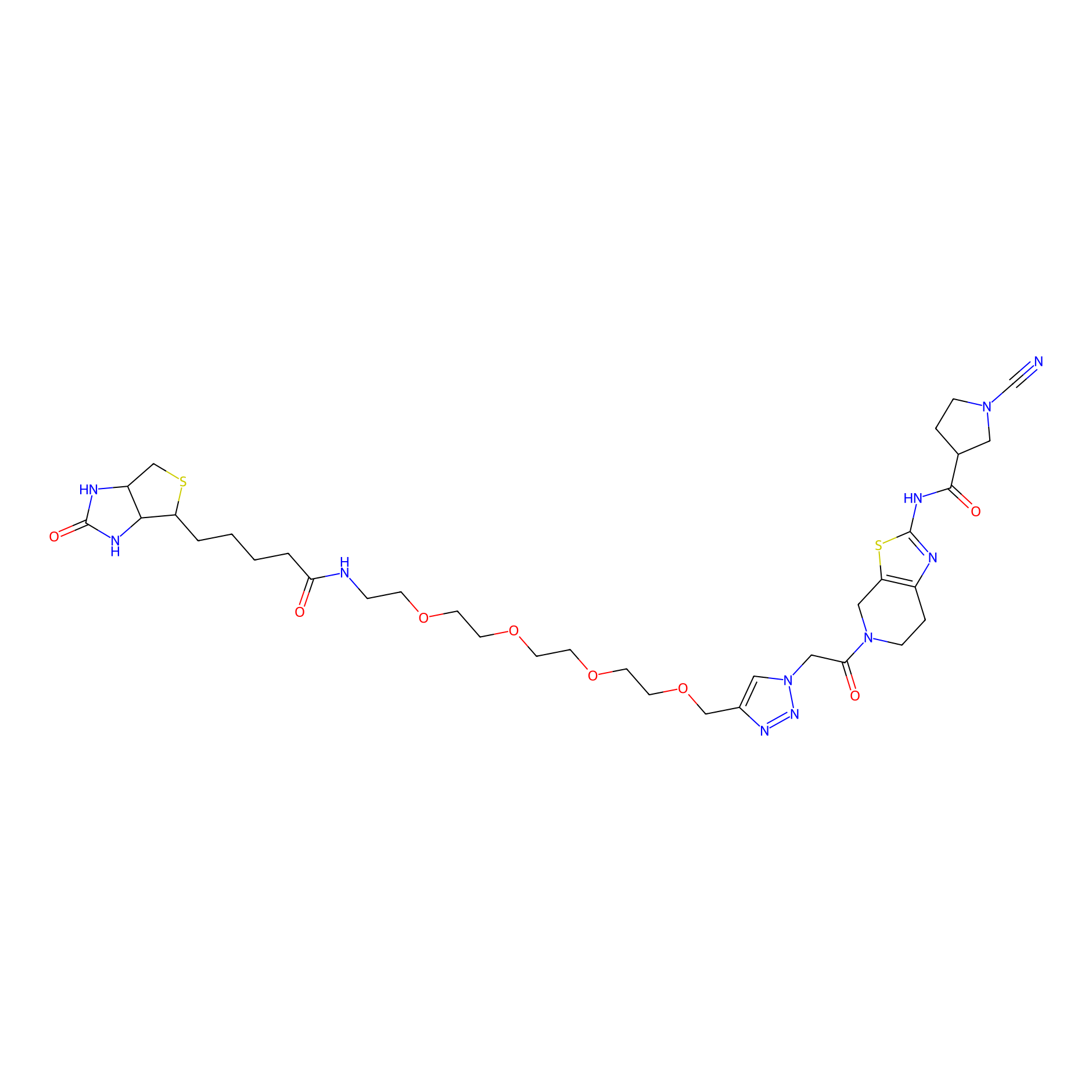 |
3.11 | LDD0328 | [3] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [4] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [4] | |
|
C-Sul Probe Info |
 |
2.71 | LDD0066 | [5] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [6] | |
|
AZ-9 Probe Info |
 |
E523(0.83); E315(1.00); E521(0.86); D160(0.70) | LDD2208 | [7] | |
|
ONAyne Probe Info |
 |
K524(0.00); K108(0.00); K77(0.00); K71(0.00) | LDD0273 | [8] | |
|
DBIA Probe Info |
 |
C17(1.65) | LDD3311 | [9] | |
|
HHS-475 Probe Info |
 |
Y545(0.74); Y15(0.82); Y431(0.87); Y525(0.90) | LDD0264 | [10] | |
|
AMP probe Probe Info |
 |
K507(0.00); K328(0.00); K524(0.00); K526(0.00) | LDD0200 | [11] | |
|
ATP probe Probe Info |
 |
K507(0.00); K328(0.00); K524(0.00); K526(0.00) | LDD0199 | [11] | |
|
IA-alkyne Probe Info |
 |
C306(0.00); C17(0.00); C574(0.00); C267(0.00) | LDD0032 | [12] | |
|
ATP probe Probe Info |
 |
K451(0.00); K348(0.00); K319(0.00); K88(0.00) | LDD0035 | [13] | |
|
IPM Probe Info |
 |
C306(0.00); C603(0.00) | LDD0025 | [14] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [14] | |
|
NAIA_5 Probe Info |
 |
C17(0.00); C306(0.00); C308(0.00) | LDD2224 | [15] | |
|
TFBX Probe Info |
 |
C17(0.00); C267(0.00) | LDD0027 | [14] | |
|
WYneN Probe Info |
 |
C603(0.00); C306(0.00) | LDD0021 | [16] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [16] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [16] | |
|
NHS Probe Info |
 |
K348(0.00); K451(0.00); K77(0.00); K328(0.00) | LDD0010 | [16] | |
|
OSF Probe Info |
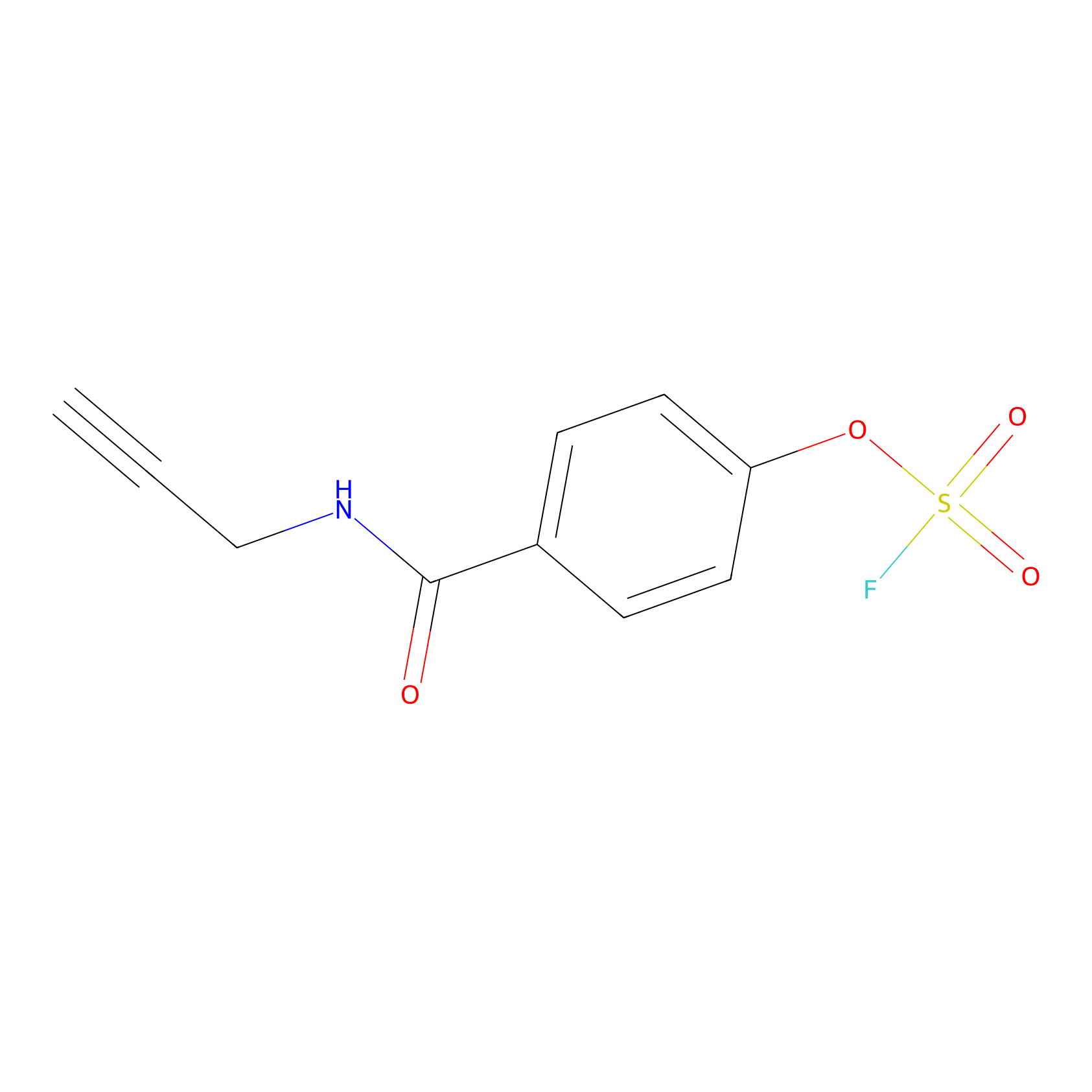 |
Y525(0.00); Y41(0.00); Y107(0.00) | LDD0029 | [17] | |
|
PPMS Probe Info |
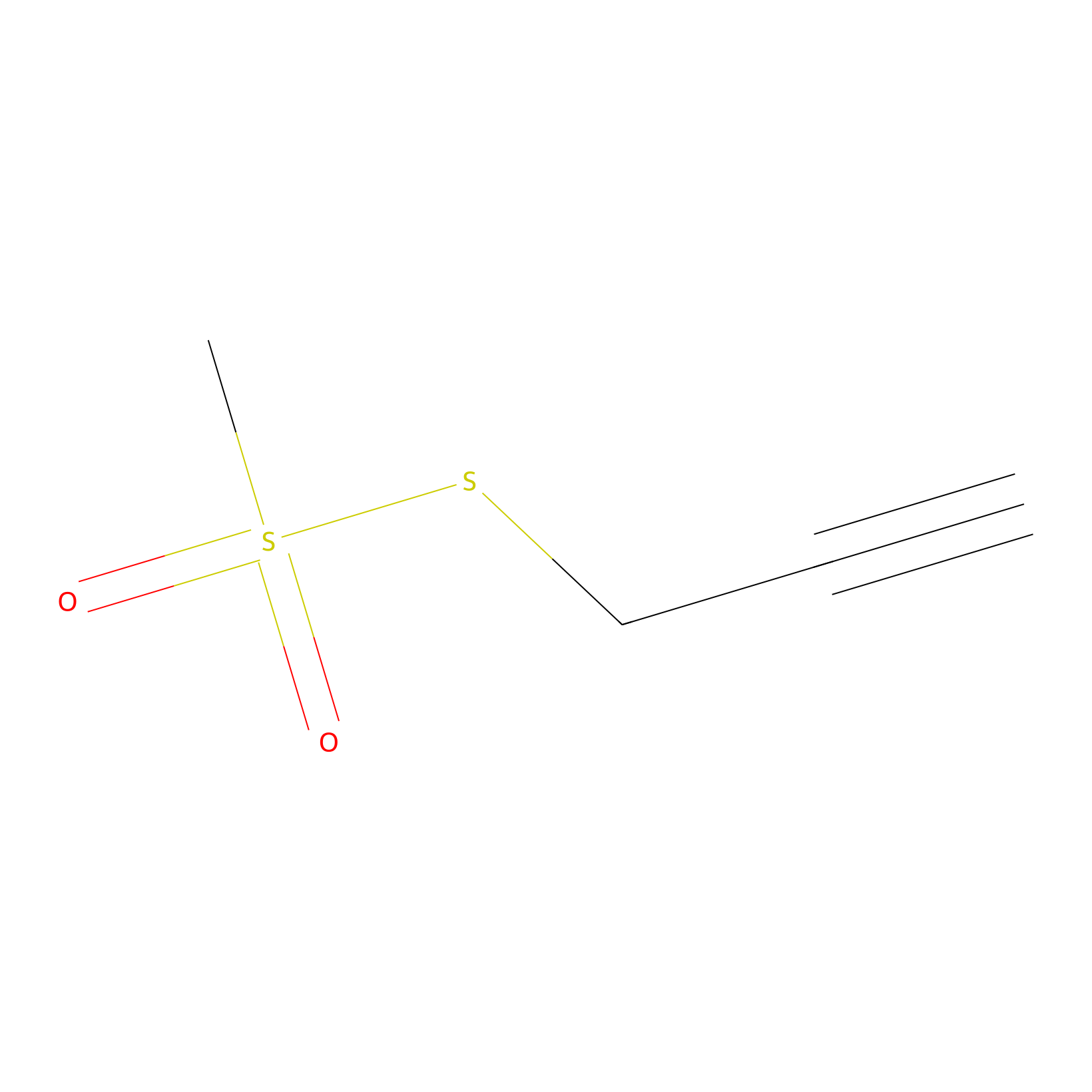 |
N.A. | LDD0008 | [16] | |
|
SF Probe Info |
 |
K348(0.00); Y41(0.00); K497(0.00); K524(0.00) | LDD0028 | [17] | |
|
STPyne Probe Info |
 |
K159(0.00); K328(0.00); K112(0.00); K319(0.00) | LDD0009 | [16] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [16] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [18] | |
|
Ox-W18 Probe Info |
 |
W90(0.00); W580(0.00) | LDD2175 | [19] | |
|
Acrolein Probe Info |
 |
C603(0.00); C306(0.00); H227(0.00); C574(0.00) | LDD0217 | [20] | |
|
Crotonaldehyde Probe Info |
 |
C306(0.00); H227(0.00); H332(0.00); H240(0.00) | LDD0219 | [20] | |
|
Methacrolein Probe Info |
 |
H332(0.00); C306(0.00); H240(0.00); H227(0.00) | LDD0218 | [20] | |
|
W1 Probe Info |
 |
C603(0.00); C306(0.00) | LDD0236 | [21] | |
|
MPP-AC Probe Info |
 |
N.A. | LDD0428 | [22] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [22] | |
|
HHS-465 Probe Info |
 |
K102(0.00); K108(0.00); K112(0.00); K159(0.00) | LDD2240 | [23] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Diazir Probe Info |
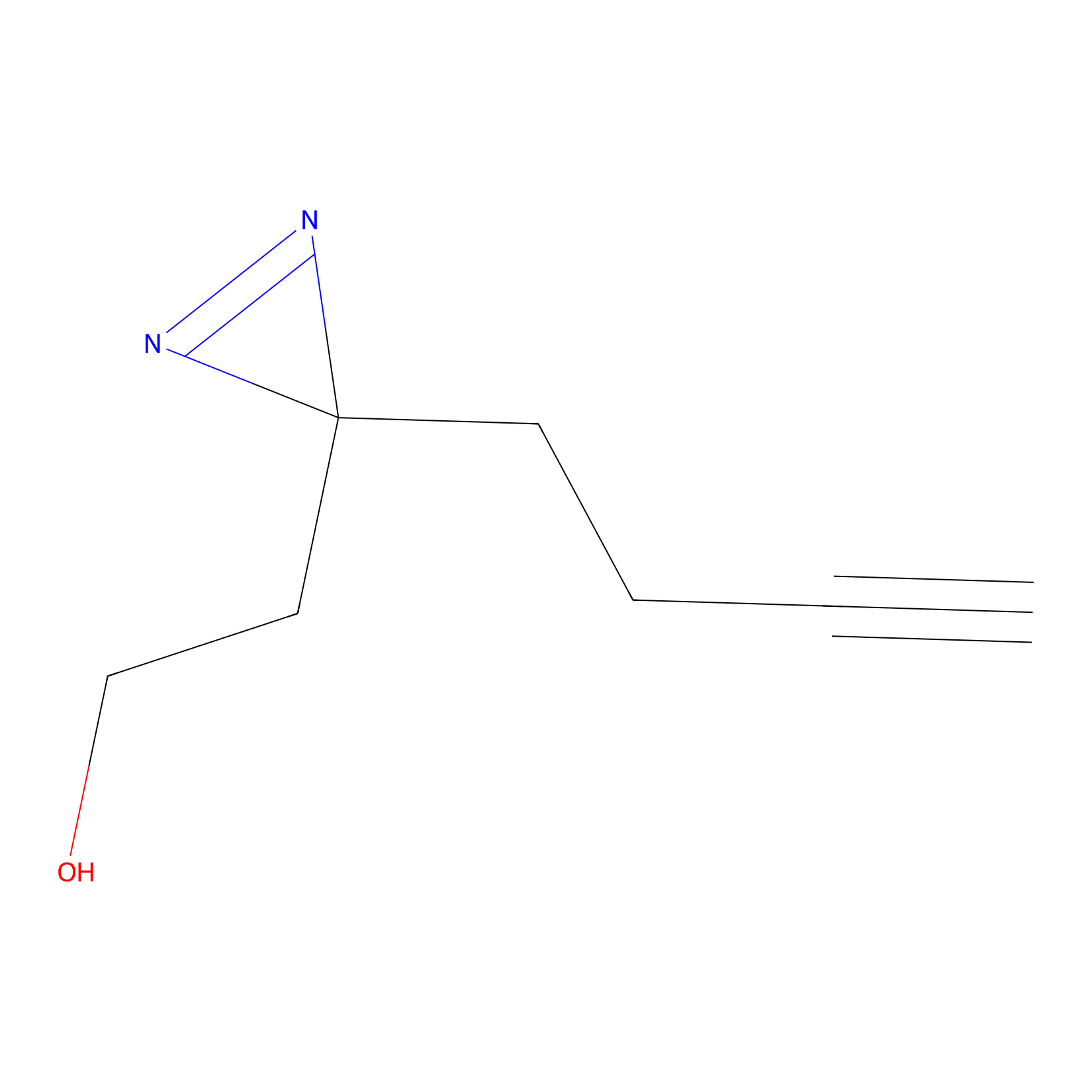 |
N.A. | LDD0011 | [16] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C17(1.00); C306(0.94); C603(0.85); C574(0.90) | LDD1507 | [24] |
| LDCM0215 | AC10 | HEK-293T | C17(0.91); C306(0.94); C603(0.84); C574(1.10) | LDD1508 | [24] |
| LDCM0226 | AC11 | HEK-293T | C17(0.98); C306(1.02); C603(0.90); C574(1.15) | LDD1509 | [24] |
| LDCM0237 | AC12 | HEK-293T | C17(0.87); C306(0.98); C603(1.03); C574(0.96) | LDD1510 | [24] |
| LDCM0259 | AC14 | HEK-293T | C17(0.99); C306(0.99); C603(0.97); C574(1.00) | LDD1512 | [24] |
| LDCM0270 | AC15 | HEK-293T | C17(0.96); C306(0.89); C603(1.07); C574(1.13) | LDD1513 | [24] |
| LDCM0276 | AC17 | HEK-293T | C17(0.91); C306(0.98); C603(0.83); C574(0.83) | LDD1515 | [24] |
| LDCM0277 | AC18 | HEK-293T | C17(1.06); C306(1.01); C603(0.86); C574(1.03) | LDD1516 | [24] |
| LDCM0278 | AC19 | HEK-293T | C17(0.98); C306(1.02); C603(0.72); C574(0.94) | LDD1517 | [24] |
| LDCM0279 | AC2 | HEK-293T | C17(1.00); C306(0.98); C603(0.88); C574(1.16) | LDD1518 | [24] |
| LDCM0280 | AC20 | HEK-293T | C17(0.98); C306(0.97); C603(0.92); C574(0.95) | LDD1519 | [24] |
| LDCM0281 | AC21 | HEK-293T | C17(1.02); C306(1.02); C603(0.93); C574(0.96) | LDD1520 | [24] |
| LDCM0282 | AC22 | HEK-293T | C17(1.03); C306(0.98); C603(0.98); C574(1.08) | LDD1521 | [24] |
| LDCM0283 | AC23 | HEK-293T | C17(0.93); C306(0.95); C603(1.08); C574(1.13) | LDD1522 | [24] |
| LDCM0284 | AC24 | HEK-293T | C17(0.89); C306(0.99); C603(0.88); C574(1.17) | LDD1523 | [24] |
| LDCM0285 | AC25 | HEK-293T | C17(0.97); C306(0.93); C603(0.93); C574(0.87) | LDD1524 | [24] |
| LDCM0286 | AC26 | HEK-293T | C17(1.01); C306(0.99); C603(1.01); C574(0.97) | LDD1525 | [24] |
| LDCM0287 | AC27 | HEK-293T | C17(1.06); C306(1.06); C603(0.86); C574(1.16) | LDD1526 | [24] |
| LDCM0288 | AC28 | HEK-293T | C17(1.01); C306(1.02); C603(0.98); C574(0.96) | LDD1527 | [24] |
| LDCM0289 | AC29 | HEK-293T | C17(1.09); C306(0.93); C603(0.96); C574(1.03) | LDD1528 | [24] |
| LDCM0290 | AC3 | HEK-293T | C17(1.05); C306(1.00); C603(0.95); C574(1.16) | LDD1529 | [24] |
| LDCM0291 | AC30 | HEK-293T | C17(1.02); C306(0.97); C603(0.93); C574(1.06) | LDD1530 | [24] |
| LDCM0292 | AC31 | HEK-293T | C17(1.01); C306(1.02); C603(1.00); C574(1.12) | LDD1531 | [24] |
| LDCM0293 | AC32 | HEK-293T | C17(0.92); C306(1.02); C603(0.91); C574(1.09) | LDD1532 | [24] |
| LDCM0294 | AC33 | HEK-293T | C17(0.92); C306(0.96); C603(0.81); C574(0.83) | LDD1533 | [24] |
| LDCM0295 | AC34 | HEK-293T | C17(0.86); C306(0.96); C603(0.88); C574(1.01) | LDD1534 | [24] |
| LDCM0296 | AC35 | HEK-293T | C17(1.00); C306(1.08); C603(0.84); C574(1.18) | LDD1535 | [24] |
| LDCM0297 | AC36 | HEK-293T | C17(0.96); C306(0.99); C603(0.99); C574(0.89) | LDD1536 | [24] |
| LDCM0298 | AC37 | HEK-293T | C17(0.99); C306(0.99); C603(1.15); C574(1.16) | LDD1537 | [24] |
| LDCM0299 | AC38 | HEK-293T | C17(0.98); C306(1.03); C603(1.06); C574(1.03) | LDD1538 | [24] |
| LDCM0300 | AC39 | HEK-293T | C17(0.98); C306(0.93); C603(1.04); C574(1.16) | LDD1539 | [24] |
| LDCM0301 | AC4 | HEK-293T | C17(0.97); C306(0.95); C603(0.93); C574(0.96) | LDD1540 | [24] |
| LDCM0302 | AC40 | HEK-293T | C17(1.02); C306(1.02); C603(0.89); C574(1.00) | LDD1541 | [24] |
| LDCM0303 | AC41 | HEK-293T | C17(0.86); C306(0.93); C603(0.79); C574(0.93) | LDD1542 | [24] |
| LDCM0304 | AC42 | HEK-293T | C17(0.93); C306(0.97); C603(1.07); C574(0.99) | LDD1543 | [24] |
| LDCM0305 | AC43 | HEK-293T | C17(1.01); C306(1.00); C603(0.88); C574(1.20) | LDD1544 | [24] |
| LDCM0306 | AC44 | HEK-293T | C17(0.91); C306(1.01); C603(0.94); C574(1.00) | LDD1545 | [24] |
| LDCM0307 | AC45 | HEK-293T | C17(1.02); C306(0.95); C603(0.86); C574(1.00) | LDD1546 | [24] |
| LDCM0308 | AC46 | HEK-293T | C17(1.02); C306(0.96); C603(0.96); C574(1.04) | LDD1547 | [24] |
| LDCM0309 | AC47 | HEK-293T | C17(0.97); C306(0.90); C603(1.14); C574(1.12) | LDD1548 | [24] |
| LDCM0310 | AC48 | HEK-293T | C17(1.01); C306(0.98); C603(1.12); C574(1.12) | LDD1549 | [24] |
| LDCM0311 | AC49 | HEK-293T | C17(0.90); C306(0.97); C603(0.82); C574(0.98) | LDD1550 | [24] |
| LDCM0312 | AC5 | HEK-293T | C17(1.02); C306(0.92); C603(1.02); C574(1.06) | LDD1551 | [24] |
| LDCM0313 | AC50 | HEK-293T | C17(0.96); C306(1.00); C603(0.93); C574(1.09) | LDD1552 | [24] |
| LDCM0314 | AC51 | HEK-293T | C17(0.95); C306(1.02); C603(1.07); C574(1.31) | LDD1553 | [24] |
| LDCM0315 | AC52 | HEK-293T | C17(0.90); C306(1.04); C603(0.91); C574(0.95) | LDD1554 | [24] |
| LDCM0316 | AC53 | HEK-293T | C17(0.94); C306(0.93); C603(0.99); C574(1.08) | LDD1555 | [24] |
| LDCM0317 | AC54 | HEK-293T | C17(0.97); C306(1.01); C603(1.02); C574(1.08) | LDD1556 | [24] |
| LDCM0318 | AC55 | HEK-293T | C17(0.93); C306(0.97); C603(1.08); C574(1.12) | LDD1557 | [24] |
| LDCM0319 | AC56 | HEK-293T | C17(0.95); C306(1.04); C603(0.90); C574(1.13) | LDD1558 | [24] |
| LDCM0320 | AC57 | HEK-293T | C17(0.92); C306(0.94); C603(0.89); C574(0.80) | LDD1559 | [24] |
| LDCM0321 | AC58 | HEK-293T | C17(0.89); C306(0.92); C603(0.95); C574(1.04) | LDD1560 | [24] |
| LDCM0322 | AC59 | HEK-293T | C17(1.01); C306(0.97); C603(0.98); C574(1.22) | LDD1561 | [24] |
| LDCM0323 | AC6 | HEK-293T | C17(0.99); C306(0.99); C603(0.96); C574(0.96) | LDD1562 | [24] |
| LDCM0324 | AC60 | HEK-293T | C17(0.91); C306(0.90); C603(0.94); C574(0.89) | LDD1563 | [24] |
| LDCM0325 | AC61 | HEK-293T | C17(1.00); C306(0.94); C603(0.97); C574(0.92) | LDD1564 | [24] |
| LDCM0326 | AC62 | HEK-293T | C17(0.96); C306(0.97); C603(0.91); C574(0.92) | LDD1565 | [24] |
| LDCM0327 | AC63 | HEK-293T | C17(0.93); C306(0.92); C603(0.91); C574(1.02) | LDD1566 | [24] |
| LDCM0328 | AC64 | HEK-293T | C17(0.97); C306(0.94); C603(1.00); C574(1.01) | LDD1567 | [24] |
| LDCM0334 | AC7 | HEK-293T | C17(1.00); C306(0.94); C603(1.06); C574(1.11) | LDD1568 | [24] |
| LDCM0345 | AC8 | HEK-293T | C17(1.00); C306(1.01); C603(0.91); C574(0.92) | LDD1569 | [24] |
| LDCM0248 | AKOS034007472 | HEK-293T | C17(1.01); C306(0.96); C603(1.09); C574(1.09) | LDD1511 | [24] |
| LDCM0356 | AKOS034007680 | HEK-293T | C17(0.95); C306(0.93); C603(0.89); C574(0.84) | LDD1570 | [24] |
| LDCM0275 | AKOS034007705 | HEK-293T | C17(1.03); C306(0.97); C603(1.02); C574(1.03) | LDD1514 | [24] |
| LDCM0108 | Chloroacetamide | HeLa | C603(0.00); C574(0.00); H227(0.00); H332(0.00) | LDD0222 | [20] |
| LDCM0367 | CL1 | HEK-293T | C17(1.05); C306(0.96); C603(1.03); C574(0.96) | LDD1571 | [24] |
| LDCM0368 | CL10 | HEK-293T | C17(1.01); C306(1.16); C603(0.73); C574(0.70) | LDD1572 | [24] |
| LDCM0369 | CL100 | HEK-293T | C17(0.99); C306(0.93); C603(0.79); C574(0.94) | LDD1573 | [24] |
| LDCM0370 | CL101 | HEK-293T | C17(0.96); C306(0.94); C603(0.98); C574(0.88) | LDD1574 | [24] |
| LDCM0371 | CL102 | HEK-293T | C17(0.94); C306(0.95); C603(0.80); C574(0.93) | LDD1575 | [24] |
| LDCM0372 | CL103 | HEK-293T | C17(0.93); C306(0.95); C603(0.96); C574(1.01) | LDD1576 | [24] |
| LDCM0373 | CL104 | HEK-293T | C17(0.98); C306(0.99); C603(0.94); C574(1.00) | LDD1577 | [24] |
| LDCM0374 | CL105 | HEK-293T | C17(0.94); C306(1.04); C603(1.05); C574(0.91) | LDD1578 | [24] |
| LDCM0375 | CL106 | HEK-293T | C17(1.03); C306(1.00); C603(1.10); C574(0.97) | LDD1579 | [24] |
| LDCM0376 | CL107 | HEK-293T | C17(1.03); C306(1.00); C603(0.97); C574(1.11) | LDD1580 | [24] |
| LDCM0377 | CL108 | HEK-293T | C17(0.92); C306(0.97); C603(0.97); C574(0.92) | LDD1581 | [24] |
| LDCM0378 | CL109 | HEK-293T | C17(0.99); C306(0.98); C603(1.03); C574(0.89) | LDD1582 | [24] |
| LDCM0379 | CL11 | HEK-293T | C17(1.06); C306(1.05); C603(1.00); C574(0.62) | LDD1583 | [24] |
| LDCM0380 | CL110 | HEK-293T | C17(0.93); C306(0.91); C603(1.01); C574(0.88) | LDD1584 | [24] |
| LDCM0381 | CL111 | HEK-293T | C17(0.97); C306(1.00); C603(0.94); C574(0.90) | LDD1585 | [24] |
| LDCM0382 | CL112 | HEK-293T | C17(0.95); C306(0.95); C603(0.86); C574(1.02) | LDD1586 | [24] |
| LDCM0383 | CL113 | HEK-293T | C17(0.98); C306(0.92); C603(1.04); C574(0.96) | LDD1587 | [24] |
| LDCM0384 | CL114 | HEK-293T | C17(0.88); C306(1.00); C603(0.84); C574(0.87) | LDD1588 | [24] |
| LDCM0385 | CL115 | HEK-293T | C17(1.02); C306(0.98); C603(1.06); C574(0.97) | LDD1589 | [24] |
| LDCM0386 | CL116 | HEK-293T | C17(0.88); C306(0.98); C603(0.86); C574(0.91) | LDD1590 | [24] |
| LDCM0387 | CL117 | HEK-293T | C17(0.97); C306(0.94); C603(0.92); C574(0.91) | LDD1591 | [24] |
| LDCM0388 | CL118 | HEK-293T | C17(0.93); C306(0.98); C603(0.96); C574(1.00) | LDD1592 | [24] |
| LDCM0389 | CL119 | HEK-293T | C17(1.00); C306(0.94); C603(0.98); C574(1.03) | LDD1593 | [24] |
| LDCM0390 | CL12 | HEK-293T | C17(1.02); C306(1.07); C603(0.89); C574(0.90) | LDD1594 | [24] |
| LDCM0391 | CL120 | HEK-293T | C17(0.90); C306(0.97); C603(0.97); C574(0.93) | LDD1595 | [24] |
| LDCM0392 | CL121 | HEK-293T | C17(0.93); C306(1.01); C603(0.95); C574(0.97) | LDD1596 | [24] |
| LDCM0393 | CL122 | HEK-293T | C17(0.93); C306(1.00); C603(0.98); C574(1.02) | LDD1597 | [24] |
| LDCM0394 | CL123 | HEK-293T | C17(0.90); C306(0.95); C603(0.84); C574(0.89) | LDD1598 | [24] |
| LDCM0395 | CL124 | HEK-293T | C17(0.88); C306(0.97); C603(0.86); C574(0.93) | LDD1599 | [24] |
| LDCM0396 | CL125 | HEK-293T | C17(1.00); C306(0.91); C603(1.05); C574(0.99) | LDD1600 | [24] |
| LDCM0397 | CL126 | HEK-293T | C17(0.97); C306(0.94); C603(0.90); C574(0.89) | LDD1601 | [24] |
| LDCM0398 | CL127 | HEK-293T | C17(1.01); C306(0.93); C603(0.96); C574(0.90) | LDD1602 | [24] |
| LDCM0399 | CL128 | HEK-293T | C17(1.01); C306(0.92); C603(0.89); C574(0.92) | LDD1603 | [24] |
| LDCM0400 | CL13 | HEK-293T | C17(0.95); C306(0.88); C603(1.13); C574(1.03) | LDD1604 | [24] |
| LDCM0401 | CL14 | HEK-293T | C17(1.03); C306(1.02); C603(0.94); C574(1.05) | LDD1605 | [24] |
| LDCM0402 | CL15 | HEK-293T | C17(0.95); C306(0.87); C603(0.95); C574(0.98) | LDD1606 | [24] |
| LDCM0403 | CL16 | HEK-293T | C17(0.91); C306(1.00); C603(0.88); C574(0.96) | LDD1607 | [24] |
| LDCM0404 | CL17 | HEK-293T | C17(0.91); C306(0.96); C603(0.89); C574(0.82) | LDD1608 | [24] |
| LDCM0405 | CL18 | HEK-293T | C17(1.02); C306(1.05); C603(0.87); C574(1.06) | LDD1609 | [24] |
| LDCM0406 | CL19 | HEK-293T | C17(1.04); C306(1.09); C603(1.06); C574(0.96) | LDD1610 | [24] |
| LDCM0407 | CL2 | HEK-293T | C17(1.05); C306(1.08); C603(1.01); C574(1.02) | LDD1611 | [24] |
| LDCM0408 | CL20 | HEK-293T | C17(0.89); C306(1.15); C603(1.05); C574(1.03) | LDD1612 | [24] |
| LDCM0409 | CL21 | HEK-293T | C17(0.94); C306(1.00); C603(0.77); C574(0.74) | LDD1613 | [24] |
| LDCM0410 | CL22 | HEK-293T | C17(1.06); C306(1.18); C603(0.95); C574(0.96) | LDD1614 | [24] |
| LDCM0411 | CL23 | HEK-293T | C17(1.01); C306(1.09); C603(1.16); C574(0.76) | LDD1615 | [24] |
| LDCM0412 | CL24 | HEK-293T | C17(0.98); C306(1.07); C603(0.93); C574(1.04) | LDD1616 | [24] |
| LDCM0413 | CL25 | HEK-293T | C17(1.01); C306(0.93); C603(0.86); C574(0.98) | LDD1617 | [24] |
| LDCM0414 | CL26 | HEK-293T | C17(1.00); C306(1.02); C603(1.06); C574(1.05) | LDD1618 | [24] |
| LDCM0415 | CL27 | HEK-293T | C17(0.99); C306(0.99); C603(1.03); C574(1.07) | LDD1619 | [24] |
| LDCM0416 | CL28 | HEK-293T | C17(0.90); C306(0.97); C603(1.05); C574(1.04) | LDD1620 | [24] |
| LDCM0417 | CL29 | HEK-293T | C17(0.97); C306(0.95); C603(0.99); C574(0.94) | LDD1621 | [24] |
| LDCM0418 | CL3 | HEK-293T | C17(1.09); C306(1.02); C603(1.01); C574(0.98) | LDD1622 | [24] |
| LDCM0419 | CL30 | HEK-293T | C17(1.01); C306(0.99); C603(1.19); C574(1.02) | LDD1623 | [24] |
| LDCM0420 | CL31 | HEK-293T | C17(1.08); C306(1.10); C603(0.87); C574(0.94) | LDD1624 | [24] |
| LDCM0421 | CL32 | HEK-293T | C17(0.98); C306(1.13); C603(0.96); C574(1.03) | LDD1625 | [24] |
| LDCM0422 | CL33 | HEK-293T | C17(0.95); C306(1.04); C603(0.84); C574(0.72) | LDD1626 | [24] |
| LDCM0423 | CL34 | HEK-293T | C17(1.04); C306(1.16); C603(0.98); C574(0.85) | LDD1627 | [24] |
| LDCM0424 | CL35 | HEK-293T | C17(0.98); C306(1.07); C603(0.99); C574(0.72) | LDD1628 | [24] |
| LDCM0425 | CL36 | HEK-293T | C17(1.05); C306(1.07); C603(1.01); C574(1.03) | LDD1629 | [24] |
| LDCM0426 | CL37 | HEK-293T | C17(1.03); C306(0.93); C603(1.06); C574(0.99) | LDD1630 | [24] |
| LDCM0428 | CL39 | HEK-293T | C17(0.98); C306(0.95); C603(1.13); C574(1.04) | LDD1632 | [24] |
| LDCM0429 | CL4 | HEK-293T | C17(0.91); C306(0.95); C603(1.10); C574(0.87) | LDD1633 | [24] |
| LDCM0430 | CL40 | HEK-293T | C17(0.95); C306(0.95); C603(1.00); C574(0.91) | LDD1634 | [24] |
| LDCM0431 | CL41 | HEK-293T | C17(1.02); C306(0.94); C603(0.83); C574(0.85) | LDD1635 | [24] |
| LDCM0432 | CL42 | HEK-293T | C17(0.99); C306(1.00); C603(0.88); C574(0.96) | LDD1636 | [24] |
| LDCM0433 | CL43 | HEK-293T | C17(0.99); C306(1.03); C603(0.92); C574(0.92) | LDD1637 | [24] |
| LDCM0434 | CL44 | HEK-293T | C17(0.91); C306(1.00); C603(1.00); C574(0.92) | LDD1638 | [24] |
| LDCM0435 | CL45 | HEK-293T | C17(1.01); C306(1.04); C603(0.86); C574(0.75) | LDD1639 | [24] |
| LDCM0436 | CL46 | HEK-293T | C17(1.00); C306(1.08); C603(0.89); C574(0.89) | LDD1640 | [24] |
| LDCM0437 | CL47 | HEK-293T | C17(1.05); C306(0.99); C603(1.02); C574(0.70) | LDD1641 | [24] |
| LDCM0438 | CL48 | HEK-293T | C17(1.04); C306(0.97); C603(1.00); C574(1.02) | LDD1642 | [24] |
| LDCM0439 | CL49 | HEK-293T | C17(1.03); C306(0.95); C603(1.19); C574(0.99) | LDD1643 | [24] |
| LDCM0440 | CL5 | HEK-293T | C17(0.94); C306(0.97); C603(0.92); C574(0.94) | LDD1644 | [24] |
| LDCM0441 | CL50 | HEK-293T | C17(0.98); C306(0.99); C603(0.93); C574(1.03) | LDD1645 | [24] |
| LDCM0443 | CL52 | HEK-293T | C17(0.87); C306(1.01); C603(1.05); C574(0.92) | LDD1646 | [24] |
| LDCM0444 | CL53 | HEK-293T | C17(0.95); C306(0.88); C603(0.89); C574(0.77) | LDD1647 | [24] |
| LDCM0445 | CL54 | HEK-293T | C17(0.90); C306(1.01); C603(0.82); C574(0.89) | LDD1648 | [24] |
| LDCM0446 | CL55 | HEK-293T | C17(1.02); C306(1.08); C603(0.87); C574(0.81) | LDD1649 | [24] |
| LDCM0447 | CL56 | HEK-293T | C17(1.00); C306(1.05); C603(0.87); C574(0.93) | LDD1650 | [24] |
| LDCM0448 | CL57 | HEK-293T | C17(0.98); C306(1.03); C603(0.77); C574(0.76) | LDD1651 | [24] |
| LDCM0449 | CL58 | HEK-293T | C17(0.99); C306(1.11); C603(0.83); C574(0.90) | LDD1652 | [24] |
| LDCM0450 | CL59 | HEK-293T | C17(0.95); C306(1.06); C603(1.21); C574(0.62) | LDD1653 | [24] |
| LDCM0451 | CL6 | HEK-293T | C17(0.97); C306(1.11); C603(0.97); C574(0.96) | LDD1654 | [24] |
| LDCM0452 | CL60 | HEK-293T | C17(1.03); C306(1.18); C603(0.85); C574(0.99) | LDD1655 | [24] |
| LDCM0453 | CL61 | HEK-293T | C17(1.03); C306(0.92); C603(1.15); C574(0.99) | LDD1656 | [24] |
| LDCM0454 | CL62 | HEK-293T | C17(0.99); C306(0.96); C603(0.94); C574(1.11) | LDD1657 | [24] |
| LDCM0455 | CL63 | HEK-293T | C17(0.96); C306(0.99); C603(1.04); C574(1.09) | LDD1658 | [24] |
| LDCM0456 | CL64 | HEK-293T | C17(1.02); C306(1.00); C603(0.87); C574(0.83) | LDD1659 | [24] |
| LDCM0457 | CL65 | HEK-293T | C17(0.95); C306(0.95); C603(0.85); C574(0.89) | LDD1660 | [24] |
| LDCM0458 | CL66 | HEK-293T | C17(0.95); C306(0.99); C603(0.99); C574(1.01) | LDD1661 | [24] |
| LDCM0459 | CL67 | HEK-293T | C17(1.00); C306(1.04); C603(0.88); C574(1.00) | LDD1662 | [24] |
| LDCM0460 | CL68 | HEK-293T | C17(0.93); C306(1.03); C603(0.98); C574(0.99) | LDD1663 | [24] |
| LDCM0461 | CL69 | HEK-293T | C17(1.04); C306(0.95); C603(0.93); C574(0.84) | LDD1664 | [24] |
| LDCM0462 | CL7 | HEK-293T | C17(1.06); C306(1.14); C603(1.01); C574(0.95) | LDD1665 | [24] |
| LDCM0463 | CL70 | HEK-293T | C17(1.00); C306(1.09); C603(0.91); C574(0.84) | LDD1666 | [24] |
| LDCM0464 | CL71 | HEK-293T | C17(1.02); C306(1.00); C603(1.12); C574(0.80) | LDD1667 | [24] |
| LDCM0465 | CL72 | HEK-293T | C17(0.98); C306(1.13); C603(0.93); C574(0.95) | LDD1668 | [24] |
| LDCM0466 | CL73 | HEK-293T | C17(0.99); C306(1.00); C603(0.89); C574(1.11) | LDD1669 | [24] |
| LDCM0467 | CL74 | HEK-293T | C17(0.99); C306(1.00); C603(0.86); C574(1.09) | LDD1670 | [24] |
| LDCM0469 | CL76 | HEK-293T | C17(0.94); C306(0.96); C603(1.03); C574(1.02) | LDD1672 | [24] |
| LDCM0470 | CL77 | HEK-293T | C17(0.94); C306(0.94); C603(0.76); C574(0.91) | LDD1673 | [24] |
| LDCM0471 | CL78 | HEK-293T | C17(1.07); C306(1.11); C603(1.07); C574(1.12) | LDD1674 | [24] |
| LDCM0472 | CL79 | HEK-293T | C17(0.98); C306(1.09); C603(0.94); C574(1.03) | LDD1675 | [24] |
| LDCM0473 | CL8 | HEK-293T | C17(0.87); C306(0.91); C603(0.69); C574(0.69) | LDD1676 | [24] |
| LDCM0474 | CL80 | HEK-293T | C17(1.04); C306(1.02); C603(1.20); C574(1.13) | LDD1677 | [24] |
| LDCM0475 | CL81 | HEK-293T | C17(1.03); C306(1.09); C603(1.03); C574(0.91) | LDD1678 | [24] |
| LDCM0476 | CL82 | HEK-293T | C17(1.06); C306(1.10); C603(0.98); C574(0.97) | LDD1679 | [24] |
| LDCM0477 | CL83 | HEK-293T | C17(1.05); C306(1.17); C603(1.04); C574(0.74) | LDD1680 | [24] |
| LDCM0478 | CL84 | HEK-293T | C17(1.01); C306(0.98); C603(1.03); C574(0.95) | LDD1681 | [24] |
| LDCM0479 | CL85 | HEK-293T | C17(1.04); C306(0.89); C603(1.04); C574(0.97) | LDD1682 | [24] |
| LDCM0480 | CL86 | HEK-293T | C17(0.97); C306(0.95); C603(1.06); C574(0.91) | LDD1683 | [24] |
| LDCM0481 | CL87 | HEK-293T | C17(1.01); C306(0.85); C603(0.99); C574(0.99) | LDD1684 | [24] |
| LDCM0482 | CL88 | HEK-293T | C17(0.77); C306(0.93); C603(0.91); C574(0.93) | LDD1685 | [24] |
| LDCM0483 | CL89 | HEK-293T | C17(0.94); C306(0.95); C603(1.08); C574(0.84) | LDD1686 | [24] |
| LDCM0484 | CL9 | HEK-293T | C17(1.04); C306(1.08); C603(0.99); C574(0.84) | LDD1687 | [24] |
| LDCM0485 | CL90 | HEK-293T | C17(0.74); C306(0.99); C603(0.68); C574(0.82) | LDD1688 | [24] |
| LDCM0486 | CL91 | HEK-293T | C17(1.07); C306(1.06); C603(1.02); C574(1.07) | LDD1689 | [24] |
| LDCM0487 | CL92 | HEK-293T | C17(0.91); C306(0.97); C603(0.90); C574(0.88) | LDD1690 | [24] |
| LDCM0488 | CL93 | HEK-293T | C17(1.01); C306(1.00); C603(0.91); C574(0.87) | LDD1691 | [24] |
| LDCM0489 | CL94 | HEK-293T | C17(1.06); C306(1.03); C603(0.88); C574(0.88) | LDD1692 | [24] |
| LDCM0490 | CL95 | HEK-293T | C17(0.87); C306(0.87); C603(0.92); C574(0.65) | LDD1693 | [24] |
| LDCM0491 | CL96 | HEK-293T | C17(0.98); C306(0.90); C603(0.91); C574(0.87) | LDD1694 | [24] |
| LDCM0492 | CL97 | HEK-293T | C17(0.99); C306(0.98); C603(0.91); C574(0.83) | LDD1695 | [24] |
| LDCM0493 | CL98 | HEK-293T | C17(0.99); C306(0.94); C603(0.92); C574(0.94) | LDD1696 | [24] |
| LDCM0494 | CL99 | HEK-293T | C17(0.94); C306(0.94); C603(0.90); C574(0.93) | LDD1697 | [24] |
| LDCM0495 | E2913 | HEK-293T | C17(0.98); C306(0.98); C603(0.99); C574(1.04) | LDD1698 | [24] |
| LDCM0468 | Fragment33 | HEK-293T | C17(1.02); C306(0.96); C603(0.96); C574(1.05) | LDD1671 | [24] |
| LDCM0427 | Fragment51 | HEK-293T | C17(0.96); C306(0.96); C603(0.84); C574(1.05) | LDD1631 | [24] |
| LDCM0116 | HHS-0101 | DM93 | Y545(0.74); Y15(0.82); Y431(0.87); Y525(0.90) | LDD0264 | [10] |
| LDCM0117 | HHS-0201 | DM93 | Y545(0.45); Y137(0.63); Y15(0.64); Y525(0.69) | LDD0265 | [10] |
| LDCM0118 | HHS-0301 | DM93 | Y545(0.55); Y525(0.71); Y15(0.80); Y431(0.81) | LDD0266 | [10] |
| LDCM0119 | HHS-0401 | DM93 | Y15(0.67); Y525(0.75); Y41(0.75); Y545(0.78) | LDD0267 | [10] |
| LDCM0120 | HHS-0701 | DM93 | Y41(0.48); Y525(0.59); Y115(0.63); Y107(0.71) | LDD0268 | [10] |
| LDCM0107 | IAA | HeLa | C603(0.00); H227(0.00); H332(0.00); H89(0.00) | LDD0221 | [20] |
| LDCM0022 | KB02 | HEK-293T | C17(1.06); C306(0.92); C603(1.30); C574(1.04) | LDD1492 | [24] |
| LDCM0023 | KB03 | HEK-293T | C17(1.03); C306(1.00); C603(1.20); C574(1.02) | LDD1497 | [24] |
| LDCM0024 | KB05 | G361 | C17(1.65) | LDD3311 | [9] |
| LDCM0109 | NEM | HeLa | H332(0.00); H227(0.00); H240(0.00); C306(0.00) | LDD0223 | [20] |
The Interaction Atlas With This Target
References
