Details of the Target
General Information of Target
| Target ID | LDTP12056 | |||||
|---|---|---|---|---|---|---|
| Target Name | Thiol S-methyltransferase TMT1A (TMT1A) | |||||
| Gene Name | TMT1A | |||||
| Gene ID | 25840 | |||||
| Synonyms |
METTL7A; Thiol S-methyltransferase TMT1A; EC 2.1.1.9; Methyltransferase-like protein 7A; N6-adenosine-methyltransferase TMT1A; EC 2.1.1.348; Protein AAM-B; Thiol methyltransferase 1A |
|||||
| 3D Structure | ||||||
| Sequence |
MAPDLASQRHSESFPSVNSRPNVILPGREGRREGLPPGGGTRGSLVPTRPVPPSPAPLGT
SPYSWSRSGPGRGGGAGSSRVPRGVPGPAVCAPGSLLHHASPTQTMAAADGSLFDNPRTF SRRPPAQASRQAKATKRKYQASSEAPPAKRRNETSFLPAKKTSVKETQRTFKGNAQKMFS PKKHSVSTSDRNQEERQCIKTSSLFKNNPDIPELHRPVVKQVQEKVFTSAAFHELGLHPH LISTINTVLKMSSMTSVQKQSIPVLLEGRDALVRSQTGSGKTLAYCIPVVQSLQAMESKI QRSDGPYALVLVPTRELALQSFDTVQKLLKPFTWIVPGVLMGGEKRKSEKARLRKGINIL ISTPGRLVDHIKSTKNIHFSRLRWLVFDEADRILDLGFEKDITVILNAVNAECQKRQNVL LSATLTEGVTRLADISLHDPVSISVLDKSHDQLNPKDKAVQEVCPPPAGDKLDSFAIPES LKQHVTVVPSKLRLVCLAAFILQKCKFEEDQKMVVFFSSCELVEFHYSLFLQTLLSSSGA PASGQLPSASMRLKFLRLHGGMEQEERTAVFQEFSHSRRGVLLCTDVAARGLDLPQVTWI VQYNAPSSPAEYIHRIGRTARIGCHGSSLLILAPSEAEYVNSLASHKINVSEIKMEDILC VLTRDDCFKGKRWGAQKSHAVGPQEIRERATVLQTVFEDYVHSSERRVSWAKKALQSFIQ AYATYPRELKHIFHVRSLHLGHVAKSFGLRDAPRNLSALTRKKRKAHVKRPDLHKKTQSK HSLAEILRSEYSSGMEADIAKVKKQNAPGEPGGRPLQHSLQPTPCFGRGKTLKWRKTQKG VQRDSKTSQKV |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Methyltransferase superfamily
|
|||||
| Subcellular location |
Lipid droplet
|
|||||
| Function |
Thiol S-methyltransferase that catalyzes the transfer of a methyl group from S-adenosyl-L-methionine to alkyl and phenolic thiol-containing acceptor substrates. Together with TMT1B accounts for most of S-thiol methylation activity in the endoplasmic reticulum of hepatocytes. Able to methylate the N6 position of adenosine residues in long non-coding RNAs (lncRNAs). May facilitate lncRNAs transfer into exosomes at the tumor-stroma interface. Promotes osteogenic and odontogenic differentiation by regulating the expression of genes involved in stem cell differentiation and survival. Targeted from the endoplasmic reticulum to lipid droplets, where it recruits cellular proteins to form functional organelles.; (Microbial infection) May be involved in the assembly and release stages of hepatitis C virus (HCV) life cycle and thus play a crucial role in HCV propagation.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [1] | |
|
STPyne Probe Info |
 |
K105(7.78); K109(5.72) | LDD0277 | [2] | |
|
OPA-S-S-alkyne Probe Info |
 |
K210(1.36) | LDD3494 | [3] | |
|
P11 Probe Info |
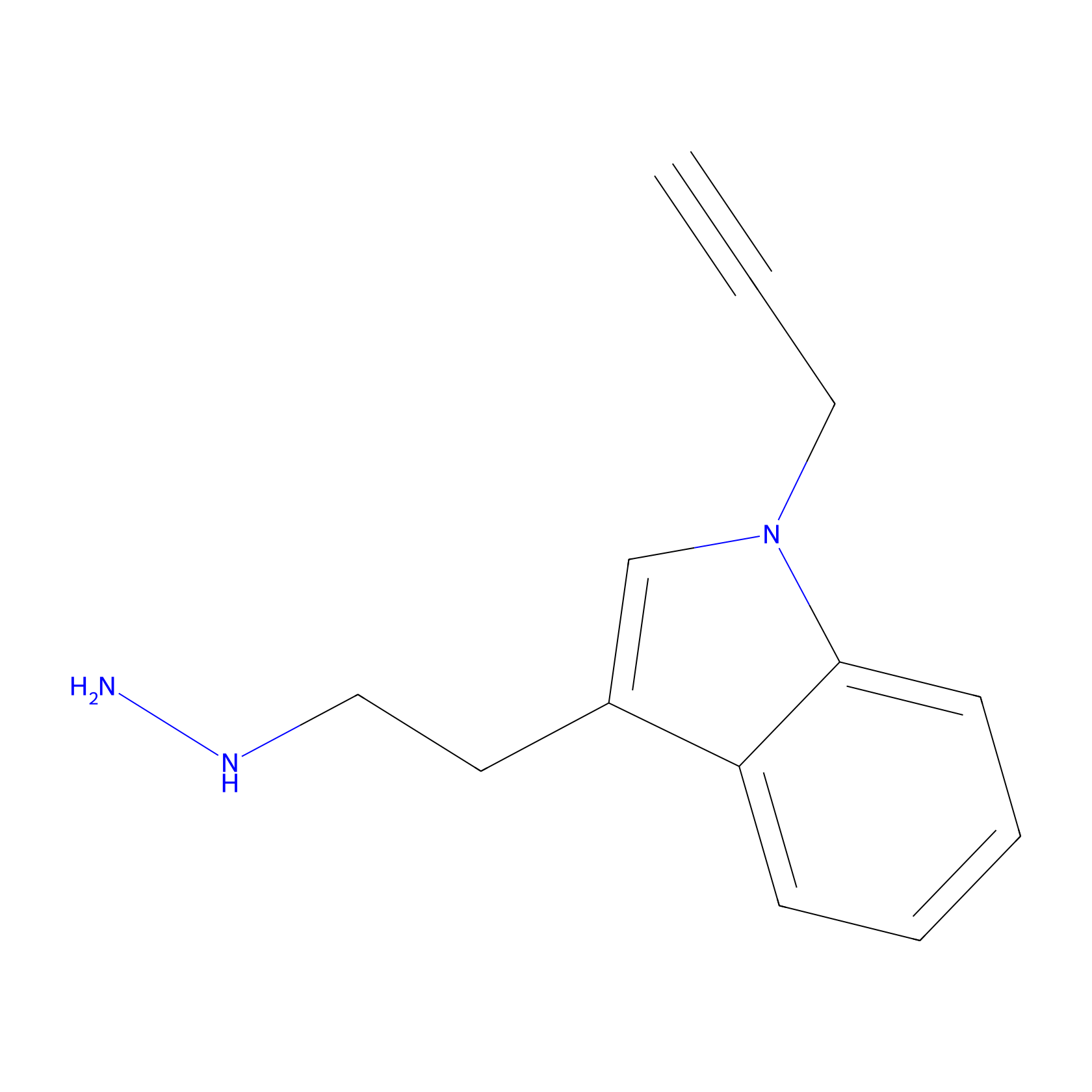 |
20.00 | LDD0201 | [4] | |
|
DBIA Probe Info |
 |
C79(1.48) | LDD0080 | [5] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [6] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [7] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [8] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [8] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [9] | |
|
Acrolein Probe Info |
 |
C79(0.00); C96(0.00) | LDD0217 | [10] | |
|
Crotonaldehyde Probe Info |
 |
C79(0.00); C96(0.00) | LDD0219 | [10] | |
|
Methacrolein Probe Info |
 |
C96(0.00); C79(0.00) | LDD0218 | [10] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [11] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [12] | |
|
NAIA_5 Probe Info |
 |
C79(0.00); C96(0.00); C92(0.00) | LDD2223 | [13] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C220 Probe Info |
 |
43.41 | LDD1894 | [14] | |
|
C246 Probe Info |
 |
14.32 | LDD1919 | [14] | |
|
C307 Probe Info |
 |
15.24 | LDD1975 | [14] | |
|
C310 Probe Info |
 |
12.73 | LDD1977 | [14] | |
|
C423 Probe Info |
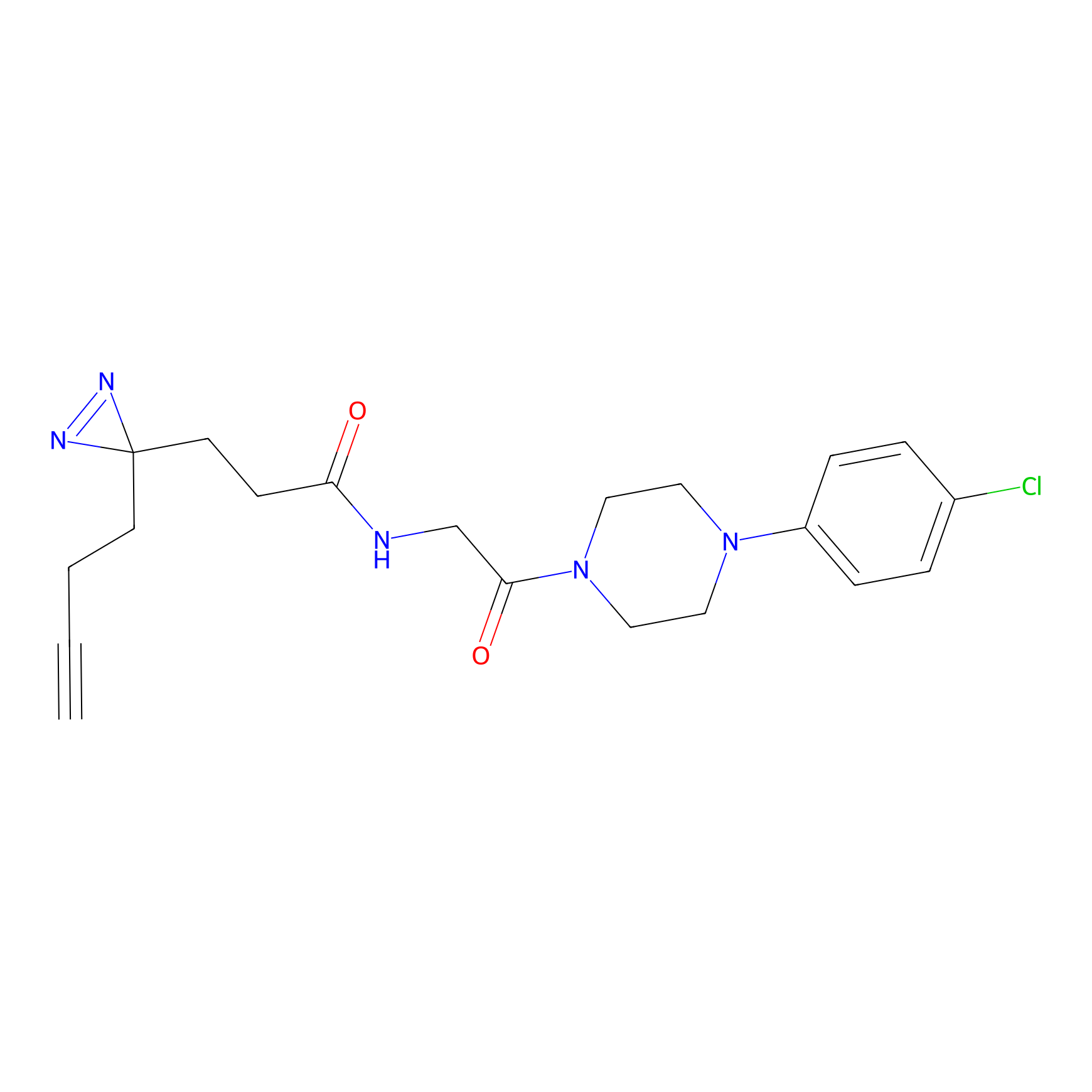 |
8.06 | LDD2078 | [14] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [15] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [15] | |
|
FFF probe14 Probe Info |
 |
20.00 | LDD0477 | [15] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0464 | [15] | |
|
JN0003 Probe Info |
 |
20.00 | LDD0484 | [15] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [16] | |
|
AEA-DA Probe Info |
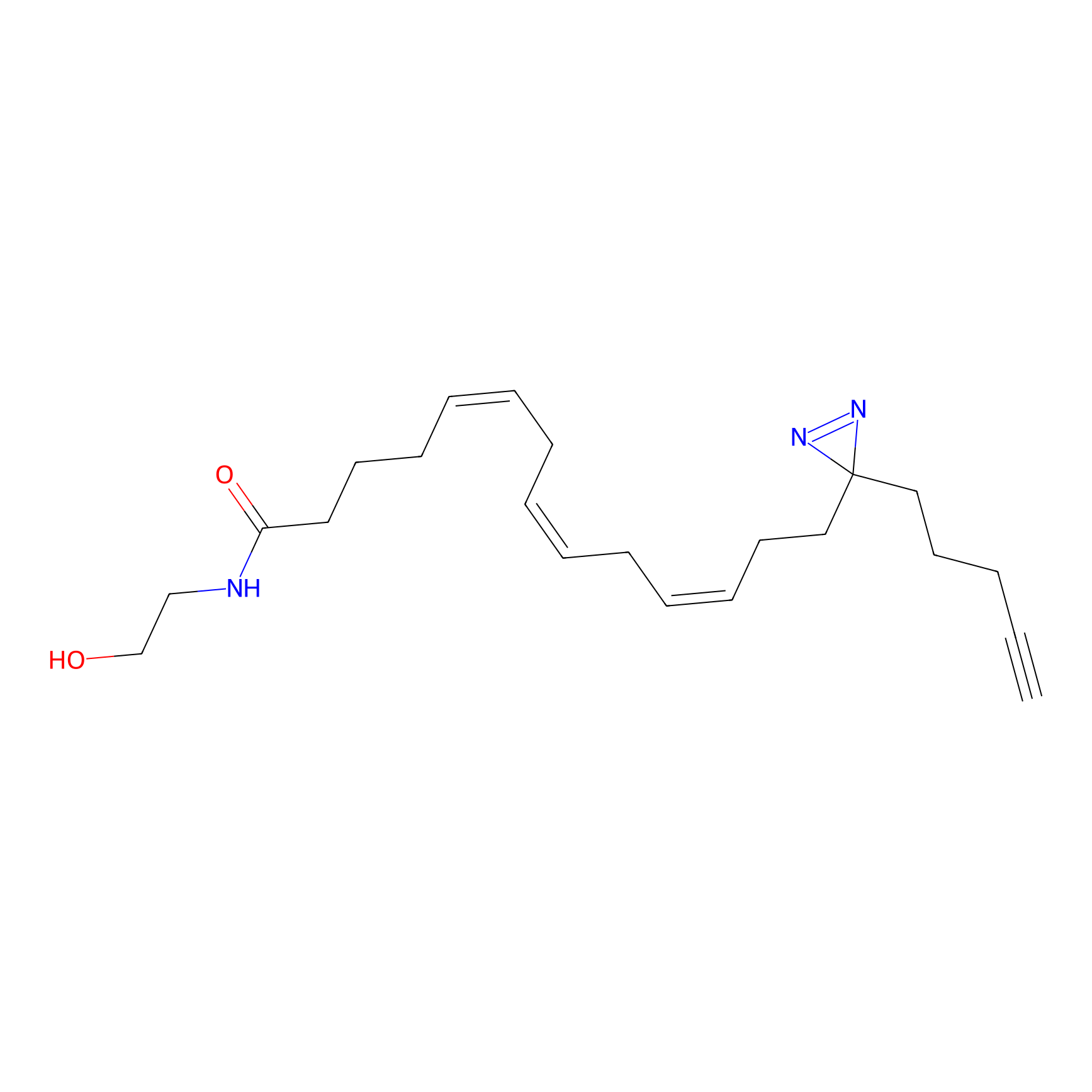 |
8.40 | LDD0333 | [17] | |
|
OEA-DA Probe Info |
 |
20.00 | LDD0046 | [17] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HCT 116 | C79(1.52) | LDD0531 | [5] |
| LDCM0215 | AC10 | HEK-293T | C79(1.08) | LDD0813 | [5] |
| LDCM0216 | AC100 | HCT 116 | C79(0.85) | LDD0533 | [5] |
| LDCM0217 | AC101 | HCT 116 | C79(0.75) | LDD0534 | [5] |
| LDCM0218 | AC102 | HCT 116 | C79(1.04) | LDD0535 | [5] |
| LDCM0219 | AC103 | HCT 116 | C79(0.85) | LDD0536 | [5] |
| LDCM0220 | AC104 | HCT 116 | C79(0.89) | LDD0537 | [5] |
| LDCM0221 | AC105 | HCT 116 | C79(0.81) | LDD0538 | [5] |
| LDCM0222 | AC106 | HCT 116 | C79(0.74) | LDD0539 | [5] |
| LDCM0223 | AC107 | HCT 116 | C79(0.88) | LDD0540 | [5] |
| LDCM0224 | AC108 | HCT 116 | C79(1.00) | LDD0541 | [5] |
| LDCM0225 | AC109 | HCT 116 | C79(0.74) | LDD0542 | [5] |
| LDCM0226 | AC11 | HEK-293T | C79(1.01) | LDD0824 | [5] |
| LDCM0227 | AC110 | HCT 116 | C79(0.72) | LDD0544 | [5] |
| LDCM0228 | AC111 | HCT 116 | C79(0.79) | LDD0545 | [5] |
| LDCM0229 | AC112 | HCT 116 | C79(0.85) | LDD0546 | [5] |
| LDCM0230 | AC113 | HEK-293T | C79(0.94) | LDD0828 | [5] |
| LDCM0231 | AC114 | HEK-293T | C79(1.06) | LDD0829 | [5] |
| LDCM0232 | AC115 | HEK-293T | C79(0.97) | LDD0830 | [5] |
| LDCM0233 | AC116 | HEK-293T | C79(0.88) | LDD0831 | [5] |
| LDCM0234 | AC117 | HEK-293T | C79(1.13) | LDD0832 | [5] |
| LDCM0235 | AC118 | HEK-293T | C79(1.01) | LDD0833 | [5] |
| LDCM0236 | AC119 | HEK-293T | C79(0.98) | LDD0834 | [5] |
| LDCM0237 | AC12 | HEK-293T | C79(1.12) | LDD0835 | [5] |
| LDCM0238 | AC120 | HEK-293T | C79(0.97) | LDD0836 | [5] |
| LDCM0239 | AC121 | HEK-293T | C79(1.06) | LDD0837 | [5] |
| LDCM0240 | AC122 | HEK-293T | C79(1.07) | LDD0838 | [5] |
| LDCM0241 | AC123 | HEK-293T | C79(0.88) | LDD0839 | [5] |
| LDCM0242 | AC124 | HEK-293T | C79(1.11) | LDD0840 | [5] |
| LDCM0243 | AC125 | HEK-293T | C79(1.30) | LDD0841 | [5] |
| LDCM0244 | AC126 | HEK-293T | C79(1.06) | LDD0842 | [5] |
| LDCM0245 | AC127 | HEK-293T | C79(0.96) | LDD0843 | [5] |
| LDCM0259 | AC14 | HEK-293T | C79(1.17) | LDD0857 | [5] |
| LDCM0270 | AC15 | HEK-293T | C79(1.53) | LDD0868 | [5] |
| LDCM0276 | AC17 | HEK-293T | C79(2.03) | LDD0874 | [5] |
| LDCM0277 | AC18 | HEK-293T | C79(1.63) | LDD0875 | [5] |
| LDCM0278 | AC19 | HEK-293T | C79(1.25) | LDD0876 | [5] |
| LDCM0279 | AC2 | HCT 116 | C79(1.61) | LDD0596 | [5] |
| LDCM0280 | AC20 | HEK-293T | C79(1.53) | LDD0878 | [5] |
| LDCM0281 | AC21 | HEK-293T | C79(1.64) | LDD0879 | [5] |
| LDCM0282 | AC22 | HEK-293T | C79(1.37) | LDD0880 | [5] |
| LDCM0283 | AC23 | HEK-293T | C79(1.54) | LDD0881 | [5] |
| LDCM0284 | AC24 | HEK-293T | C79(1.08) | LDD0882 | [5] |
| LDCM0285 | AC25 | HEK-293T | C79(1.00) | LDD0883 | [5] |
| LDCM0286 | AC26 | HEK-293T | C79(0.93) | LDD0884 | [5] |
| LDCM0287 | AC27 | HEK-293T | C79(1.07) | LDD0885 | [5] |
| LDCM0288 | AC28 | HEK-293T | C79(1.98) | LDD0886 | [5] |
| LDCM0289 | AC29 | HEK-293T | C79(1.25) | LDD0887 | [5] |
| LDCM0290 | AC3 | HCT 116 | C79(1.53) | LDD0607 | [5] |
| LDCM0291 | AC30 | HEK-293T | C79(1.06) | LDD0889 | [5] |
| LDCM0292 | AC31 | HEK-293T | C79(0.90) | LDD0890 | [5] |
| LDCM0293 | AC32 | HEK-293T | C79(0.94) | LDD0891 | [5] |
| LDCM0294 | AC33 | HEK-293T | C79(1.19) | LDD0892 | [5] |
| LDCM0295 | AC34 | HEK-293T | C79(0.89) | LDD0893 | [5] |
| LDCM0296 | AC35 | HEK-293T | C92(1.02); C96(1.04); C79(0.90); C148(1.04) | LDD1535 | [18] |
| LDCM0297 | AC36 | HEK-293T | C92(1.02); C96(1.13); C79(0.94); C148(0.93) | LDD1536 | [18] |
| LDCM0298 | AC37 | HEK-293T | C92(1.06); C96(1.10); C79(1.01); C148(1.09) | LDD1537 | [18] |
| LDCM0299 | AC38 | HEK-293T | C92(1.12); C96(1.01); C79(0.92); C148(0.92) | LDD1538 | [18] |
| LDCM0300 | AC39 | HEK-293T | C92(1.08); C96(1.03); C79(0.88); C148(0.97) | LDD1539 | [18] |
| LDCM0301 | AC4 | HCT 116 | C79(1.73) | LDD0618 | [5] |
| LDCM0302 | AC40 | HEK-293T | C92(1.07); C96(0.99); C79(0.92); C148(1.08) | LDD1541 | [18] |
| LDCM0303 | AC41 | HEK-293T | C92(1.05); C96(1.08); C79(1.06); C148(1.08) | LDD1542 | [18] |
| LDCM0304 | AC42 | HEK-293T | C92(1.02); C96(0.84); C79(0.93); C148(1.00) | LDD1543 | [18] |
| LDCM0305 | AC43 | HEK-293T | C92(1.02); C96(0.96); C79(0.92); C148(0.83) | LDD1544 | [18] |
| LDCM0306 | AC44 | HEK-293T | C92(0.99); C96(1.13); C79(0.97); C148(1.04) | LDD1545 | [18] |
| LDCM0307 | AC45 | HEK-293T | C92(1.03); C96(1.04); C79(0.97); C148(1.00) | LDD1546 | [18] |
| LDCM0308 | AC46 | HEK-293T | C79(0.98) | LDD0906 | [5] |
| LDCM0309 | AC47 | HEK-293T | C79(1.26) | LDD0907 | [5] |
| LDCM0310 | AC48 | HEK-293T | C79(1.49) | LDD0908 | [5] |
| LDCM0311 | AC49 | HEK-293T | C79(1.21) | LDD0909 | [5] |
| LDCM0312 | AC5 | HCT 116 | C79(1.82) | LDD0629 | [5] |
| LDCM0313 | AC50 | HEK-293T | C79(1.35) | LDD0911 | [5] |
| LDCM0314 | AC51 | HEK-293T | C79(1.56) | LDD0912 | [5] |
| LDCM0315 | AC52 | HEK-293T | C79(1.31) | LDD0913 | [5] |
| LDCM0316 | AC53 | HEK-293T | C79(1.22) | LDD0914 | [5] |
| LDCM0317 | AC54 | HEK-293T | C79(1.65) | LDD0915 | [5] |
| LDCM0318 | AC55 | HEK-293T | C79(1.48) | LDD0916 | [5] |
| LDCM0319 | AC56 | HEK-293T | C79(0.94) | LDD0917 | [5] |
| LDCM0320 | AC57 | HEK-293T | C79(0.97) | LDD0918 | [5] |
| LDCM0321 | AC58 | HEK-293T | C79(0.75) | LDD0919 | [5] |
| LDCM0322 | AC59 | HEK-293T | C79(1.05) | LDD0920 | [5] |
| LDCM0323 | AC6 | HEK-293T | C79(0.77) | LDD0921 | [5] |
| LDCM0324 | AC60 | HEK-293T | C79(1.75) | LDD0922 | [5] |
| LDCM0325 | AC61 | HEK-293T | C79(1.50) | LDD0923 | [5] |
| LDCM0326 | AC62 | HEK-293T | C79(0.67) | LDD0924 | [5] |
| LDCM0327 | AC63 | HEK-293T | C79(1.17) | LDD0925 | [5] |
| LDCM0328 | AC64 | HEK-293T | C79(0.94) | LDD0926 | [5] |
| LDCM0329 | AC65 | HEK-293T | C79(1.21) | LDD0927 | [5] |
| LDCM0330 | AC66 | HEK-293T | C79(0.75) | LDD0928 | [5] |
| LDCM0331 | AC67 | HEK-293T | C79(1.05) | LDD0929 | [5] |
| LDCM0332 | AC68 | HEK-293T | C79(1.09) | LDD0930 | [5] |
| LDCM0333 | AC69 | HEK-293T | C79(1.16) | LDD0931 | [5] |
| LDCM0334 | AC7 | HEK-293T | C79(1.16) | LDD0932 | [5] |
| LDCM0335 | AC70 | HEK-293T | C79(1.24) | LDD0933 | [5] |
| LDCM0336 | AC71 | HEK-293T | C79(1.16) | LDD0934 | [5] |
| LDCM0337 | AC72 | HEK-293T | C79(1.62) | LDD0935 | [5] |
| LDCM0338 | AC73 | HEK-293T | C79(1.51) | LDD0936 | [5] |
| LDCM0339 | AC74 | HEK-293T | C79(1.17) | LDD0937 | [5] |
| LDCM0340 | AC75 | HEK-293T | C79(1.18) | LDD0938 | [5] |
| LDCM0341 | AC76 | HEK-293T | C79(0.99) | LDD0939 | [5] |
| LDCM0342 | AC77 | HEK-293T | C79(1.07) | LDD0940 | [5] |
| LDCM0343 | AC78 | HEK-293T | C79(1.19) | LDD0941 | [5] |
| LDCM0344 | AC79 | HEK-293T | C79(1.15) | LDD0942 | [5] |
| LDCM0345 | AC8 | HEK-293T | C79(1.10) | LDD0943 | [5] |
| LDCM0346 | AC80 | HEK-293T | C79(1.14) | LDD0944 | [5] |
| LDCM0347 | AC81 | HEK-293T | C79(1.16) | LDD0945 | [5] |
| LDCM0348 | AC82 | HEK-293T | C79(1.25) | LDD0946 | [5] |
| LDCM0349 | AC83 | HEK-293T | C79(1.06) | LDD0947 | [5] |
| LDCM0350 | AC84 | HEK-293T | C79(1.13) | LDD0948 | [5] |
| LDCM0351 | AC85 | HEK-293T | C79(1.16) | LDD0949 | [5] |
| LDCM0352 | AC86 | HEK-293T | C79(1.25) | LDD0950 | [5] |
| LDCM0353 | AC87 | HEK-293T | C79(1.04) | LDD0951 | [5] |
| LDCM0354 | AC88 | HEK-293T | C79(1.19) | LDD0952 | [5] |
| LDCM0355 | AC89 | HEK-293T | C79(1.40) | LDD0953 | [5] |
| LDCM0357 | AC90 | HEK-293T | C79(1.39) | LDD0955 | [5] |
| LDCM0358 | AC91 | HEK-293T | C79(1.14) | LDD0956 | [5] |
| LDCM0359 | AC92 | HEK-293T | C79(1.12) | LDD0957 | [5] |
| LDCM0360 | AC93 | HEK-293T | C79(1.25) | LDD0958 | [5] |
| LDCM0361 | AC94 | HEK-293T | C79(1.21) | LDD0959 | [5] |
| LDCM0362 | AC95 | HEK-293T | C79(1.04) | LDD0960 | [5] |
| LDCM0363 | AC96 | HEK-293T | C79(1.14) | LDD0961 | [5] |
| LDCM0364 | AC97 | HEK-293T | C79(1.78) | LDD0962 | [5] |
| LDCM0365 | AC98 | HCT 116 | C79(0.89) | LDD0682 | [5] |
| LDCM0366 | AC99 | HCT 116 | C79(0.73) | LDD0683 | [5] |
| LDCM0248 | AKOS034007472 | HEK-293T | C79(1.08) | LDD0846 | [5] |
| LDCM0356 | AKOS034007680 | HEK-293T | C79(1.90) | LDD0954 | [5] |
| LDCM0275 | AKOS034007705 | HEK-293T | C79(1.42) | LDD0873 | [5] |
| LDCM0088 | C45 | HEK-293T | 20.00 | LDD0201 | [4] |
| LDCM0108 | Chloroacetamide | HeLa | C79(0.00); C96(0.00) | LDD0222 | [10] |
| LDCM0632 | CL-Sc | Hep-G2 | C79(1.87) | LDD2227 | [13] |
| LDCM0367 | CL1 | HEK-293T | C79(0.95) | LDD0965 | [5] |
| LDCM0368 | CL10 | HEK-293T | C79(2.51) | LDD0966 | [5] |
| LDCM0369 | CL100 | HCT 116 | C79(1.65) | LDD0686 | [5] |
| LDCM0370 | CL101 | HEK-293T | C79(1.67) | LDD0968 | [5] |
| LDCM0371 | CL102 | HEK-293T | C79(1.66) | LDD0969 | [5] |
| LDCM0372 | CL103 | HEK-293T | C79(1.36) | LDD0970 | [5] |
| LDCM0373 | CL104 | HEK-293T | C79(1.38) | LDD0971 | [5] |
| LDCM0374 | CL105 | HEK-293T | C79(1.28) | LDD0972 | [5] |
| LDCM0375 | CL106 | HEK-293T | C79(1.44) | LDD0973 | [5] |
| LDCM0376 | CL107 | HEK-293T | C79(1.35) | LDD0974 | [5] |
| LDCM0377 | CL108 | HEK-293T | C79(1.63) | LDD0975 | [5] |
| LDCM0378 | CL109 | HEK-293T | C79(1.20) | LDD0976 | [5] |
| LDCM0379 | CL11 | HEK-293T | C79(2.14) | LDD0977 | [5] |
| LDCM0380 | CL110 | HEK-293T | C79(1.37) | LDD0978 | [5] |
| LDCM0381 | CL111 | HEK-293T | C79(1.13) | LDD0979 | [5] |
| LDCM0382 | CL112 | HEK-293T | C79(1.25) | LDD0980 | [5] |
| LDCM0383 | CL113 | HEK-293T | C79(0.97) | LDD0981 | [5] |
| LDCM0384 | CL114 | HEK-293T | C79(1.51) | LDD0982 | [5] |
| LDCM0385 | CL115 | HEK-293T | C79(1.19) | LDD0983 | [5] |
| LDCM0386 | CL116 | HEK-293T | C79(1.10) | LDD0984 | [5] |
| LDCM0387 | CL117 | HEK-293T | C92(1.02); C96(1.04); C79(0.88); C148(0.99) | LDD1591 | [18] |
| LDCM0388 | CL118 | HEK-293T | C92(1.06); C96(1.03); C79(0.87); C148(0.97) | LDD1592 | [18] |
| LDCM0389 | CL119 | HEK-293T | C92(1.06); C96(1.02); C79(1.03); C148(0.82) | LDD1593 | [18] |
| LDCM0390 | CL12 | HEK-293T | C79(0.94) | LDD0988 | [5] |
| LDCM0391 | CL120 | HEK-293T | C92(1.03); C96(0.98); C79(0.97); C148(0.88) | LDD1595 | [18] |
| LDCM0392 | CL121 | HEK-293T | C79(1.70) | LDD0990 | [5] |
| LDCM0393 | CL122 | HEK-293T | C79(1.10) | LDD0991 | [5] |
| LDCM0394 | CL123 | HEK-293T | C79(2.15) | LDD0992 | [5] |
| LDCM0395 | CL124 | HEK-293T | C79(1.29) | LDD0993 | [5] |
| LDCM0396 | CL125 | HEK-293T | C79(1.10) | LDD0994 | [5] |
| LDCM0397 | CL126 | HEK-293T | C79(1.47) | LDD0995 | [5] |
| LDCM0398 | CL127 | HEK-293T | C79(1.05) | LDD0996 | [5] |
| LDCM0399 | CL128 | HEK-293T | C79(0.52) | LDD0997 | [5] |
| LDCM0400 | CL13 | HEK-293T | C79(0.99) | LDD0998 | [5] |
| LDCM0401 | CL14 | HEK-293T | C79(1.09) | LDD0999 | [5] |
| LDCM0402 | CL15 | HEK-293T | C79(1.94) | LDD1000 | [5] |
| LDCM0403 | CL16 | HCT 116 | C79(1.19) | LDD0720 | [5] |
| LDCM0404 | CL17 | HCT 116 | C79(1.07) | LDD0721 | [5] |
| LDCM0405 | CL18 | HCT 116 | C79(1.12) | LDD0722 | [5] |
| LDCM0406 | CL19 | HCT 116 | C79(1.33) | LDD0723 | [5] |
| LDCM0407 | CL2 | HEK-293T | C79(0.89) | LDD1005 | [5] |
| LDCM0408 | CL20 | HCT 116 | C79(1.17) | LDD0725 | [5] |
| LDCM0409 | CL21 | HCT 116 | C79(1.29) | LDD0726 | [5] |
| LDCM0410 | CL22 | HCT 116 | C79(1.83) | LDD0727 | [5] |
| LDCM0411 | CL23 | HCT 116 | C79(1.16) | LDD0728 | [5] |
| LDCM0412 | CL24 | HCT 116 | C79(1.30) | LDD0729 | [5] |
| LDCM0413 | CL25 | HCT 116 | C79(1.31) | LDD0730 | [5] |
| LDCM0414 | CL26 | HCT 116 | C79(1.32) | LDD0731 | [5] |
| LDCM0415 | CL27 | HCT 116 | C79(1.06) | LDD0732 | [5] |
| LDCM0416 | CL28 | HCT 116 | C79(1.24) | LDD0733 | [5] |
| LDCM0417 | CL29 | HCT 116 | C79(1.35) | LDD0734 | [5] |
| LDCM0418 | CL3 | HEK-293T | C79(1.07) | LDD1016 | [5] |
| LDCM0419 | CL30 | HCT 116 | C79(1.08) | LDD0736 | [5] |
| LDCM0420 | CL31 | HEK-293T | C79(1.21) | LDD1018 | [5] |
| LDCM0421 | CL32 | HEK-293T | C79(1.21) | LDD1019 | [5] |
| LDCM0422 | CL33 | HEK-293T | C79(2.66) | LDD1020 | [5] |
| LDCM0423 | CL34 | HEK-293T | C79(1.50) | LDD1021 | [5] |
| LDCM0424 | CL35 | HEK-293T | C79(1.30) | LDD1022 | [5] |
| LDCM0425 | CL36 | HEK-293T | C79(1.28) | LDD1023 | [5] |
| LDCM0426 | CL37 | HEK-293T | C79(1.43) | LDD1024 | [5] |
| LDCM0428 | CL39 | HEK-293T | C79(2.76) | LDD1026 | [5] |
| LDCM0429 | CL4 | HEK-293T | C79(0.95) | LDD1027 | [5] |
| LDCM0430 | CL40 | HEK-293T | C79(1.65) | LDD1028 | [5] |
| LDCM0431 | CL41 | HEK-293T | C79(2.15) | LDD1029 | [5] |
| LDCM0432 | CL42 | HEK-293T | C79(2.16) | LDD1030 | [5] |
| LDCM0433 | CL43 | HEK-293T | C79(2.97) | LDD1031 | [5] |
| LDCM0434 | CL44 | HEK-293T | C79(1.31) | LDD1032 | [5] |
| LDCM0435 | CL45 | HEK-293T | C79(2.88) | LDD1033 | [5] |
| LDCM0436 | CL46 | HEK-293T | C79(1.03) | LDD1034 | [5] |
| LDCM0437 | CL47 | HEK-293T | C79(0.97) | LDD1035 | [5] |
| LDCM0438 | CL48 | HEK-293T | C79(0.78) | LDD1036 | [5] |
| LDCM0439 | CL49 | HEK-293T | C79(1.17) | LDD1037 | [5] |
| LDCM0440 | CL5 | HEK-293T | C79(1.05) | LDD1038 | [5] |
| LDCM0441 | CL50 | HEK-293T | C79(1.60) | LDD1039 | [5] |
| LDCM0442 | CL51 | HEK-293T | C79(1.34) | LDD1040 | [5] |
| LDCM0443 | CL52 | HEK-293T | C79(0.68) | LDD1041 | [5] |
| LDCM0444 | CL53 | HEK-293T | C79(1.67) | LDD1042 | [5] |
| LDCM0445 | CL54 | HEK-293T | C79(1.29) | LDD1043 | [5] |
| LDCM0446 | CL55 | HEK-293T | C79(1.13) | LDD1044 | [5] |
| LDCM0447 | CL56 | HEK-293T | C79(0.88) | LDD1045 | [5] |
| LDCM0448 | CL57 | HEK-293T | C79(1.46) | LDD1046 | [5] |
| LDCM0449 | CL58 | HEK-293T | C79(1.33) | LDD1047 | [5] |
| LDCM0450 | CL59 | HEK-293T | C79(1.35) | LDD1048 | [5] |
| LDCM0451 | CL6 | HEK-293T | C79(1.28) | LDD1049 | [5] |
| LDCM0452 | CL60 | HEK-293T | C79(1.79) | LDD1050 | [5] |
| LDCM0453 | CL61 | HCT 116 | C79(1.06) | LDD0770 | [5] |
| LDCM0454 | CL62 | HCT 116 | C79(1.24) | LDD0771 | [5] |
| LDCM0455 | CL63 | HCT 116 | C79(1.18) | LDD0772 | [5] |
| LDCM0456 | CL64 | HCT 116 | C79(1.44) | LDD0773 | [5] |
| LDCM0457 | CL65 | HCT 116 | C79(1.23) | LDD0774 | [5] |
| LDCM0458 | CL66 | HCT 116 | C79(1.56) | LDD0775 | [5] |
| LDCM0459 | CL67 | HCT 116 | C79(1.42) | LDD0776 | [5] |
| LDCM0460 | CL68 | HCT 116 | C79(1.33) | LDD0777 | [5] |
| LDCM0461 | CL69 | HCT 116 | C79(1.14) | LDD0778 | [5] |
| LDCM0462 | CL7 | HEK-293T | C79(1.20) | LDD1060 | [5] |
| LDCM0463 | CL70 | HCT 116 | C79(1.30) | LDD0780 | [5] |
| LDCM0464 | CL71 | HCT 116 | C79(1.28) | LDD0781 | [5] |
| LDCM0465 | CL72 | HCT 116 | C79(1.28) | LDD0782 | [5] |
| LDCM0466 | CL73 | HCT 116 | C79(1.37) | LDD0783 | [5] |
| LDCM0467 | CL74 | HCT 116 | C79(1.28) | LDD0784 | [5] |
| LDCM0469 | CL76 | HEK-293T | C79(1.36) | LDD1067 | [5] |
| LDCM0470 | CL77 | HEK-293T | C79(1.41) | LDD1068 | [5] |
| LDCM0471 | CL78 | HEK-293T | C79(1.39) | LDD1069 | [5] |
| LDCM0472 | CL79 | HEK-293T | C79(1.34) | LDD1070 | [5] |
| LDCM0473 | CL8 | HEK-293T | C79(1.15) | LDD1071 | [5] |
| LDCM0474 | CL80 | HEK-293T | C79(1.31) | LDD1072 | [5] |
| LDCM0475 | CL81 | HEK-293T | C79(1.27) | LDD1073 | [5] |
| LDCM0476 | CL82 | HEK-293T | C79(1.08) | LDD1074 | [5] |
| LDCM0477 | CL83 | HEK-293T | C79(1.28) | LDD1075 | [5] |
| LDCM0478 | CL84 | HEK-293T | C79(1.40) | LDD1076 | [5] |
| LDCM0479 | CL85 | HEK-293T | C79(1.49) | LDD1077 | [5] |
| LDCM0480 | CL86 | HEK-293T | C79(1.62) | LDD1078 | [5] |
| LDCM0481 | CL87 | HEK-293T | C79(1.30) | LDD1079 | [5] |
| LDCM0482 | CL88 | HEK-293T | C79(1.49) | LDD1080 | [5] |
| LDCM0483 | CL89 | HEK-293T | C79(1.28) | LDD1081 | [5] |
| LDCM0484 | CL9 | HEK-293T | C79(1.11) | LDD1082 | [5] |
| LDCM0485 | CL90 | HEK-293T | C79(1.77) | LDD1083 | [5] |
| LDCM0486 | CL91 | HCT 116 | C79(1.06) | LDD0803 | [5] |
| LDCM0487 | CL92 | HCT 116 | C79(1.02) | LDD0804 | [5] |
| LDCM0488 | CL93 | HCT 116 | C79(1.03) | LDD0805 | [5] |
| LDCM0489 | CL94 | HCT 116 | C79(1.13) | LDD0806 | [5] |
| LDCM0490 | CL95 | HCT 116 | C79(1.15) | LDD0807 | [5] |
| LDCM0491 | CL96 | HCT 116 | C79(1.22) | LDD0808 | [5] |
| LDCM0492 | CL97 | HCT 116 | C79(1.93) | LDD0809 | [5] |
| LDCM0493 | CL98 | HCT 116 | C79(1.66) | LDD0810 | [5] |
| LDCM0494 | CL99 | HCT 116 | C79(1.65) | LDD0811 | [5] |
| LDCM0495 | E2913 | HEK-293T | C92(1.01); C96(1.08); C79(0.92); C148(0.85) | LDD1698 | [18] |
| LDCM0625 | F8 | Ramos | C96(2.00); C79(1.04); C92(1.33) | LDD2187 | [19] |
| LDCM0572 | Fragment10 | Ramos | C96(1.01); C79(2.39); C92(1.28) | LDD2189 | [19] |
| LDCM0573 | Fragment11 | Ramos | C96(0.01); C79(0.04); C92(0.00) | LDD2190 | [19] |
| LDCM0574 | Fragment12 | Ramos | C96(1.42); C79(2.04); C92(1.51) | LDD2191 | [19] |
| LDCM0575 | Fragment13 | Ramos | C96(0.91); C79(1.27); C92(1.01) | LDD2192 | [19] |
| LDCM0576 | Fragment14 | Ramos | C96(1.62); C79(1.65); C92(1.76) | LDD2193 | [19] |
| LDCM0579 | Fragment20 | Ramos | C96(1.75); C79(2.68); C92(2.60) | LDD2194 | [19] |
| LDCM0580 | Fragment21 | Ramos | C96(0.94); C79(1.17); C92(1.08) | LDD2195 | [19] |
| LDCM0582 | Fragment23 | Ramos | C96(0.68); C79(1.03); C92(1.03) | LDD2196 | [19] |
| LDCM0578 | Fragment27 | Ramos | C96(0.91); C79(0.99); C92(0.96) | LDD2197 | [19] |
| LDCM0586 | Fragment28 | Ramos | C96(0.90); C79(1.44); C92(0.73) | LDD2198 | [19] |
| LDCM0588 | Fragment30 | Ramos | C96(1.06); C79(0.81); C92(0.96) | LDD2199 | [19] |
| LDCM0589 | Fragment31 | Ramos | C96(0.79); C79(0.70); C92(0.91) | LDD2200 | [19] |
| LDCM0590 | Fragment32 | Ramos | C96(0.97); C79(1.10); C92(0.83) | LDD2201 | [19] |
| LDCM0468 | Fragment33 | HCT 116 | C79(1.24) | LDD0785 | [5] |
| LDCM0596 | Fragment38 | Ramos | C96(0.98); C79(0.91); C92(0.95) | LDD2203 | [19] |
| LDCM0566 | Fragment4 | Ramos | C96(1.31); C79(1.67); C92(1.30) | LDD2184 | [19] |
| LDCM0427 | Fragment51 | HEK-293T | C79(1.29) | LDD1025 | [5] |
| LDCM0610 | Fragment52 | Ramos | C96(0.97); C79(0.79); C92(1.15) | LDD2204 | [19] |
| LDCM0614 | Fragment56 | Ramos | C96(1.23); C79(0.66); C92(1.07) | LDD2205 | [19] |
| LDCM0569 | Fragment7 | Ramos | C96(1.40); C79(1.38); C92(1.43) | LDD2186 | [19] |
| LDCM0571 | Fragment9 | Ramos | C96(3.36); C79(5.63); C92(2.93) | LDD2188 | [19] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [10] |
| LDCM0022 | KB02 | HCT 116 | C79(1.48) | LDD0080 | [5] |
| LDCM0023 | KB03 | HCT 116 | C79(2.40) | LDD0081 | [5] |
| LDCM0024 | KB05 | HCT 116 | C79(1.47) | LDD0082 | [5] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0225 | [10] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C79(0.61) | LDD2206 | [20] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C96(0.76); C79(0.50) | LDD2207 | [20] |
| LDCM0131 | RA190 | MM1.R | C79(1.43); C96(1.41) | LDD0304 | [21] |
| LDCM0136 | SR-4559 | HEK-293T | 8.40 | LDD0333 | [17] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| 14-3-3 protein gamma (YWHAG) | 14-3-3 family | P61981 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Gelsolin (GSN) | Villin/gelsolin family | P06396 | |||
References
