Details of the Target
General Information of Target
| Target ID | LDTP11993 | |||||
|---|---|---|---|---|---|---|
| Target Name | Nucleolar protein 6 (NOL6) | |||||
| Gene Name | NOL6 | |||||
| Gene ID | 65083 | |||||
| Synonyms |
Nucleolar protein 6; Nucleolar RNA-associated protein; Nrap |
|||||
| 3D Structure | ||||||
| Sequence |
MKPSWLQCRKVTSAGGLGGPLPGSSPARGAGAALRALVVPGPRGGLGGRGCRALSSGSGS
EYKTHFAASVTDPERFWGKAAEQISWYKPWTKTLENKHSPSTRWFVEGMLNICYNAVDRH IENGKGDKIAIIYDSPVTNTKATFTYKEVLEQVSKLAGVLVKHGIKKGDTVVIYMPMIPQ AMYTMLACARIGAIHSLIFGGFASKELSSRIDHVKPKVVVTASFGIEPGRRVEYVPLVEE ALKIGQHKPDKILIYNRPNMEAVPLAPGRDLDWDEEMAKAQSHDCVPVLSEHPLYILYTS GTTGLPKGVIRPTGGYAVMLHWSMSSIYGLQPGEVWWAASDLGWVVGHSYICYGPLLHGN TTVLYEGKPVGTPDAGAYFRVLAEHGVAALFTAPTAIRAIRQQDPGAALGKQYSLTRFKT LFVAGERCDVETLEWSKNVFRVPVLDHWWQTETGSPITASCVGLGNSKTPPPGQAGKSVP GYNVMILDDNMQKLKARCLGNIVVKLPLPPGAFSGLWKNQEAFKHLYFEKFPGYYDTMDA GYMDEEGYLYVMSRVDDVINVAGHRISAGAIEESILSHGTVADCAVVGKEDPLKGHVPLA LCVLRKDINATEEQVLEEIVKHVRQNIGPVAAFRNAVFVKQLPKTRSGKIPRSALSAIVN GKPYKITSTIEDPSIFGHVEEMLKQA |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
NRAP family
|
|||||
| Subcellular location |
Nucleus, nucleolus
|
|||||
| Function |
Part of the small subunit (SSU) processome, first precursor of the small eukaryotic ribosomal subunit. During the assembly of the SSU processome in the nucleolus, many ribosome biogenesis factors, an RNA chaperone and ribosomal proteins associate with the nascent pre-rRNA and work in concert to generate RNA folding, modifications, rearrangements and cleavage as well as targeted degradation of pre-ribosomal RNA by the RNA exosome.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| 22RV1 | Deletion: p.Q953KfsTer32 | DBIA Probe Info | |||
| 786O | SNV: p.R521W | DBIA Probe Info | |||
| ECC2 | SNV: p.H527Y | DBIA Probe Info | |||
| HCT116 | Deletion: p.Q953KfsTer32 | DBIA Probe Info | |||
| HCT15 | SNV: p.R173Ter; p.R186M | DBIA Probe Info | |||
| IGROV1 | Deletion: p.G639AfsTer10 | DBIA Probe Info | |||
| JURKAT | SNV: p.Q410Ter | Compound 10 Probe Info | |||
| MCC13 | SNV: p.P255S | DBIA Probe Info | |||
| MDAMB468 | SNV: p.Q865P | DBIA Probe Info | |||
| MOLT4 | SNV: p.G48C; p.Q282H; p.R442L | IA-alkyne Probe Info | |||
| P31FUJ | SNV: p.R153P | DBIA Probe Info | |||
| PF382 | SNV: p.R856W | DBIA Probe Info | |||
| RL952 | Deletion: p.G639AfsTer10 | DBIA Probe Info | |||
| SKES1 | SNV: p.D789G | DBIA Probe Info | |||
| TOV21G | SNV: p.Q1092H | DBIA Probe Info | |||
| WM793 | SNV: p.P615S | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [2] | |
|
BTD Probe Info |
 |
C486(2.27) | LDD1700 | [3] | |
|
AHL-Pu-1 Probe Info |
 |
C265(2.76) | LDD0169 | [4] | |
|
DBIA Probe Info |
 |
C1034(0.91) | LDD0078 | [5] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [6] | |
|
N1 Probe Info |
 |
N.A. | LDD0245 | [7] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C777(0.00); C486(0.00); C665(0.00) | LDD0038 | [8] | |
|
IA-alkyne Probe Info |
 |
C665(0.00); C777(0.00) | LDD0036 | [8] | |
|
Lodoacetamide azide Probe Info |
 |
C777(0.00); C665(0.00); C486(0.00); C1034(0.00) | LDD0037 | [8] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [9] | |
|
TFBX Probe Info |
 |
C1034(0.00); C665(0.00) | LDD0027 | [9] | |
|
WYneN Probe Info |
 |
C265(0.00); C1034(0.00) | LDD0021 | [10] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [10] | |
|
aHNE Probe Info |
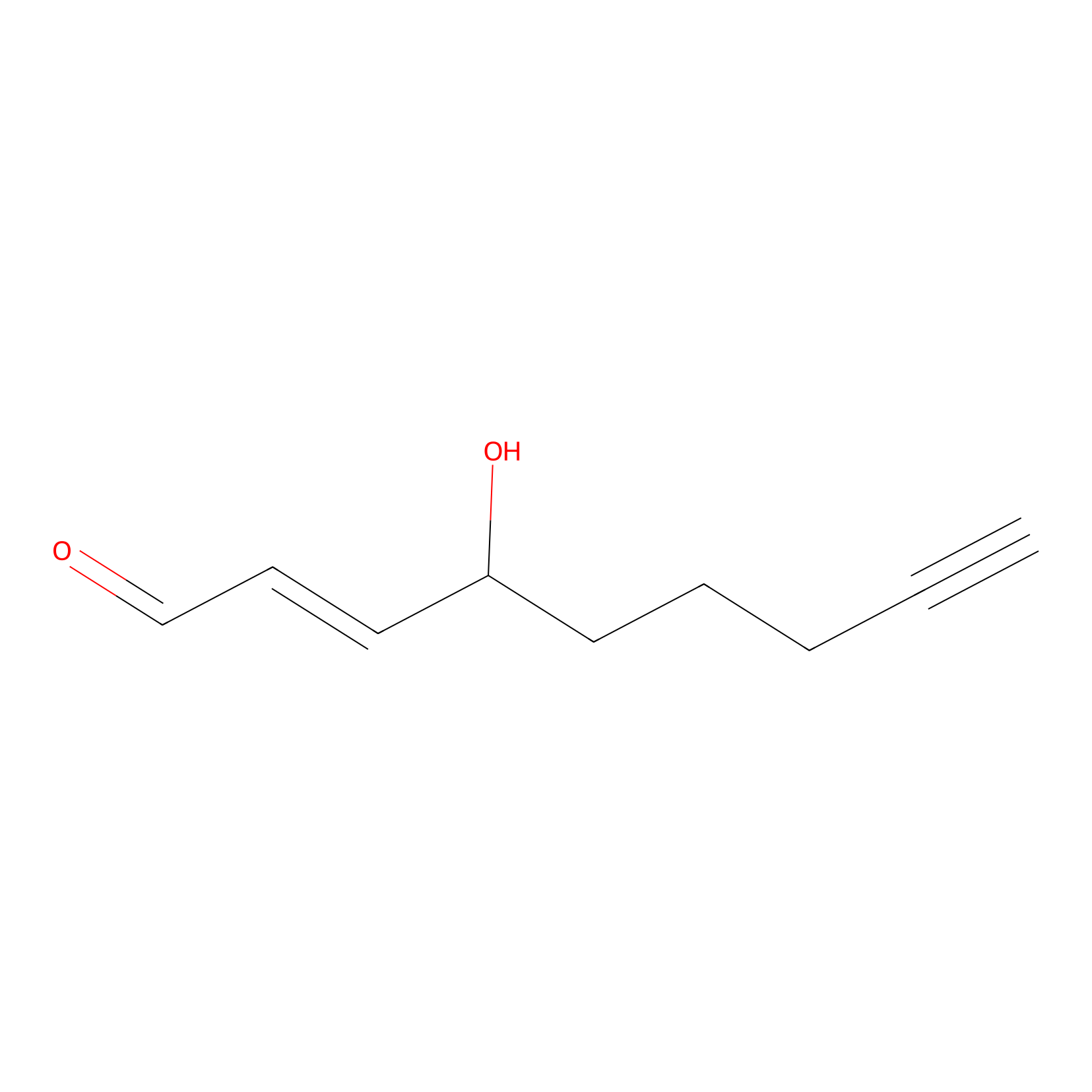 |
N.A. | LDD0001 | [10] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [11] | |
|
Compound 11 Probe Info |
 |
N.A. | LDD2213 | [11] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [10] | |
|
IPM Probe Info |
 |
C1034(0.00); C265(0.00) | LDD0005 | [10] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [12] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [13] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [14] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [15] | |
|
Crotonaldehyde Probe Info |
 |
C777(0.00); C486(0.00); C1034(0.00) | LDD0219 | [15] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [15] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [16] | |
|
NAIA_5 Probe Info |
 |
C777(0.00); C486(0.00) | LDD2223 | [17] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C486(0.48) | LDD2095 | [3] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C486(0.81) | LDD2130 | [3] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C486(0.75) | LDD2117 | [3] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C486(1.63) | LDD2152 | [3] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C486(0.77) | LDD2131 | [3] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C265(2.76) | LDD0169 | [4] |
| LDCM0214 | AC1 | HCT 116 | C1034(1.05); C256(1.68); C265(1.57); C665(1.55) | LDD0531 | [5] |
| LDCM0215 | AC10 | HCT 116 | C1034(1.19); C256(1.38); C265(1.38); C665(1.36) | LDD0532 | [5] |
| LDCM0216 | AC100 | HCT 116 | C728(0.93); C739(0.93); C1034(1.23); C256(0.71) | LDD0533 | [5] |
| LDCM0217 | AC101 | HCT 116 | C728(0.85); C739(0.85); C1034(1.24); C256(0.68) | LDD0534 | [5] |
| LDCM0218 | AC102 | HCT 116 | C728(0.75); C739(0.75); C1034(1.43); C256(0.80) | LDD0535 | [5] |
| LDCM0219 | AC103 | HCT 116 | C728(1.10); C739(1.10); C1034(1.41); C256(0.61) | LDD0536 | [5] |
| LDCM0220 | AC104 | HCT 116 | C728(0.86); C739(0.86); C1034(0.99); C256(0.90) | LDD0537 | [5] |
| LDCM0221 | AC105 | HCT 116 | C728(0.74); C739(0.74); C1034(1.47); C256(0.71) | LDD0538 | [5] |
| LDCM0222 | AC106 | HCT 116 | C728(1.21); C739(1.21); C1034(1.45); C256(0.61) | LDD0539 | [5] |
| LDCM0223 | AC107 | HCT 116 | C728(1.09); C739(1.09); C1034(1.34); C256(0.75) | LDD0540 | [5] |
| LDCM0224 | AC108 | HCT 116 | C728(0.79); C739(0.79); C1034(1.26); C256(0.65) | LDD0541 | [5] |
| LDCM0225 | AC109 | HCT 116 | C728(0.74); C739(0.74); C1034(1.18); C256(0.90) | LDD0542 | [5] |
| LDCM0226 | AC11 | HCT 116 | C1034(1.21); C256(1.63); C265(1.36); C665(1.66) | LDD0543 | [5] |
| LDCM0227 | AC110 | HCT 116 | C728(0.95); C739(0.95); C1034(1.33); C256(0.68) | LDD0544 | [5] |
| LDCM0228 | AC111 | HCT 116 | C728(0.91); C739(0.91); C1034(1.48); C256(0.65) | LDD0545 | [5] |
| LDCM0229 | AC112 | HCT 116 | C728(0.77); C739(0.77); C1034(1.34); C256(0.54) | LDD0546 | [5] |
| LDCM0230 | AC113 | HCT 116 | C728(1.09); C739(1.09); C1034(0.93); C256(1.21) | LDD0547 | [5] |
| LDCM0231 | AC114 | HCT 116 | C728(1.12); C739(0.90); C1034(0.98); C256(1.40) | LDD0548 | [5] |
| LDCM0232 | AC115 | HCT 116 | C728(1.14); C739(0.86); C1034(1.19); C256(1.20) | LDD0549 | [5] |
| LDCM0233 | AC116 | HCT 116 | C728(0.91); C739(1.01); C1034(1.11); C256(1.35) | LDD0550 | [5] |
| LDCM0234 | AC117 | HCT 116 | C728(1.04); C739(0.88); C1034(0.88); C256(1.21) | LDD0551 | [5] |
| LDCM0235 | AC118 | HCT 116 | C728(1.05); C739(1.00); C1034(0.83); C256(1.53) | LDD0552 | [5] |
| LDCM0236 | AC119 | HCT 116 | C728(1.12); C739(0.88); C1034(1.03); C256(1.31) | LDD0553 | [5] |
| LDCM0237 | AC12 | HCT 116 | C1034(1.09); C256(1.48); C265(1.20); C665(1.61) | LDD0554 | [5] |
| LDCM0238 | AC120 | HCT 116 | C728(1.05); C739(0.89); C1034(0.97); C256(1.35) | LDD0555 | [5] |
| LDCM0239 | AC121 | HCT 116 | C728(0.92); C739(0.96); C1034(0.85); C256(1.82) | LDD0556 | [5] |
| LDCM0240 | AC122 | HCT 116 | C728(1.05); C739(0.83); C1034(0.80); C256(1.05) | LDD0557 | [5] |
| LDCM0241 | AC123 | HCT 116 | C728(0.96); C739(0.98); C1034(0.81); C256(1.66) | LDD0558 | [5] |
| LDCM0242 | AC124 | HCT 116 | C728(1.12); C739(0.92); C1034(0.76); C256(1.69) | LDD0559 | [5] |
| LDCM0243 | AC125 | HCT 116 | C728(1.12); C739(0.90); C1034(0.93); C256(1.56) | LDD0560 | [5] |
| LDCM0244 | AC126 | HCT 116 | C728(1.08); C739(0.99); C1034(0.99); C256(1.60) | LDD0561 | [5] |
| LDCM0245 | AC127 | HCT 116 | C728(1.02); C739(0.92); C1034(1.32); C256(1.73) | LDD0562 | [5] |
| LDCM0246 | AC128 | HCT 116 | C728(1.59); C739(1.59); C1034(0.97); C265(1.28) | LDD0563 | [5] |
| LDCM0247 | AC129 | HCT 116 | C728(1.10); C739(1.10); C1034(1.05); C265(1.34) | LDD0564 | [5] |
| LDCM0249 | AC130 | HCT 116 | C728(0.91); C739(0.91); C1034(0.90); C265(1.37) | LDD0566 | [5] |
| LDCM0250 | AC131 | HCT 116 | C728(1.19); C739(1.19); C1034(0.91); C265(1.28) | LDD0567 | [5] |
| LDCM0251 | AC132 | HCT 116 | C728(0.56); C739(0.56); C1034(1.20); C265(1.24) | LDD0568 | [5] |
| LDCM0252 | AC133 | HCT 116 | C728(0.74); C739(0.74); C1034(1.13); C265(1.34) | LDD0569 | [5] |
| LDCM0253 | AC134 | HCT 116 | C728(0.72); C739(0.72); C1034(1.63); C265(1.50) | LDD0570 | [5] |
| LDCM0254 | AC135 | HCT 116 | C728(0.69); C739(0.69); C1034(1.03); C265(1.41) | LDD0571 | [5] |
| LDCM0255 | AC136 | HCT 116 | C728(0.70); C739(0.70); C1034(1.15); C265(1.31) | LDD0572 | [5] |
| LDCM0256 | AC137 | HCT 116 | C728(0.82); C739(0.82); C1034(1.32); C265(1.36) | LDD0573 | [5] |
| LDCM0257 | AC138 | HCT 116 | C728(0.63); C739(0.63); C1034(1.68); C265(1.72) | LDD0574 | [5] |
| LDCM0258 | AC139 | HCT 116 | C728(0.47); C739(0.47); C1034(1.26); C265(1.40) | LDD0575 | [5] |
| LDCM0259 | AC14 | HCT 116 | C1034(0.91); C256(1.24); C265(1.22); C665(1.62) | LDD0576 | [5] |
| LDCM0260 | AC140 | HCT 116 | C728(0.46); C739(0.46); C1034(1.65); C265(1.60) | LDD0577 | [5] |
| LDCM0261 | AC141 | HCT 116 | C728(0.48); C739(0.48); C1034(1.28); C265(1.63) | LDD0578 | [5] |
| LDCM0262 | AC142 | HCT 116 | C728(0.60); C739(0.60); C1034(1.48); C265(1.68) | LDD0579 | [5] |
| LDCM0263 | AC143 | HCT 116 | C256(0.73); C665(0.81); C265(0.91); C777(0.94) | LDD0580 | [5] |
| LDCM0264 | AC144 | HCT 116 | C777(0.52); C256(0.86); C665(0.86); C265(0.91) | LDD0581 | [5] |
| LDCM0265 | AC145 | HCT 116 | C777(0.64); C265(0.90); C256(0.93); C665(0.94) | LDD0582 | [5] |
| LDCM0266 | AC146 | HCT 116 | C777(0.61); C256(0.67); C665(0.78); C265(0.94) | LDD0583 | [5] |
| LDCM0267 | AC147 | HCT 116 | C777(0.48); C265(0.93); C665(0.94); C256(1.11) | LDD0584 | [5] |
| LDCM0268 | AC148 | HCT 116 | C777(0.34); C665(0.62); C256(0.75); C265(1.27) | LDD0585 | [5] |
| LDCM0269 | AC149 | HCT 116 | C777(0.48); C665(0.77); C256(0.80); C265(0.98) | LDD0586 | [5] |
| LDCM0270 | AC15 | HCT 116 | C777(0.77); C1034(1.07); C256(1.11); C665(1.34) | LDD0587 | [5] |
| LDCM0271 | AC150 | HCT 116 | C256(0.76); C777(0.77); C665(0.84); C265(1.06) | LDD0588 | [5] |
| LDCM0272 | AC151 | HCT 116 | C777(0.76); C665(0.79); C256(0.85); C265(0.91) | LDD0589 | [5] |
| LDCM0273 | AC152 | HCT 116 | C777(0.40); C665(0.75); C256(0.77); C265(0.97) | LDD0590 | [5] |
| LDCM0274 | AC153 | HCT 116 | C777(0.37); C256(0.40); C665(0.67); C265(1.30) | LDD0591 | [5] |
| LDCM0621 | AC154 | HCT 116 | C1034(1.33); C256(0.68); C265(0.98); C665(0.95) | LDD2158 | [5] |
| LDCM0622 | AC155 | HCT 116 | C1034(1.34); C256(0.62); C265(0.70); C665(0.66) | LDD2159 | [5] |
| LDCM0623 | AC156 | HCT 116 | C1034(1.03); C256(0.68); C265(0.81); C665(0.80) | LDD2160 | [5] |
| LDCM0624 | AC157 | HCT 116 | C1034(1.01); C256(0.68); C265(0.94); C665(0.74) | LDD2161 | [5] |
| LDCM0276 | AC17 | HCT 116 | C256(0.98); C265(1.01); C777(1.03); C665(1.03) | LDD0593 | [5] |
| LDCM0277 | AC18 | HCT 116 | C256(0.67); C265(0.75); C777(1.15); C1034(1.27) | LDD0594 | [5] |
| LDCM0278 | AC19 | HCT 116 | C256(0.90); C265(0.91); C777(0.95); C665(1.00) | LDD0595 | [5] |
| LDCM0279 | AC2 | HCT 116 | C1034(1.00); C665(1.25); C777(1.27); C256(1.31) | LDD0596 | [5] |
| LDCM0280 | AC20 | HCT 116 | C665(0.69); C256(0.87); C777(0.90); C265(0.95) | LDD0597 | [5] |
| LDCM0281 | AC21 | HCT 116 | C256(0.81); C265(0.88); C777(1.14); C1034(1.17) | LDD0598 | [5] |
| LDCM0282 | AC22 | HCT 116 | C665(0.85); C777(1.04); C256(1.05); C265(1.08) | LDD0599 | [5] |
| LDCM0283 | AC23 | HCT 116 | C777(0.89); C256(0.90); C665(0.97); C265(1.02) | LDD0600 | [5] |
| LDCM0284 | AC24 | HCT 116 | C665(0.92); C256(1.04); C777(1.07); C1034(1.11) | LDD0601 | [5] |
| LDCM0285 | AC25 | HCT 116 | C728(0.63); C739(0.63); C777(0.94); C1034(0.97) | LDD0602 | [5] |
| LDCM0286 | AC26 | HCT 116 | C728(0.40); C739(0.40); C777(0.68); C665(0.78) | LDD0603 | [5] |
| LDCM0287 | AC27 | HCT 116 | C777(0.67); C728(0.68); C739(0.68); C265(0.84) | LDD0604 | [5] |
| LDCM0288 | AC28 | HCT 116 | C728(0.36); C739(0.36); C777(0.68); C665(0.73) | LDD0605 | [5] |
| LDCM0289 | AC29 | HCT 116 | C728(0.33); C739(0.33); C777(0.62); C665(0.73) | LDD0606 | [5] |
| LDCM0290 | AC3 | HCT 116 | C1034(1.05); C265(1.44); C777(1.69); C665(1.84) | LDD0607 | [5] |
| LDCM0291 | AC30 | HCT 116 | C728(0.30); C739(0.30); C777(0.53); C665(0.64) | LDD0608 | [5] |
| LDCM0292 | AC31 | HCT 116 | C728(0.41); C739(0.41); C777(0.68); C665(0.89) | LDD0609 | [5] |
| LDCM0293 | AC32 | HCT 116 | C728(0.23); C739(0.23); C777(0.49); C665(0.65) | LDD0610 | [5] |
| LDCM0294 | AC33 | HCT 116 | C777(0.63); C665(0.93); C265(0.99); C728(1.01) | LDD0611 | [5] |
| LDCM0295 | AC34 | HCT 116 | C777(0.57); C265(0.96); C665(1.05); C1034(1.57) | LDD0612 | [5] |
| LDCM0296 | AC35 | HCT 116 | C1034(0.89); C265(1.15); C256(1.16); C728(1.20) | LDD0613 | [5] |
| LDCM0297 | AC36 | HCT 116 | C728(0.83); C739(0.83); C1034(0.87); C256(1.00) | LDD0614 | [5] |
| LDCM0298 | AC37 | HCT 116 | C1034(0.84); C256(0.88); C265(0.99); C777(1.05) | LDD0615 | [5] |
| LDCM0299 | AC38 | HCT 116 | C1034(0.90); C256(0.94); C265(1.05); C777(1.10) | LDD0616 | [5] |
| LDCM0300 | AC39 | HCT 116 | C728(0.58); C739(0.58); C256(0.82); C265(0.86) | LDD0617 | [5] |
| LDCM0301 | AC4 | HCT 116 | C1034(1.17); C265(1.41); C777(2.05); C256(2.08) | LDD0618 | [5] |
| LDCM0302 | AC40 | HCT 116 | C256(0.63); C777(0.72); C265(0.82); C728(0.93) | LDD0619 | [5] |
| LDCM0303 | AC41 | HCT 116 | C256(0.84); C728(0.89); C739(0.89); C777(0.95) | LDD0620 | [5] |
| LDCM0304 | AC42 | HCT 116 | C256(0.76); C665(0.90); C265(0.92); C777(0.95) | LDD0621 | [5] |
| LDCM0305 | AC43 | HCT 116 | C256(0.73); C665(0.85); C265(0.87); C777(0.99) | LDD0622 | [5] |
| LDCM0306 | AC44 | HCT 116 | C256(0.75); C265(0.85); C777(0.85); C1034(0.96) | LDD0623 | [5] |
| LDCM0307 | AC45 | HCT 116 | C256(0.59); C728(0.72); C739(0.72); C265(0.74) | LDD0624 | [5] |
| LDCM0308 | AC46 | HCT 116 | C665(0.69); C256(0.95); C777(1.05); C728(1.10) | LDD0625 | [5] |
| LDCM0309 | AC47 | HCT 116 | C256(0.85); C265(0.94); C728(0.94); C739(0.94) | LDD0626 | [5] |
| LDCM0310 | AC48 | HCT 116 | C728(0.48); C739(0.48); C665(0.71); C777(0.91) | LDD0627 | [5] |
| LDCM0311 | AC49 | HCT 116 | C777(0.52); C728(0.62); C739(0.62); C665(0.68) | LDD0628 | [5] |
| LDCM0312 | AC5 | HCT 116 | C1034(1.16); C265(1.79); C777(1.91); C256(2.08) | LDD0629 | [5] |
| LDCM0313 | AC50 | HCT 116 | C665(0.54); C777(0.62); C728(0.70); C739(0.70) | LDD0630 | [5] |
| LDCM0314 | AC51 | HCT 116 | C665(0.56); C256(0.94); C728(1.12); C739(1.12) | LDD0631 | [5] |
| LDCM0315 | AC52 | HCT 116 | C665(0.71); C256(0.92); C728(0.93); C739(0.93) | LDD0632 | [5] |
| LDCM0316 | AC53 | HCT 116 | C665(0.72); C728(0.77); C739(0.77); C256(0.84) | LDD0633 | [5] |
| LDCM0317 | AC54 | HCT 116 | C777(0.73); C665(0.85); C728(0.96); C739(0.96) | LDD0634 | [5] |
| LDCM0318 | AC55 | HCT 116 | C256(0.66); C728(0.76); C739(0.76); C665(0.76) | LDD0635 | [5] |
| LDCM0319 | AC56 | HCT 116 | C665(0.57); C777(0.63); C728(0.73); C739(0.73) | LDD0636 | [5] |
| LDCM0320 | AC57 | HCT 116 | C777(0.79); C265(0.90); C256(0.94); C665(1.10) | LDD0637 | [5] |
| LDCM0321 | AC58 | HCT 116 | C777(0.69); C265(0.84); C665(0.87); C256(0.92) | LDD0638 | [5] |
| LDCM0322 | AC59 | HCT 116 | C777(0.66); C256(0.68); C265(0.90); C665(1.02) | LDD0639 | [5] |
| LDCM0323 | AC6 | HCT 116 | C256(0.78); C777(0.81); C665(1.01); C265(1.03) | LDD0640 | [5] |
| LDCM0324 | AC60 | HCT 116 | C256(0.61); C777(0.70); C265(0.97); C665(1.13) | LDD0641 | [5] |
| LDCM0325 | AC61 | HCT 116 | C777(0.79); C256(0.92); C265(1.01); C665(1.03) | LDD0642 | [5] |
| LDCM0326 | AC62 | HCT 116 | C777(0.68); C256(0.70); C265(0.94); C665(1.08) | LDD0643 | [5] |
| LDCM0327 | AC63 | HCT 116 | C256(0.51); C777(0.69); C265(0.84); C1034(1.30) | LDD0644 | [5] |
| LDCM0328 | AC64 | HCT 116 | C256(0.67); C777(0.70); C265(0.97); C665(1.30) | LDD0645 | [5] |
| LDCM0329 | AC65 | HCT 116 | C777(0.75); C665(0.92); C256(0.92); C265(1.18) | LDD0646 | [5] |
| LDCM0330 | AC66 | HCT 116 | C777(0.79); C665(0.88); C256(0.92); C1034(1.27) | LDD0647 | [5] |
| LDCM0331 | AC67 | HCT 116 | C256(0.59); C777(0.60); C265(0.98); C665(1.04) | LDD0648 | [5] |
| LDCM0332 | AC68 | HCT 116 | C728(0.41); C739(0.41); C665(0.64); C256(0.68) | LDD0649 | [5] |
| LDCM0333 | AC69 | HCT 116 | C728(0.51); C739(0.51); C256(0.53); C265(0.73) | LDD0650 | [5] |
| LDCM0334 | AC7 | HCT 116 | C777(0.94); C1034(1.08); C665(1.24); C256(1.38) | LDD0651 | [5] |
| LDCM0335 | AC70 | HCT 116 | C728(0.28); C739(0.28); C256(0.36); C665(0.49) | LDD0652 | [5] |
| LDCM0336 | AC71 | HCT 116 | C728(0.48); C739(0.48); C256(0.78); C777(0.87) | LDD0653 | [5] |
| LDCM0337 | AC72 | HCT 116 | C256(0.55); C665(0.57); C777(0.74); C728(0.76) | LDD0654 | [5] |
| LDCM0338 | AC73 | HCT 116 | C256(0.29); C728(0.33); C739(0.33); C665(0.49) | LDD0655 | [5] |
| LDCM0339 | AC74 | HCT 116 | C256(0.30); C728(0.33); C739(0.33); C777(0.64) | LDD0656 | [5] |
| LDCM0340 | AC75 | HCT 116 | C256(0.32); C728(0.39); C739(0.39); C665(0.53) | LDD0657 | [5] |
| LDCM0341 | AC76 | HCT 116 | C728(0.40); C739(0.40); C256(0.44); C777(0.61) | LDD0658 | [5] |
| LDCM0342 | AC77 | HCT 116 | C256(0.39); C728(0.43); C739(0.43); C665(0.55) | LDD0659 | [5] |
| LDCM0343 | AC78 | HCT 116 | C665(0.61); C728(0.62); C739(0.62); C256(0.77) | LDD0660 | [5] |
| LDCM0344 | AC79 | HCT 116 | C665(0.46); C728(0.55); C739(0.55); C256(0.75) | LDD0661 | [5] |
| LDCM0345 | AC8 | HCT 116 | C777(0.82); C256(0.97); C1034(1.19); C665(1.31) | LDD0662 | [5] |
| LDCM0346 | AC80 | HCT 116 | C256(0.49); C728(0.55); C739(0.55); C265(0.70) | LDD0663 | [5] |
| LDCM0347 | AC81 | HCT 116 | C665(0.71); C777(0.75); C1034(1.05); C265(1.08) | LDD0664 | [5] |
| LDCM0348 | AC82 | HCT 116 | C728(0.29); C739(0.29); C256(0.40); C777(0.65) | LDD0665 | [5] |
| LDCM0349 | AC83 | HCT 116 | C728(0.53); C739(0.53); C777(0.68); C665(0.84) | LDD0666 | [5] |
| LDCM0350 | AC84 | HCT 116 | C728(0.64); C739(0.64); C777(0.66); C665(0.82) | LDD0667 | [5] |
| LDCM0351 | AC85 | HCT 116 | C728(0.59); C739(0.59); C665(0.75); C777(0.79) | LDD0668 | [5] |
| LDCM0352 | AC86 | HCT 116 | C728(0.36); C739(0.36); C665(0.70); C777(0.78) | LDD0669 | [5] |
| LDCM0353 | AC87 | HCT 116 | C728(0.40); C739(0.40); C777(0.77); C665(0.82) | LDD0670 | [5] |
| LDCM0354 | AC88 | HCT 116 | C728(0.41); C739(0.41); C665(0.72); C777(0.73) | LDD0671 | [5] |
| LDCM0355 | AC89 | HCT 116 | C728(0.38); C739(0.38); C777(0.71); C665(0.76) | LDD0672 | [5] |
| LDCM0357 | AC90 | HCT 116 | C728(0.40); C739(0.40); C265(0.82); C665(0.84) | LDD0674 | [5] |
| LDCM0358 | AC91 | HCT 116 | C728(0.66); C739(0.66); C777(0.73); C665(0.96) | LDD0675 | [5] |
| LDCM0359 | AC92 | HCT 116 | C728(0.58); C739(0.58); C777(0.70); C665(0.84) | LDD0676 | [5] |
| LDCM0360 | AC93 | HCT 116 | C777(0.82); C728(0.84); C739(0.84); C665(0.88) | LDD0677 | [5] |
| LDCM0361 | AC94 | HCT 116 | C777(0.94); C265(0.96); C1034(1.25); C665(1.44) | LDD0678 | [5] |
| LDCM0362 | AC95 | HCT 116 | C728(0.45); C739(0.45); C665(0.78); C777(0.97) | LDD0679 | [5] |
| LDCM0363 | AC96 | HCT 116 | C777(0.87); C265(0.87); C1034(1.16); C665(1.18) | LDD0680 | [5] |
| LDCM0364 | AC97 | HCT 116 | C728(0.36); C739(0.36); C777(0.64); C665(0.67) | LDD0681 | [5] |
| LDCM0365 | AC98 | HCT 116 | C777(0.33); C665(0.46); C256(0.60); C728(0.82) | LDD0682 | [5] |
| LDCM0366 | AC99 | HCT 116 | C665(0.53); C777(0.59); C265(0.64); C256(0.70) | LDD0683 | [5] |
| LDCM0545 | Acetamide | MDA-MB-231 | C486(0.30) | LDD2138 | [3] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C486(0.75) | LDD2113 | [3] |
| LDCM0248 | AKOS034007472 | HCT 116 | C1034(1.05); C256(1.27); C265(1.33); C665(1.50) | LDD0565 | [5] |
| LDCM0356 | AKOS034007680 | HCT 116 | C777(0.97); C1034(1.11); C256(1.51); C265(1.57) | LDD0673 | [5] |
| LDCM0275 | AKOS034007705 | HCT 116 | C256(0.61); C777(0.69); C665(1.08); C265(1.31) | LDD0592 | [5] |
| LDCM0020 | ARS-1620 | HCC44 | C1034(0.91) | LDD0078 | [5] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C486(0.57) | LDD2091 | [3] |
| LDCM0108 | Chloroacetamide | HeLa | C777(0.00); C486(0.00); C256(0.00); C265(0.00) | LDD0222 | [15] |
| LDCM0367 | CL1 | HCT 116 | C265(0.77); C665(0.84); C256(0.95); C1034(0.95) | LDD0684 | [5] |
| LDCM0368 | CL10 | HCT 116 | C777(0.82); C665(1.05); C265(1.07); C256(1.10) | LDD0685 | [5] |
| LDCM0369 | CL100 | HCT 116 | C1034(1.31); C777(1.43); C256(1.49); C665(1.55) | LDD0686 | [5] |
| LDCM0370 | CL101 | HCT 116 | C777(0.74); C256(1.05); C1034(1.16); C665(1.32) | LDD0687 | [5] |
| LDCM0371 | CL102 | HCT 116 | C777(0.91); C1034(1.02); C665(1.42); C265(1.46) | LDD0688 | [5] |
| LDCM0372 | CL103 | HCT 116 | C777(0.98); C1034(1.02); C265(1.41); C665(1.56) | LDD0689 | [5] |
| LDCM0373 | CL104 | HCT 116 | C777(0.87); C1034(1.23); C256(1.24); C265(1.47) | LDD0690 | [5] |
| LDCM0374 | CL105 | HCT 116 | C256(0.78); C265(0.80); C777(0.97); C1034(1.23) | LDD0691 | [5] |
| LDCM0375 | CL106 | HCT 116 | C265(0.66); C256(0.67); C777(1.17); C665(1.66) | LDD0692 | [5] |
| LDCM0376 | CL107 | HCT 116 | C256(0.71); C265(0.73); C777(0.89); C665(1.44) | LDD0693 | [5] |
| LDCM0377 | CL108 | HCT 116 | C265(0.70); C256(0.70); C777(1.10); C665(1.33) | LDD0694 | [5] |
| LDCM0378 | CL109 | HCT 116 | C256(0.83); C265(0.86); C777(0.96); C665(1.08) | LDD0695 | [5] |
| LDCM0379 | CL11 | HCT 116 | C777(0.67); C665(0.89); C256(1.27); C1034(1.46) | LDD0696 | [5] |
| LDCM0380 | CL110 | HCT 116 | C256(0.75); C777(0.92); C265(0.95); C665(1.03) | LDD0697 | [5] |
| LDCM0381 | CL111 | HCT 116 | C256(0.83); C777(0.93); C665(0.95); C265(1.14) | LDD0698 | [5] |
| LDCM0382 | CL112 | HCT 116 | C665(0.74); C777(0.84); C728(0.86); C739(0.86) | LDD0699 | [5] |
| LDCM0383 | CL113 | HCT 116 | C728(0.43); C739(0.43); C777(0.56); C665(0.71) | LDD0700 | [5] |
| LDCM0384 | CL114 | HCT 116 | C728(0.45); C739(0.45); C777(0.62); C665(0.74) | LDD0701 | [5] |
| LDCM0385 | CL115 | HCT 116 | C728(0.49); C739(0.49); C777(0.60); C665(0.97) | LDD0702 | [5] |
| LDCM0386 | CL116 | HCT 116 | C777(0.74); C665(0.83); C265(1.06); C1034(1.15) | LDD0703 | [5] |
| LDCM0387 | CL117 | HCT 116 | C256(0.48); C777(0.59); C265(0.68); C728(0.72) | LDD0704 | [5] |
| LDCM0388 | CL118 | HCT 116 | C728(0.84); C739(0.84); C265(0.87); C256(0.87) | LDD0705 | [5] |
| LDCM0389 | CL119 | HCT 116 | C256(0.74); C777(0.76); C265(0.81); C665(0.92) | LDD0706 | [5] |
| LDCM0390 | CL12 | HCT 116 | C777(0.59); C665(0.92); C256(1.15); C1034(1.31) | LDD0707 | [5] |
| LDCM0391 | CL120 | HCT 116 | C256(0.89); C728(0.94); C739(0.94); C1034(0.98) | LDD0708 | [5] |
| LDCM0392 | CL121 | HCT 116 | C665(0.79); C777(0.83); C256(0.93); C265(1.14) | LDD0709 | [5] |
| LDCM0393 | CL122 | HCT 116 | C665(0.72); C728(0.76); C739(0.76); C777(0.97) | LDD0710 | [5] |
| LDCM0394 | CL123 | HCT 116 | C777(0.76); C728(0.83); C739(0.83); C665(0.85) | LDD0711 | [5] |
| LDCM0395 | CL124 | HCT 116 | C777(0.69); C665(0.69); C728(0.71); C739(0.71) | LDD0712 | [5] |
| LDCM0396 | CL125 | HCT 116 | C256(0.85); C665(0.93); C777(0.97); C1034(1.04) | LDD0713 | [5] |
| LDCM0397 | CL126 | HCT 116 | C777(0.77); C665(0.91); C265(1.01); C256(1.05) | LDD0714 | [5] |
| LDCM0398 | CL127 | HCT 116 | C777(0.82); C256(0.86); C265(0.89); C665(1.02) | LDD0715 | [5] |
| LDCM0399 | CL128 | HCT 116 | C265(0.77); C777(0.78); C256(0.91); C665(1.11) | LDD0716 | [5] |
| LDCM0400 | CL13 | HCT 116 | C777(0.75); C665(1.03); C265(1.04); C1034(1.17) | LDD0717 | [5] |
| LDCM0401 | CL14 | HCT 116 | C265(0.79); C665(0.97); C1034(1.05); C777(1.06) | LDD0718 | [5] |
| LDCM0402 | CL15 | HCT 116 | C777(0.87); C665(0.88); C265(1.10); C1034(1.10) | LDD0719 | [5] |
| LDCM0403 | CL16 | HCT 116 | C256(0.79); C665(0.84); C1034(1.06); C265(1.11) | LDD0720 | [5] |
| LDCM0404 | CL17 | HCT 116 | C256(0.85); C1034(0.97); C665(0.98); C265(1.08) | LDD0721 | [5] |
| LDCM0405 | CL18 | HCT 116 | C256(0.81); C1034(1.06); C665(1.06); C265(1.21) | LDD0722 | [5] |
| LDCM0406 | CL19 | HCT 116 | C256(0.73); C728(0.86); C739(0.86); C1034(1.07) | LDD0723 | [5] |
| LDCM0407 | CL2 | HCT 116 | C265(0.82); C665(0.89); C1034(0.96); C256(1.07) | LDD0724 | [5] |
| LDCM0408 | CL20 | HCT 116 | C256(0.64); C728(0.85); C739(0.85); C665(0.94) | LDD0725 | [5] |
| LDCM0409 | CL21 | HCT 116 | C728(0.58); C739(0.58); C256(0.75); C665(0.96) | LDD0726 | [5] |
| LDCM0410 | CL22 | HCT 116 | C728(0.27); C739(0.27); C256(0.66); C665(0.97) | LDD0727 | [5] |
| LDCM0411 | CL23 | HCT 116 | C256(0.73); C728(0.76); C739(0.76); C665(0.79) | LDD0728 | [5] |
| LDCM0412 | CL24 | HCT 116 | C728(0.42); C739(0.42); C256(0.64); C665(0.99) | LDD0729 | [5] |
| LDCM0413 | CL25 | HCT 116 | C728(0.45); C739(0.45); C1034(1.44); C256(0.73) | LDD0730 | [5] |
| LDCM0414 | CL26 | HCT 116 | C728(0.73); C739(0.73); C1034(1.14); C256(0.69) | LDD0731 | [5] |
| LDCM0415 | CL27 | HCT 116 | C728(0.59); C739(0.59); C1034(0.99); C256(0.75) | LDD0732 | [5] |
| LDCM0416 | CL28 | HCT 116 | C728(1.16); C739(1.16); C1034(1.21); C256(0.66) | LDD0733 | [5] |
| LDCM0417 | CL29 | HCT 116 | C728(0.62); C739(0.62); C1034(1.20); C256(0.71) | LDD0734 | [5] |
| LDCM0418 | CL3 | HCT 116 | C1034(0.93); C256(1.11); C265(0.94); C665(0.83) | LDD0735 | [5] |
| LDCM0419 | CL30 | HCT 116 | C728(0.91); C739(0.91); C1034(0.99); C256(0.75) | LDD0736 | [5] |
| LDCM0420 | CL31 | HCT 116 | C728(1.12); C739(1.12); C1034(1.17); C256(1.16) | LDD0737 | [5] |
| LDCM0421 | CL32 | HCT 116 | C728(1.74); C739(1.74); C1034(1.16); C256(1.04) | LDD0738 | [5] |
| LDCM0422 | CL33 | HCT 116 | C728(1.77); C739(1.77); C1034(1.22); C256(1.22) | LDD0739 | [5] |
| LDCM0423 | CL34 | HCT 116 | C728(0.89); C739(0.89); C1034(1.36); C256(0.95) | LDD0740 | [5] |
| LDCM0424 | CL35 | HCT 116 | C728(1.77); C739(1.77); C1034(1.57); C256(1.00) | LDD0741 | [5] |
| LDCM0425 | CL36 | HCT 116 | C728(1.15); C739(1.15); C1034(1.50); C256(0.95) | LDD0742 | [5] |
| LDCM0426 | CL37 | HCT 116 | C728(1.20); C739(1.20); C1034(1.61); C256(1.00) | LDD0743 | [5] |
| LDCM0428 | CL39 | HCT 116 | C728(1.01); C739(1.01); C1034(1.44); C256(1.11) | LDD0745 | [5] |
| LDCM0429 | CL4 | HCT 116 | C1034(0.96); C256(1.19); C265(0.98); C665(0.90) | LDD0746 | [5] |
| LDCM0430 | CL40 | HCT 116 | C728(1.76); C739(1.76); C1034(1.37); C256(1.16) | LDD0747 | [5] |
| LDCM0431 | CL41 | HCT 116 | C728(1.00); C739(1.00); C1034(1.32); C256(1.08) | LDD0748 | [5] |
| LDCM0432 | CL42 | HCT 116 | C728(0.94); C739(0.94); C1034(2.20); C256(1.15) | LDD0749 | [5] |
| LDCM0433 | CL43 | HCT 116 | C728(1.08); C739(1.08); C1034(1.63); C256(1.09) | LDD0750 | [5] |
| LDCM0434 | CL44 | HCT 116 | C728(1.35); C739(1.35); C1034(1.39); C256(1.24) | LDD0751 | [5] |
| LDCM0435 | CL45 | HCT 116 | C728(0.70); C739(0.70); C1034(1.58); C256(1.30) | LDD0752 | [5] |
| LDCM0436 | CL46 | HCT 116 | C728(0.78); C739(0.78); C1034(1.13); C265(0.62) | LDD0753 | [5] |
| LDCM0437 | CL47 | HCT 116 | C728(0.86); C739(0.86); C1034(1.20); C265(1.02) | LDD0754 | [5] |
| LDCM0438 | CL48 | HCT 116 | C728(0.98); C739(0.98); C1034(1.27); C265(0.87) | LDD0755 | [5] |
| LDCM0439 | CL49 | HCT 116 | C728(1.15); C739(1.15); C1034(1.07); C265(0.96) | LDD0756 | [5] |
| LDCM0440 | CL5 | HCT 116 | C1034(0.98); C256(1.17); C265(0.96); C665(1.03) | LDD0757 | [5] |
| LDCM0441 | CL50 | HCT 116 | C728(0.86); C739(0.86); C1034(1.19); C265(0.87) | LDD0758 | [5] |
| LDCM0442 | CL51 | HCT 116 | C728(0.73); C739(0.73); C1034(1.27); C265(0.76) | LDD0759 | [5] |
| LDCM0443 | CL52 | HCT 116 | C728(0.70); C739(0.70); C1034(1.35); C265(0.89) | LDD0760 | [5] |
| LDCM0444 | CL53 | HCT 116 | C728(1.31); C739(1.31); C1034(1.17); C265(1.00) | LDD0761 | [5] |
| LDCM0445 | CL54 | HCT 116 | C728(0.94); C739(0.94); C1034(1.26); C265(0.70) | LDD0762 | [5] |
| LDCM0446 | CL55 | HCT 116 | C728(0.78); C739(0.78); C1034(1.10); C265(0.78) | LDD0763 | [5] |
| LDCM0447 | CL56 | HCT 116 | C728(1.70); C739(1.70); C1034(1.12); C265(1.06) | LDD0764 | [5] |
| LDCM0448 | CL57 | HCT 116 | C728(1.05); C739(1.05); C1034(1.06); C265(0.98) | LDD0765 | [5] |
| LDCM0449 | CL58 | HCT 116 | C728(0.82); C739(0.82); C1034(1.09); C265(0.95) | LDD0766 | [5] |
| LDCM0450 | CL59 | HCT 116 | C728(0.93); C739(0.93); C1034(1.02); C265(0.84) | LDD0767 | [5] |
| LDCM0451 | CL6 | HCT 116 | C1034(1.05); C256(1.19); C265(0.94); C665(0.94) | LDD0768 | [5] |
| LDCM0452 | CL60 | HCT 116 | C728(0.81); C739(0.81); C1034(1.10); C265(0.92) | LDD0769 | [5] |
| LDCM0453 | CL61 | HCT 116 | C728(1.33); C739(1.33); C1034(1.19); C256(0.88) | LDD0770 | [5] |
| LDCM0454 | CL62 | HCT 116 | C728(0.81); C739(0.81); C1034(1.28); C256(0.90) | LDD0771 | [5] |
| LDCM0455 | CL63 | HCT 116 | C728(1.03); C739(1.03); C1034(1.31); C256(0.76) | LDD0772 | [5] |
| LDCM0456 | CL64 | HCT 116 | C728(1.40); C739(1.40); C1034(1.39); C256(0.78) | LDD0773 | [5] |
| LDCM0457 | CL65 | HCT 116 | C728(1.27); C739(1.27); C1034(1.17); C256(0.94) | LDD0774 | [5] |
| LDCM0458 | CL66 | HCT 116 | C728(0.98); C739(0.98); C1034(1.60); C256(0.71) | LDD0775 | [5] |
| LDCM0459 | CL67 | HCT 116 | C728(0.92); C739(0.92); C1034(1.51); C256(0.78) | LDD0776 | [5] |
| LDCM0460 | CL68 | HCT 116 | C728(1.02); C739(1.02); C1034(1.23); C256(0.76) | LDD0777 | [5] |
| LDCM0461 | CL69 | HCT 116 | C728(1.16); C739(1.16); C1034(1.12); C256(0.98) | LDD0778 | [5] |
| LDCM0462 | CL7 | HCT 116 | C1034(1.20); C256(1.22); C265(0.96); C665(1.20) | LDD0779 | [5] |
| LDCM0463 | CL70 | HCT 116 | C728(2.08); C739(2.08); C1034(1.21); C256(0.77) | LDD0780 | [5] |
| LDCM0464 | CL71 | HCT 116 | C728(0.96); C739(0.96); C1034(1.41); C256(0.65) | LDD0781 | [5] |
| LDCM0465 | CL72 | HCT 116 | C728(1.18); C739(1.18); C1034(1.09); C256(0.69) | LDD0782 | [5] |
| LDCM0466 | CL73 | HCT 116 | C728(0.84); C739(0.84); C1034(1.35); C256(0.70) | LDD0783 | [5] |
| LDCM0467 | CL74 | HCT 116 | C728(1.06); C739(1.06); C1034(1.28); C256(0.72) | LDD0784 | [5] |
| LDCM0469 | CL76 | HCT 116 | C1034(1.17); C256(1.30); C265(1.19); C665(0.90) | LDD0786 | [5] |
| LDCM0470 | CL77 | HCT 116 | C1034(1.26); C256(1.19); C265(1.05); C665(0.70) | LDD0787 | [5] |
| LDCM0471 | CL78 | HCT 116 | C1034(1.26); C256(0.89); C265(0.99); C665(0.82) | LDD0788 | [5] |
| LDCM0472 | CL79 | HCT 116 | C1034(1.53); C256(1.28); C265(1.23); C665(0.69) | LDD0789 | [5] |
| LDCM0473 | CL8 | HCT 116 | C1034(1.71); C256(0.92); C265(1.18); C665(0.67) | LDD0790 | [5] |
| LDCM0474 | CL80 | HCT 116 | C1034(1.03); C256(1.23); C265(1.18); C665(0.96) | LDD0791 | [5] |
| LDCM0475 | CL81 | HCT 116 | C1034(1.31); C256(1.20); C265(1.08); C665(0.78) | LDD0792 | [5] |
| LDCM0476 | CL82 | HCT 116 | C1034(1.71); C256(1.62); C265(1.35); C665(0.70) | LDD0793 | [5] |
| LDCM0477 | CL83 | HCT 116 | C1034(1.58); C256(1.62); C265(1.09); C665(0.67) | LDD0794 | [5] |
| LDCM0478 | CL84 | HCT 116 | C1034(2.29); C256(1.37); C265(1.16); C665(0.68) | LDD0795 | [5] |
| LDCM0479 | CL85 | HCT 116 | C1034(1.31); C256(1.37); C265(1.17); C665(0.78) | LDD0796 | [5] |
| LDCM0480 | CL86 | HCT 116 | C1034(1.18); C256(1.28); C265(0.99); C665(0.86) | LDD0797 | [5] |
| LDCM0481 | CL87 | HCT 116 | C1034(1.21); C256(1.52); C265(1.01); C665(0.84) | LDD0798 | [5] |
| LDCM0482 | CL88 | HCT 116 | C1034(1.52); C256(1.73); C265(1.13); C665(0.71) | LDD0799 | [5] |
| LDCM0483 | CL89 | HCT 116 | C1034(1.99); C256(1.08); C265(1.15); C665(0.67) | LDD0800 | [5] |
| LDCM0484 | CL9 | HCT 116 | C1034(1.27); C256(1.10); C265(0.84); C665(0.80) | LDD0801 | [5] |
| LDCM0485 | CL90 | HCT 116 | C1034(1.13); C256(1.87); C265(1.18); C665(0.79) | LDD0802 | [5] |
| LDCM0486 | CL91 | HCT 116 | C1034(1.10); C256(0.66); C265(1.39); C665(1.01) | LDD0803 | [5] |
| LDCM0487 | CL92 | HCT 116 | C1034(1.03); C256(0.84); C265(1.42); C665(1.10) | LDD0804 | [5] |
| LDCM0488 | CL93 | HCT 116 | C1034(1.06); C256(1.20); C265(1.15); C665(1.32) | LDD0805 | [5] |
| LDCM0489 | CL94 | HCT 116 | C1034(1.36); C256(1.94); C265(1.55); C665(1.62) | LDD0806 | [5] |
| LDCM0490 | CL95 | HCT 116 | C1034(1.34); C256(0.97); C265(1.43); C665(1.13) | LDD0807 | [5] |
| LDCM0491 | CL96 | HCT 116 | C1034(1.07); C256(0.89); C265(1.39); C665(1.16) | LDD0808 | [5] |
| LDCM0492 | CL97 | HCT 116 | C1034(1.10); C256(0.88); C265(1.35); C665(1.06) | LDD0809 | [5] |
| LDCM0493 | CL98 | HCT 116 | C1034(1.23); C256(1.18); C265(1.61); C665(1.37) | LDD0810 | [5] |
| LDCM0494 | CL99 | HCT 116 | C1034(1.25); C256(1.58); C265(1.46); C665(1.59) | LDD0811 | [5] |
| LDCM0634 | CY-0357 | Hep-G2 | C1034(0.56) | LDD2228 | [17] |
| LDCM0495 | E2913 | HEK-293T | C1034(1.15); C171(0.89); C486(1.00); C665(1.07) | LDD1698 | [18] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C265(1.67); C486(1.37); C256(1.30) | LDD1702 | [3] |
| LDCM0574 | Fragment12 | Ramos | C1034(0.83) | LDD2191 | [19] |
| LDCM0575 | Fragment13 | Ramos | C1034(1.71) | LDD2192 | [19] |
| LDCM0582 | Fragment23 | Ramos | C1034(1.47) | LDD2196 | [19] |
| LDCM0586 | Fragment28 | Ramos | C1034(1.11) | LDD2198 | [19] |
| LDCM0468 | Fragment33 | HCT 116 | C728(1.31); C739(1.31); C1034(1.18); C256(0.76) | LDD0785 | [5] |
| LDCM0596 | Fragment38 | Ramos | C1034(0.51) | LDD2203 | [19] |
| LDCM0427 | Fragment51 | HCT 116 | C728(0.72); C739(0.72); C1034(2.01); C256(0.93) | LDD0744 | [5] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [15] |
| LDCM0022 | KB02 | HCT 116 | C1034(3.21) | LDD0080 | [5] |
| LDCM0023 | KB03 | HCT 116 | C1034(1.16) | LDD0081 | [5] |
| LDCM0024 | KB05 | HCT 116 | C1034(2.06) | LDD0082 | [5] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C486(0.43) | LDD2121 | [3] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0224 | [15] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C486(0.38) | LDD2089 | [3] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C486(0.88) | LDD2093 | [3] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C486(0.82) | LDD2097 | [3] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C486(0.95) | LDD2099 | [3] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C486(0.92) | LDD2101 | [3] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C486(1.02) | LDD2107 | [3] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C486(0.51) | LDD2109 | [3] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C486(0.83) | LDD2111 | [3] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C486(0.38) | LDD2115 | [3] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C486(2.10) | LDD2119 | [3] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C486(0.73) | LDD2123 | [3] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C486(0.62) | LDD2125 | [3] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C486(0.84); C1034(0.86) | LDD2127 | [3] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C486(0.94) | LDD2129 | [3] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C486(0.42) | LDD2133 | [3] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C486(0.40) | LDD2134 | [3] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C486(1.25) | LDD2135 | [3] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C486(1.14); C777(0.84) | LDD2136 | [3] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C486(1.00) | LDD2137 | [3] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C486(2.27) | LDD1700 | [3] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C486(0.69); C777(0.65); C1034(1.10) | LDD2140 | [3] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C486(0.50) | LDD2143 | [3] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C486(1.82); C777(2.98) | LDD2144 | [3] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C486(0.86); C777(0.89) | LDD2146 | [3] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C486(0.49); C777(0.36) | LDD2150 | [3] |
| LDCM0131 | RA190 | MM1.R | C1034(1.14) | LDD0304 | [20] |
| LDCM0170 | Structure8 | Ramos | 5.25 | LDD0433 | [21] |
| LDCM0021 | THZ1 | HCT 116 | C1034(1.21); C665(1.17); C777(1.00); C265(0.93) | LDD2173 | [5] |
The Interaction Atlas With This Target
References
