Details of the Target
General Information of Target
| Target ID | LDTP12665 | |||||
|---|---|---|---|---|---|---|
| Target Name | Cell cycle control protein 50A (TMEM30A) | |||||
| Gene Name | TMEM30A | |||||
| Gene ID | 55754 | |||||
| Synonyms |
C6orf67; CDC50A; Cell cycle control protein 50A; P4-ATPase flippase complex beta subunit TMEM30A; Transmembrane protein 30A |
|||||
| 3D Structure | ||||||
| Sequence |
MARRRSQRVCASGPSMLNSARGAPELLRGTATNAEVSAAAAGATGSEELPPGDRGCRNGG
GRGPAATTSSTGVAVGAEHGEDSLSRKPDPEPGRMDHHQPGTGRYQVLLNEEDNSESSAI EQPPTSNPAPQIVQAASSAPALETDSSPPPYSSITVEVPTTSDTEVYGEFYPVPPPYSVA TSLPTYDEAEKAKAAAMAAAAAETSQRIQEEECPPRDDFSDADQLRVGNDGIFMLAFFMA FIFNWLGFCLSFCITNTIAGRYGAICGFGLSLIKWILIVRFSDYFTGYFNGQYWLWWIFL VLGLLLFFRGFVNYLKVRNMSESMAAAHRTRYFFLL |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
CDC50/LEM3 family
|
|||||
| Subcellular location |
Membrane
|
|||||
| Function |
Accessory component of a P4-ATPase flippase complex which catalyzes the hydrolysis of ATP coupled to the transport of aminophospholipids from the outer to the inner leaflet of various membranes and ensures the maintenance of asymmetric distribution of phospholipids. Phospholipid translocation seems also to be implicated in vesicle formation and in uptake of lipid signaling molecules. The beta subunit may assist in binding of the phospholipid substrate. Required for the proper folding, assembly and ER to Golgi exit of the ATP8A2:TMEM30A flippase complex. ATP8A2:TMEM30A may be involved in regulation of neurite outgrowth, and, reconstituted to liposomes, predomiminantly transports phosphatidylserine (PS) and to a lesser extent phosphatidylethanolamine (PE). The ATP8A1:TMEM30A flippase complex seems to play a role in regulation of cell migration probably involving flippase-mediated translocation of phosphatidylethanolamine (PE) at the plasma membrane. Required for the formation of the ATP8A2, ATP8B1 and ATP8B2 P-type ATPAse intermediate phosphoenzymes. Involved in uptake of platelet-activating factor (PAF), synthetic drug alkylphospholipid edelfosine, and, probably in association with ATP8B1, of perifosine. Also mediates the export of alpha subunits ATP8A1, ATP8B1, ATP8B2, ATP8B4, ATP10A, ATP10B, ATP10D, ATP11A, ATP11B and ATP11C from the ER to other membrane localizations.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| JURKAT | SNV: p.S177N | . | |||
| RPMI8226 | SNV: p.L187F | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DA-P2 Probe Info |
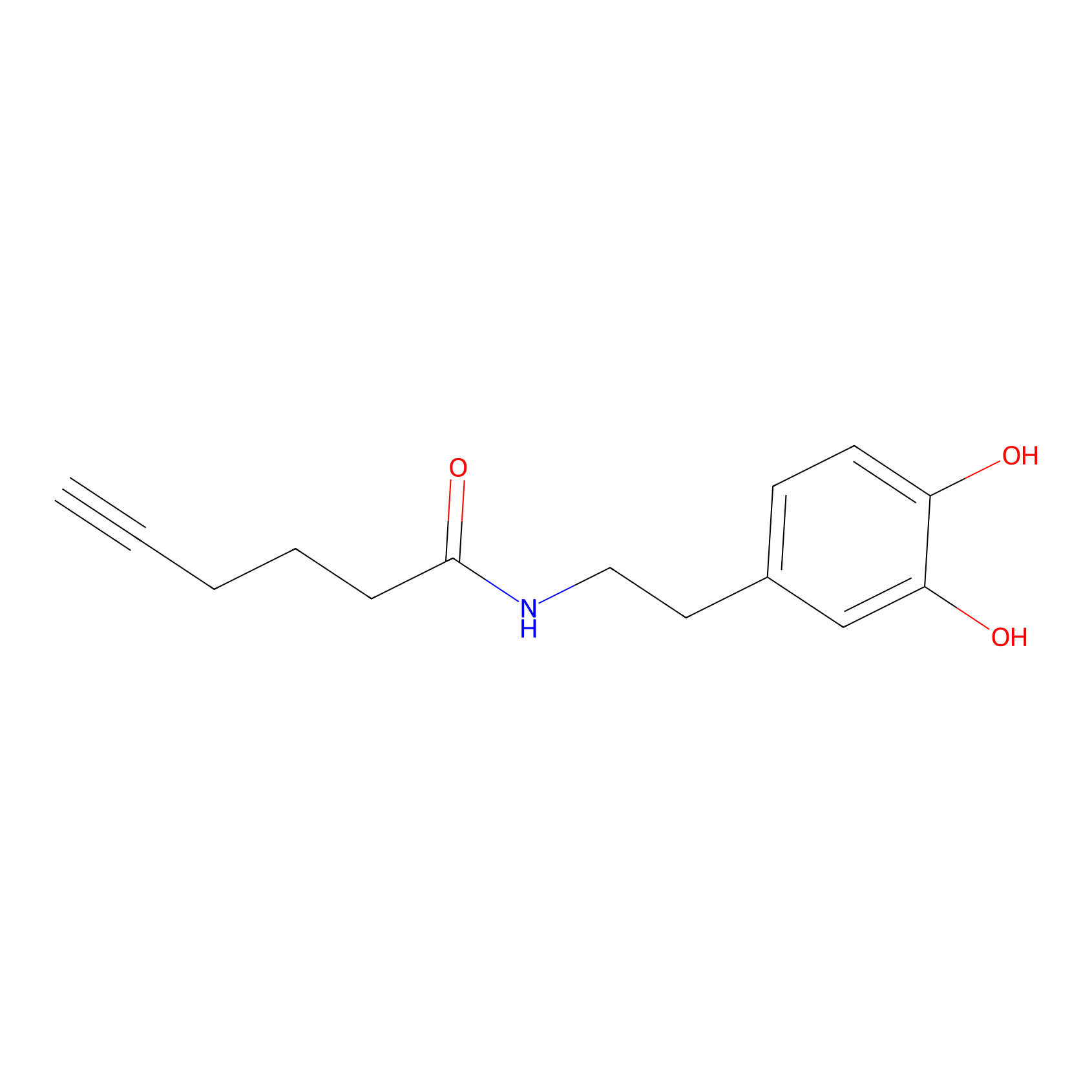 |
9.03 | LDD0348 | [1] | |
|
BTD Probe Info |
 |
C17(1.97) | LDD1699 | [2] | |
|
ONAyne Probe Info |
 |
K279(10.00) | LDD0275 | [3] | |
|
OPA-S-S-alkyne Probe Info |
 |
K279(1.73); K155(2.09) | LDD3494 | [4] | |
|
AHL-Pu-1 Probe Info |
 |
C17(2.05) | LDD0169 | [5] | |
|
DBIA Probe Info |
 |
C17(1.65) | LDD0080 | [6] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0165 | [7] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [8] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2224 | [8] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [9] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [9] | |
|
IPM Probe Info |
 |
N.A. | LDD0147 | [10] | |
|
PPMS Probe Info |
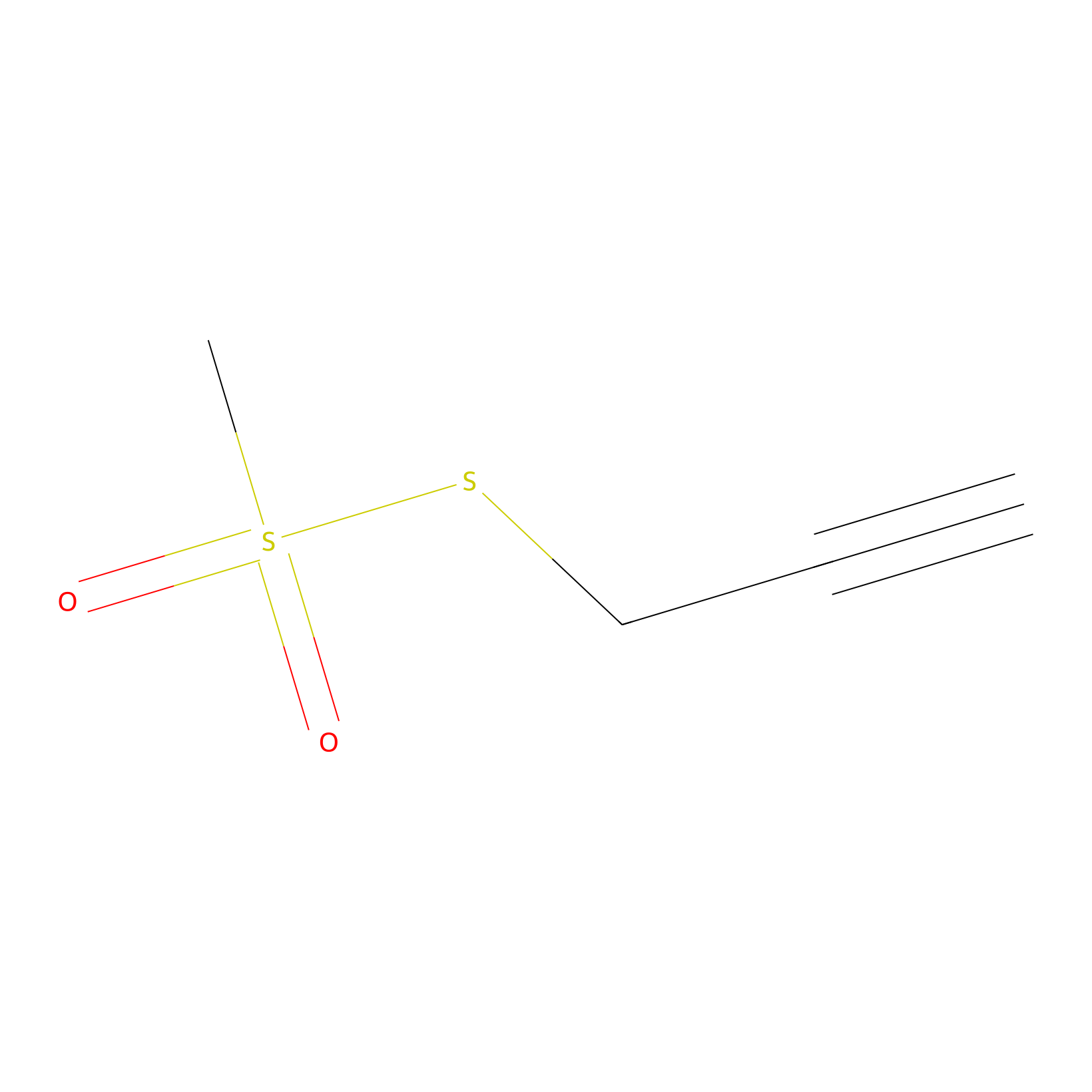 |
N.A. | LDD0008 | [9] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [9] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [9] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [11] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [12] | |
|
AOyne Probe Info |
 |
13.30 | LDD0443 | [13] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C106 Probe Info |
 |
18.38 | LDD1793 | [14] | |
|
C107 Probe Info |
 |
5.50 | LDD1794 | [14] | |
|
C170 Probe Info |
 |
8.82 | LDD1850 | [14] | |
|
C310 Probe Info |
 |
19.16 | LDD1977 | [14] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C17(2.05) | LDD0169 | [5] |
| LDCM0214 | AC1 | HEK-293T | C17(0.99) | LDD1507 | [15] |
| LDCM0215 | AC10 | HCT 116 | C17(1.17) | LDD0532 | [6] |
| LDCM0216 | AC100 | PaTu 8988t | C17(1.07) | LDD1095 | [6] |
| LDCM0217 | AC101 | PaTu 8988t | C17(0.95) | LDD1096 | [6] |
| LDCM0218 | AC102 | PaTu 8988t | C17(0.97) | LDD1097 | [6] |
| LDCM0219 | AC103 | PaTu 8988t | C17(1.28) | LDD1098 | [6] |
| LDCM0220 | AC104 | PaTu 8988t | C17(0.88) | LDD1099 | [6] |
| LDCM0221 | AC105 | PaTu 8988t | C17(1.01) | LDD1100 | [6] |
| LDCM0222 | AC106 | PaTu 8988t | C17(1.12) | LDD1101 | [6] |
| LDCM0223 | AC107 | PaTu 8988t | C17(0.94) | LDD1102 | [6] |
| LDCM0224 | AC108 | PaTu 8988t | C17(1.09) | LDD1103 | [6] |
| LDCM0225 | AC109 | PaTu 8988t | C17(1.24) | LDD1104 | [6] |
| LDCM0226 | AC11 | HCT 116 | C17(0.95) | LDD0543 | [6] |
| LDCM0227 | AC110 | PaTu 8988t | C17(1.23) | LDD1106 | [6] |
| LDCM0228 | AC111 | PaTu 8988t | C17(1.16) | LDD1107 | [6] |
| LDCM0229 | AC112 | PaTu 8988t | C17(1.03) | LDD1108 | [6] |
| LDCM0237 | AC12 | HCT 116 | C17(0.96) | LDD0554 | [6] |
| LDCM0259 | AC14 | HCT 116 | C17(1.02) | LDD0576 | [6] |
| LDCM0263 | AC143 | PaTu 8988t | C17(1.35) | LDD1142 | [6] |
| LDCM0264 | AC144 | PaTu 8988t | C17(0.89) | LDD1143 | [6] |
| LDCM0265 | AC145 | PaTu 8988t | C17(1.05) | LDD1144 | [6] |
| LDCM0266 | AC146 | PaTu 8988t | C17(0.86) | LDD1145 | [6] |
| LDCM0267 | AC147 | PaTu 8988t | C17(0.99) | LDD1146 | [6] |
| LDCM0268 | AC148 | PaTu 8988t | C17(1.16) | LDD1147 | [6] |
| LDCM0269 | AC149 | PaTu 8988t | C17(1.06) | LDD1148 | [6] |
| LDCM0270 | AC15 | HCT 116 | C17(0.99) | LDD0587 | [6] |
| LDCM0271 | AC150 | PaTu 8988t | C17(0.89) | LDD1150 | [6] |
| LDCM0272 | AC151 | PaTu 8988t | C17(1.16) | LDD1151 | [6] |
| LDCM0273 | AC152 | PaTu 8988t | C17(1.30) | LDD1152 | [6] |
| LDCM0274 | AC153 | PaTu 8988t | C17(1.16) | LDD1153 | [6] |
| LDCM0621 | AC154 | PaTu 8988t | C17(0.98) | LDD2166 | [6] |
| LDCM0622 | AC155 | PaTu 8988t | C17(0.85) | LDD2167 | [6] |
| LDCM0623 | AC156 | PaTu 8988t | C17(1.25) | LDD2168 | [6] |
| LDCM0624 | AC157 | PaTu 8988t | C17(0.75) | LDD2169 | [6] |
| LDCM0276 | AC17 | PaTu 8988t | C17(0.95) | LDD1155 | [6] |
| LDCM0277 | AC18 | PaTu 8988t | C17(1.33) | LDD1156 | [6] |
| LDCM0278 | AC19 | PaTu 8988t | C17(1.08) | LDD1157 | [6] |
| LDCM0279 | AC2 | HEK-293T | C17(1.06) | LDD1518 | [15] |
| LDCM0280 | AC20 | PaTu 8988t | C17(1.54) | LDD1159 | [6] |
| LDCM0281 | AC21 | PaTu 8988t | C17(1.28) | LDD1160 | [6] |
| LDCM0282 | AC22 | PaTu 8988t | C17(1.39) | LDD1161 | [6] |
| LDCM0283 | AC23 | PaTu 8988t | C17(1.82) | LDD1162 | [6] |
| LDCM0284 | AC24 | PaTu 8988t | C17(1.69) | LDD1163 | [6] |
| LDCM0285 | AC25 | HEK-293T | C17(1.10) | LDD1524 | [15] |
| LDCM0286 | AC26 | HEK-293T | C17(1.01) | LDD1525 | [15] |
| LDCM0287 | AC27 | HEK-293T | C17(1.08) | LDD1526 | [15] |
| LDCM0288 | AC28 | HEK-293T | C17(1.03); C157(1.06) | LDD1527 | [15] |
| LDCM0289 | AC29 | HEK-293T | C17(0.96) | LDD1528 | [15] |
| LDCM0290 | AC3 | HEK-293T | C17(0.96) | LDD1529 | [15] |
| LDCM0291 | AC30 | HEK-293T | C17(0.97); C91(1.15) | LDD1530 | [15] |
| LDCM0292 | AC31 | HEK-293T | C17(1.21); C157(0.95) | LDD1531 | [15] |
| LDCM0293 | AC32 | HEK-293T | C17(0.98); C94(1.12) | LDD1532 | [15] |
| LDCM0294 | AC33 | HEK-293T | C17(0.88) | LDD1533 | [15] |
| LDCM0295 | AC34 | HEK-293T | C17(1.07) | LDD1534 | [15] |
| LDCM0296 | AC35 | HEK-293T | C17(0.86) | LDD1535 | [15] |
| LDCM0297 | AC36 | HEK-293T | C17(0.92); C157(1.20) | LDD1536 | [15] |
| LDCM0298 | AC37 | HEK-293T | C17(1.04) | LDD1537 | [15] |
| LDCM0299 | AC38 | HEK-293T | C17(0.99); C91(1.10) | LDD1538 | [15] |
| LDCM0300 | AC39 | HEK-293T | C17(1.03); C157(0.90) | LDD1539 | [15] |
| LDCM0301 | AC4 | HEK-293T | C17(0.98); C157(1.29) | LDD1540 | [15] |
| LDCM0302 | AC40 | HEK-293T | C17(1.02); C94(0.96) | LDD1541 | [15] |
| LDCM0303 | AC41 | HEK-293T | C17(0.99) | LDD1542 | [15] |
| LDCM0304 | AC42 | HEK-293T | C17(1.09) | LDD1543 | [15] |
| LDCM0305 | AC43 | HEK-293T | C17(0.94) | LDD1544 | [15] |
| LDCM0306 | AC44 | HEK-293T | C17(0.96); C157(1.13) | LDD1545 | [15] |
| LDCM0307 | AC45 | HEK-293T | C17(1.11) | LDD1546 | [15] |
| LDCM0308 | AC46 | PaTu 8988t | C17(0.96) | LDD1187 | [6] |
| LDCM0309 | AC47 | PaTu 8988t | C17(0.99) | LDD1188 | [6] |
| LDCM0310 | AC48 | PaTu 8988t | C17(1.57) | LDD1189 | [6] |
| LDCM0311 | AC49 | PaTu 8988t | C17(1.29) | LDD1190 | [6] |
| LDCM0312 | AC5 | HEK-293T | C17(0.99) | LDD1551 | [15] |
| LDCM0313 | AC50 | PaTu 8988t | C17(1.33) | LDD1192 | [6] |
| LDCM0314 | AC51 | PaTu 8988t | C17(1.16) | LDD1193 | [6] |
| LDCM0315 | AC52 | PaTu 8988t | C17(1.39) | LDD1194 | [6] |
| LDCM0316 | AC53 | PaTu 8988t | C17(1.68) | LDD1195 | [6] |
| LDCM0317 | AC54 | PaTu 8988t | C17(1.91) | LDD1196 | [6] |
| LDCM0318 | AC55 | PaTu 8988t | C17(1.53) | LDD1197 | [6] |
| LDCM0319 | AC56 | PaTu 8988t | C17(1.49) | LDD1198 | [6] |
| LDCM0320 | AC57 | HCT 116 | C17(1.08) | LDD0637 | [6] |
| LDCM0321 | AC58 | HCT 116 | C17(1.23) | LDD0638 | [6] |
| LDCM0322 | AC59 | HCT 116 | C17(1.14) | LDD0639 | [6] |
| LDCM0323 | AC6 | HCT 116 | C17(1.11) | LDD0640 | [6] |
| LDCM0324 | AC60 | HCT 116 | C17(0.95) | LDD0641 | [6] |
| LDCM0325 | AC61 | HCT 116 | C17(1.10) | LDD0642 | [6] |
| LDCM0326 | AC62 | HCT 116 | C17(1.03) | LDD0643 | [6] |
| LDCM0327 | AC63 | HCT 116 | C17(1.01) | LDD0644 | [6] |
| LDCM0328 | AC64 | HCT 116 | C17(1.31) | LDD0645 | [6] |
| LDCM0329 | AC65 | HCT 116 | C17(1.60) | LDD0646 | [6] |
| LDCM0330 | AC66 | HCT 116 | C17(1.44) | LDD0647 | [6] |
| LDCM0331 | AC67 | HCT 116 | C17(1.07) | LDD0648 | [6] |
| LDCM0334 | AC7 | HCT 116 | C17(0.95) | LDD0651 | [6] |
| LDCM0345 | AC8 | HCT 116 | C17(1.01) | LDD0662 | [6] |
| LDCM0365 | AC98 | PaTu 8988t | C17(1.00) | LDD1244 | [6] |
| LDCM0366 | AC99 | PaTu 8988t | C17(1.03) | LDD1245 | [6] |
| LDCM0248 | AKOS034007472 | HCT 116 | C17(1.10) | LDD0565 | [6] |
| LDCM0356 | AKOS034007680 | HCT 116 | C17(0.97) | LDD0673 | [6] |
| LDCM0275 | AKOS034007705 | HCT 116 | C17(1.25) | LDD0592 | [6] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [11] |
| LDCM0367 | CL1 | HEK-293T | C17(0.90) | LDD1571 | [15] |
| LDCM0368 | CL10 | HEK-293T | C17(0.99); C91(1.32) | LDD1572 | [15] |
| LDCM0369 | CL100 | HEK-293T | C17(1.02); C94(1.06) | LDD1573 | [15] |
| LDCM0370 | CL101 | HCT 116 | C17(1.09) | LDD0687 | [6] |
| LDCM0371 | CL102 | HCT 116 | C17(0.89) | LDD0688 | [6] |
| LDCM0372 | CL103 | HCT 116 | C17(0.92) | LDD0689 | [6] |
| LDCM0373 | CL104 | HCT 116 | C17(0.85) | LDD0690 | [6] |
| LDCM0374 | CL105 | PaTu 8988t | C17(1.29) | LDD1253 | [6] |
| LDCM0375 | CL106 | PaTu 8988t | C17(1.19) | LDD1254 | [6] |
| LDCM0376 | CL107 | PaTu 8988t | C17(1.16) | LDD1255 | [6] |
| LDCM0377 | CL108 | PaTu 8988t | C17(1.37) | LDD1256 | [6] |
| LDCM0378 | CL109 | PaTu 8988t | C17(1.55) | LDD1257 | [6] |
| LDCM0379 | CL11 | HEK-293T | C17(1.08); C157(0.40) | LDD1583 | [15] |
| LDCM0380 | CL110 | PaTu 8988t | C17(1.65) | LDD1259 | [6] |
| LDCM0381 | CL111 | PaTu 8988t | C17(1.90) | LDD1260 | [6] |
| LDCM0382 | CL112 | HEK-293T | C17(1.01); C94(0.98) | LDD1586 | [15] |
| LDCM0383 | CL113 | HEK-293T | C17(0.91) | LDD1587 | [15] |
| LDCM0384 | CL114 | HEK-293T | C17(0.86) | LDD1588 | [15] |
| LDCM0385 | CL115 | HEK-293T | C17(1.13); C94(1.03) | LDD1589 | [15] |
| LDCM0386 | CL116 | HEK-293T | C17(1.03); C94(0.90) | LDD1590 | [15] |
| LDCM0387 | CL117 | HEK-293T | C17(0.93) | LDD1591 | [15] |
| LDCM0388 | CL118 | HEK-293T | C17(0.87) | LDD1592 | [15] |
| LDCM0389 | CL119 | HEK-293T | C17(0.89); C94(1.13) | LDD1593 | [15] |
| LDCM0390 | CL12 | HEK-293T | C17(1.01); C94(0.92) | LDD1594 | [15] |
| LDCM0391 | CL120 | HEK-293T | C17(1.02); C94(0.86) | LDD1595 | [15] |
| LDCM0392 | CL121 | PaTu 8988t | C17(1.26) | LDD1271 | [6] |
| LDCM0393 | CL122 | PaTu 8988t | C17(1.45) | LDD1272 | [6] |
| LDCM0394 | CL123 | PaTu 8988t | C17(1.60) | LDD1273 | [6] |
| LDCM0395 | CL124 | PaTu 8988t | C17(1.32) | LDD1274 | [6] |
| LDCM0396 | CL125 | HCT 116 | C17(1.38) | LDD0713 | [6] |
| LDCM0397 | CL126 | HCT 116 | C17(1.34) | LDD0714 | [6] |
| LDCM0398 | CL127 | HCT 116 | C17(1.42) | LDD0715 | [6] |
| LDCM0399 | CL128 | HCT 116 | C17(1.09) | LDD0716 | [6] |
| LDCM0400 | CL13 | HEK-293T | C17(0.94) | LDD1604 | [15] |
| LDCM0401 | CL14 | HEK-293T | C17(0.91) | LDD1605 | [15] |
| LDCM0402 | CL15 | HEK-293T | C17(1.05); C94(1.16) | LDD1606 | [15] |
| LDCM0403 | CL16 | PaTu 8988t | C17(0.93) | LDD1282 | [6] |
| LDCM0404 | CL17 | PaTu 8988t | C17(0.87) | LDD1283 | [6] |
| LDCM0405 | CL18 | PaTu 8988t | C17(0.94) | LDD1284 | [6] |
| LDCM0406 | CL19 | PaTu 8988t | C17(0.80) | LDD1285 | [6] |
| LDCM0407 | CL2 | HEK-293T | C17(0.90) | LDD1611 | [15] |
| LDCM0408 | CL20 | PaTu 8988t | C17(0.84) | LDD1287 | [6] |
| LDCM0409 | CL21 | PaTu 8988t | C17(1.09) | LDD1288 | [6] |
| LDCM0410 | CL22 | PaTu 8988t | C17(1.20) | LDD1289 | [6] |
| LDCM0411 | CL23 | PaTu 8988t | C17(1.04) | LDD1290 | [6] |
| LDCM0412 | CL24 | PaTu 8988t | C17(0.86) | LDD1291 | [6] |
| LDCM0413 | CL25 | PaTu 8988t | C17(0.89) | LDD1292 | [6] |
| LDCM0414 | CL26 | PaTu 8988t | C17(0.97) | LDD1293 | [6] |
| LDCM0415 | CL27 | PaTu 8988t | C17(1.16) | LDD1294 | [6] |
| LDCM0416 | CL28 | PaTu 8988t | C17(1.04) | LDD1295 | [6] |
| LDCM0417 | CL29 | PaTu 8988t | C17(0.91) | LDD1296 | [6] |
| LDCM0418 | CL3 | HEK-293T | C17(1.00); C94(0.97) | LDD1622 | [15] |
| LDCM0419 | CL30 | PaTu 8988t | C17(0.98) | LDD1298 | [6] |
| LDCM0420 | CL31 | HCT 116 | C17(1.01) | LDD0737 | [6] |
| LDCM0421 | CL32 | HCT 116 | C17(0.92) | LDD0738 | [6] |
| LDCM0422 | CL33 | HCT 116 | C17(1.08) | LDD0739 | [6] |
| LDCM0423 | CL34 | HCT 116 | C17(1.09) | LDD0740 | [6] |
| LDCM0424 | CL35 | HCT 116 | C17(1.03) | LDD0741 | [6] |
| LDCM0425 | CL36 | HCT 116 | C17(1.19) | LDD0742 | [6] |
| LDCM0426 | CL37 | HCT 116 | C17(0.85) | LDD0743 | [6] |
| LDCM0428 | CL39 | HCT 116 | C17(0.95) | LDD0745 | [6] |
| LDCM0429 | CL4 | HEK-293T | C17(0.99); C94(1.16) | LDD1633 | [15] |
| LDCM0430 | CL40 | HCT 116 | C17(1.12) | LDD0747 | [6] |
| LDCM0431 | CL41 | HCT 116 | C17(0.84) | LDD0748 | [6] |
| LDCM0432 | CL42 | HCT 116 | C17(0.99) | LDD0749 | [6] |
| LDCM0433 | CL43 | HCT 116 | C17(1.15) | LDD0750 | [6] |
| LDCM0434 | CL44 | HCT 116 | C17(0.91) | LDD0751 | [6] |
| LDCM0435 | CL45 | HCT 116 | C17(0.94) | LDD0752 | [6] |
| LDCM0436 | CL46 | HEK-293T | C17(1.00); C91(1.31) | LDD1640 | [15] |
| LDCM0437 | CL47 | HEK-293T | C17(1.09); C157(0.44) | LDD1641 | [15] |
| LDCM0438 | CL48 | HEK-293T | C17(1.12); C94(1.00) | LDD1642 | [15] |
| LDCM0439 | CL49 | HEK-293T | C17(0.86) | LDD1643 | [15] |
| LDCM0440 | CL5 | HEK-293T | C17(1.07) | LDD1644 | [15] |
| LDCM0441 | CL50 | HEK-293T | C17(0.87) | LDD1645 | [15] |
| LDCM0443 | CL52 | HEK-293T | C17(1.20); C94(1.05) | LDD1646 | [15] |
| LDCM0444 | CL53 | HEK-293T | C17(0.92) | LDD1647 | [15] |
| LDCM0445 | CL54 | HEK-293T | C17(0.92) | LDD1648 | [15] |
| LDCM0446 | CL55 | HEK-293T | C17(1.03) | LDD1649 | [15] |
| LDCM0447 | CL56 | HEK-293T | C17(0.93); C157(0.46) | LDD1650 | [15] |
| LDCM0448 | CL57 | HEK-293T | C17(1.00) | LDD1651 | [15] |
| LDCM0449 | CL58 | HEK-293T | C17(0.99); C91(1.44) | LDD1652 | [15] |
| LDCM0450 | CL59 | HEK-293T | C17(1.11); C157(0.46) | LDD1653 | [15] |
| LDCM0451 | CL6 | HEK-293T | C17(1.01) | LDD1654 | [15] |
| LDCM0452 | CL60 | HEK-293T | C17(0.97); C94(1.14) | LDD1655 | [15] |
| LDCM0453 | CL61 | HEK-293T | C17(0.89) | LDD1656 | [15] |
| LDCM0454 | CL62 | HEK-293T | C17(0.92) | LDD1657 | [15] |
| LDCM0455 | CL63 | HEK-293T | C17(0.97); C94(0.77) | LDD1658 | [15] |
| LDCM0456 | CL64 | HEK-293T | C17(0.91); C94(1.09) | LDD1659 | [15] |
| LDCM0457 | CL65 | HEK-293T | C17(0.94) | LDD1660 | [15] |
| LDCM0458 | CL66 | HEK-293T | C17(1.07) | LDD1661 | [15] |
| LDCM0459 | CL67 | HEK-293T | C17(1.00) | LDD1662 | [15] |
| LDCM0460 | CL68 | HEK-293T | C17(0.95); C157(0.58) | LDD1663 | [15] |
| LDCM0461 | CL69 | HEK-293T | C17(1.14) | LDD1664 | [15] |
| LDCM0462 | CL7 | HEK-293T | C17(1.07) | LDD1665 | [15] |
| LDCM0463 | CL70 | HEK-293T | C17(1.04); C91(1.28) | LDD1666 | [15] |
| LDCM0464 | CL71 | HEK-293T | C17(1.10); C157(0.46) | LDD1667 | [15] |
| LDCM0465 | CL72 | HEK-293T | C17(1.15); C94(1.15) | LDD1668 | [15] |
| LDCM0466 | CL73 | HEK-293T | C17(0.88) | LDD1669 | [15] |
| LDCM0467 | CL74 | HEK-293T | C17(0.94) | LDD1670 | [15] |
| LDCM0469 | CL76 | HCT 116 | C17(1.00) | LDD0786 | [6] |
| LDCM0470 | CL77 | HCT 116 | C17(0.83) | LDD0787 | [6] |
| LDCM0471 | CL78 | HCT 116 | C17(0.93) | LDD0788 | [6] |
| LDCM0472 | CL79 | HCT 116 | C17(0.87) | LDD0789 | [6] |
| LDCM0473 | CL8 | HEK-293T | C17(0.80); C157(0.55) | LDD1676 | [15] |
| LDCM0474 | CL80 | HCT 116 | C17(0.71) | LDD0791 | [6] |
| LDCM0475 | CL81 | HCT 116 | C17(0.95) | LDD0792 | [6] |
| LDCM0476 | CL82 | HCT 116 | C17(0.90) | LDD0793 | [6] |
| LDCM0477 | CL83 | HCT 116 | C17(0.84) | LDD0794 | [6] |
| LDCM0478 | CL84 | HCT 116 | C17(0.84) | LDD0795 | [6] |
| LDCM0479 | CL85 | HCT 116 | C17(0.84) | LDD0796 | [6] |
| LDCM0480 | CL86 | HCT 116 | C17(0.91) | LDD0797 | [6] |
| LDCM0481 | CL87 | HCT 116 | C17(1.11) | LDD0798 | [6] |
| LDCM0482 | CL88 | HCT 116 | C17(0.95) | LDD0799 | [6] |
| LDCM0483 | CL89 | HCT 116 | C17(0.83) | LDD0800 | [6] |
| LDCM0484 | CL9 | HEK-293T | C17(1.18) | LDD1687 | [15] |
| LDCM0485 | CL90 | HCT 116 | C17(1.30) | LDD0802 | [6] |
| LDCM0486 | CL91 | HEK-293T | C17(0.92) | LDD1689 | [15] |
| LDCM0487 | CL92 | HEK-293T | C17(0.96); C157(0.78) | LDD1690 | [15] |
| LDCM0488 | CL93 | HEK-293T | C17(1.04) | LDD1691 | [15] |
| LDCM0489 | CL94 | HEK-293T | C17(1.03); C91(1.26) | LDD1692 | [15] |
| LDCM0490 | CL95 | HEK-293T | C17(0.85); C157(0.67) | LDD1693 | [15] |
| LDCM0491 | CL96 | HEK-293T | C17(0.95); C94(1.02) | LDD1694 | [15] |
| LDCM0492 | CL97 | HEK-293T | C17(0.85) | LDD1695 | [15] |
| LDCM0493 | CL98 | HEK-293T | C17(0.94) | LDD1696 | [15] |
| LDCM0494 | CL99 | HEK-293T | C17(0.96); C94(1.01) | LDD1697 | [15] |
| LDCM0495 | E2913 | HEK-293T | C17(0.89); C94(0.98) | LDD1698 | [15] |
| LDCM0468 | Fragment33 | HEK-293T | C17(1.02); C94(1.06) | LDD1671 | [15] |
| LDCM0427 | Fragment51 | HCT 116 | C17(1.41) | LDD0744 | [6] |
| LDCM0022 | KB02 | HCT 116 | C17(1.65) | LDD0080 | [6] |
| LDCM0023 | KB03 | HCT 116 | C17(1.27) | LDD0081 | [6] |
| LDCM0024 | KB05 | HCT 116 | C17(1.47) | LDD0082 | [6] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C17(0.83) | LDD2141 | [2] |
| LDCM0131 | RA190 | MM1.R | C17(1.21) | LDD0304 | [16] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Huntingtin (HTT) | Huntingtin family | P42858 | |||
| Myelin and lymphocyte protein (MAL) | MAL family | P21145 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Ataxin-1 (ATXN1) | ATXN1 family | P54253 | |||
References
