Details of the Target
General Information of Target
| Target ID | LDTP08389 | |||||
|---|---|---|---|---|---|---|
| Target Name | Syntaxin-12 (STX12) | |||||
| Gene Name | STX12 | |||||
| Gene ID | 23673 | |||||
| Synonyms |
Syntaxin-12 |
|||||
| 3D Structure | ||||||
| Sequence |
MSYGPLDMYRNPGPSGPQLRDFSSIIQTCSGNIQRISQATAQIKNLMSQLGTKQDSSKLQ
ENLQQLQHSTNQLAKETNELLKELGSLPLPLSTSEQRQQRLQKERLMNDFSAALNNFQAV QRRVSEKEKESIARARAGSRLSAEERQREEQLVSFDSHEEWNQMQSQEDEVAITEQDLEL IKERETAIRQLEADILDVNQIFKDLAMMIHDQGDLIDSIEANVESSEVHVERATEQLQRA AYYQKKSRKKMCILVLVLSVIILILGLIIWLVYKTK |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Syntaxin family
|
|||||
| Subcellular location |
Endosome membrane
|
|||||
| Function |
SNARE promoting fusion of transport vesicles with target membranes. Together with SNARE STX6, promotes movement of vesicles from endosomes to the cell membrane, and may therefore function in the endocytic recycling pathway. Through complex formation with GRIP1, GRIA2 and NSG1 controls the intracellular fate of AMPAR and the endosomal sorting of the GRIA2 subunit toward recycling and membrane targeting.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FBPP2 Probe Info |
 |
3.13 | LDD0318 | [1] | |
|
FBP2 Probe Info |
 |
2.18 | LDD0317 | [1] | |
|
Probe 1 Probe Info |
 |
Y242(56.67); Y243(111.25) | LDD3495 | [2] | |
|
BTD Probe Info |
 |
C29(0.44) | LDD2112 | [3] | |
|
D5yne Probe Info |
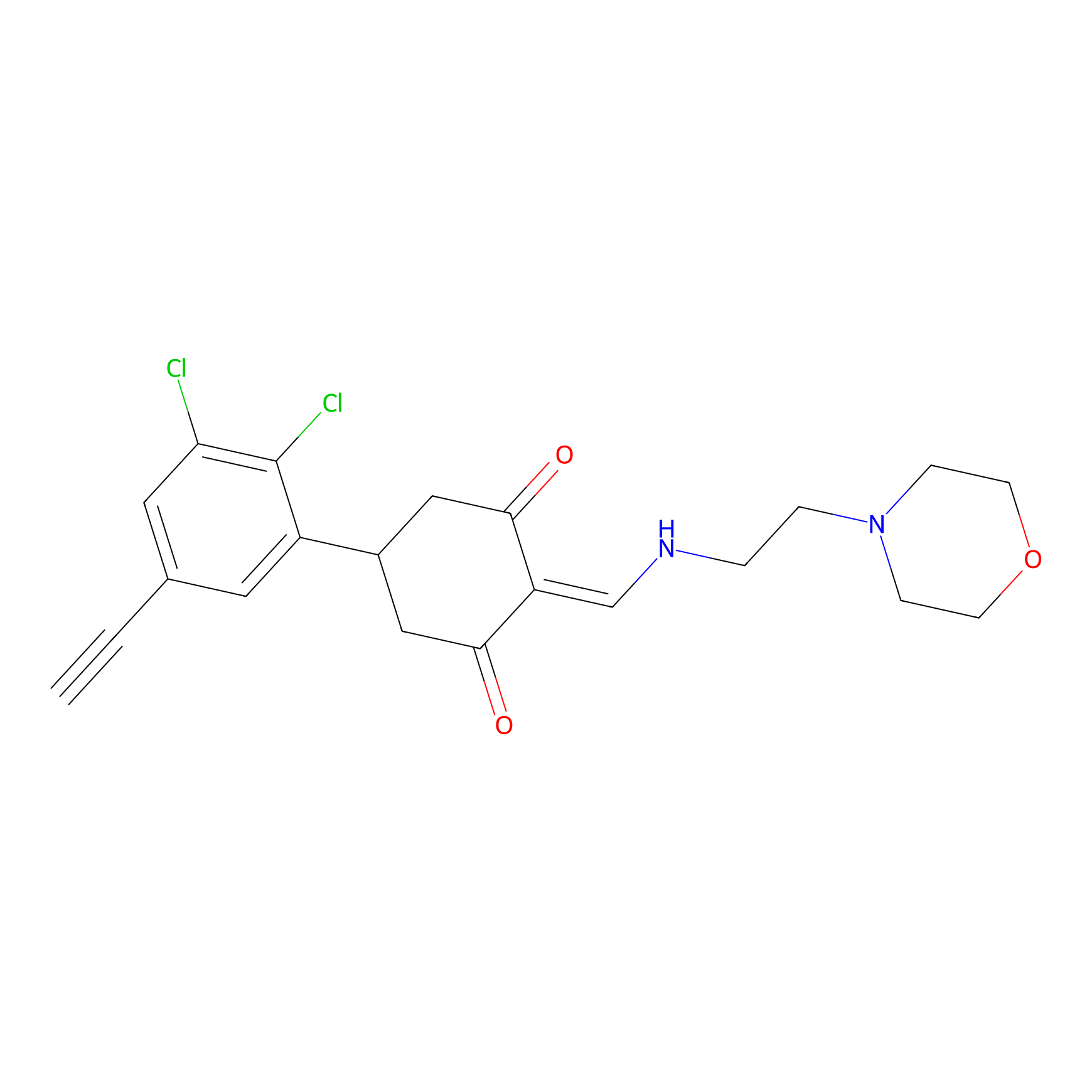 |
2.26 | LDD0055 | [4] | |
|
DBIA Probe Info |
 |
C29(1.02) | LDD1492 | [5] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [6] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [7] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0165 | [8] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [7] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [9] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [9] | |
|
1c-yne Probe Info |
 |
K44(0.00); K82(0.00) | LDD0228 | [10] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [11] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [11] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C170 Probe Info |
 |
9.06 | LDD1850 | [12] | |
|
C231 Probe Info |
 |
12.04 | LDD1904 | [12] | |
|
C310 Probe Info |
 |
11.55 | LDD1977 | [12] | |
|
C362 Probe Info |
 |
26.54 | LDD2023 | [12] | |
|
FFF probe11 Probe Info |
 |
18.79 | LDD0471 | [13] | |
|
FFF probe12 Probe Info |
 |
13.85 | LDD0473 | [13] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [13] | |
|
FFF probe14 Probe Info |
 |
20.00 | LDD0477 | [13] | |
|
FFF probe2 Probe Info |
 |
15.69 | LDD0463 | [13] | |
|
FFF probe3 Probe Info |
 |
17.80 | LDD0464 | [13] | |
|
JN0003 Probe Info |
 |
20.00 | LDD0469 | [13] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C29(0.34) | LDD2142 | [3] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C29(0.44) | LDD2112 | [3] |
| LDCM0214 | AC1 | HEK-293T | C29(0.90) | LDD1507 | [5] |
| LDCM0215 | AC10 | HEK-293T | C29(0.87) | LDD1508 | [5] |
| LDCM0226 | AC11 | HEK-293T | C29(0.82) | LDD1509 | [5] |
| LDCM0237 | AC12 | HEK-293T | C29(0.96) | LDD1510 | [5] |
| LDCM0259 | AC14 | HEK-293T | C29(0.91) | LDD1512 | [5] |
| LDCM0270 | AC15 | HEK-293T | C29(0.91) | LDD1513 | [5] |
| LDCM0276 | AC17 | HEK-293T | C29(0.92) | LDD1515 | [5] |
| LDCM0277 | AC18 | HEK-293T | C29(0.89) | LDD1516 | [5] |
| LDCM0278 | AC19 | HEK-293T | C29(0.81) | LDD1517 | [5] |
| LDCM0279 | AC2 | HEK-293T | C29(0.98) | LDD1518 | [5] |
| LDCM0280 | AC20 | HEK-293T | C29(1.02) | LDD1519 | [5] |
| LDCM0281 | AC21 | HEK-293T | C29(1.11) | LDD1520 | [5] |
| LDCM0282 | AC22 | HEK-293T | C29(0.95) | LDD1521 | [5] |
| LDCM0283 | AC23 | HEK-293T | C29(0.85) | LDD1522 | [5] |
| LDCM0284 | AC24 | HEK-293T | C29(0.89) | LDD1523 | [5] |
| LDCM0285 | AC25 | HEK-293T | C29(0.96) | LDD1524 | [5] |
| LDCM0286 | AC26 | HEK-293T | C29(0.82) | LDD1525 | [5] |
| LDCM0287 | AC27 | HEK-293T | C29(0.84) | LDD1526 | [5] |
| LDCM0288 | AC28 | HEK-293T | C29(0.84) | LDD1527 | [5] |
| LDCM0289 | AC29 | HEK-293T | C29(0.92) | LDD1528 | [5] |
| LDCM0290 | AC3 | HEK-293T | C29(0.82) | LDD1529 | [5] |
| LDCM0291 | AC30 | HEK-293T | C29(0.91) | LDD1530 | [5] |
| LDCM0292 | AC31 | HEK-293T | C29(0.96) | LDD1531 | [5] |
| LDCM0293 | AC32 | HEK-293T | C29(0.87) | LDD1532 | [5] |
| LDCM0294 | AC33 | HEK-293T | C29(0.98) | LDD1533 | [5] |
| LDCM0295 | AC34 | HEK-293T | C29(0.86) | LDD1534 | [5] |
| LDCM0296 | AC35 | HEK-293T | C29(0.94) | LDD1535 | [5] |
| LDCM0297 | AC36 | HEK-293T | C29(0.99) | LDD1536 | [5] |
| LDCM0298 | AC37 | HEK-293T | C29(0.97) | LDD1537 | [5] |
| LDCM0299 | AC38 | HEK-293T | C29(0.96) | LDD1538 | [5] |
| LDCM0300 | AC39 | HEK-293T | C29(0.93) | LDD1539 | [5] |
| LDCM0301 | AC4 | HEK-293T | C29(0.86) | LDD1540 | [5] |
| LDCM0302 | AC40 | HEK-293T | C29(1.06) | LDD1541 | [5] |
| LDCM0303 | AC41 | HEK-293T | C29(0.89) | LDD1542 | [5] |
| LDCM0304 | AC42 | HEK-293T | C29(0.82) | LDD1543 | [5] |
| LDCM0305 | AC43 | HEK-293T | C29(0.93) | LDD1544 | [5] |
| LDCM0306 | AC44 | HEK-293T | C29(0.97) | LDD1545 | [5] |
| LDCM0307 | AC45 | HEK-293T | C29(1.03) | LDD1546 | [5] |
| LDCM0308 | AC46 | HEK-293T | C29(0.98) | LDD1547 | [5] |
| LDCM0309 | AC47 | HEK-293T | C29(0.91) | LDD1548 | [5] |
| LDCM0310 | AC48 | HEK-293T | C29(0.98) | LDD1549 | [5] |
| LDCM0311 | AC49 | HEK-293T | C29(0.85) | LDD1550 | [5] |
| LDCM0312 | AC5 | HEK-293T | C29(1.03) | LDD1551 | [5] |
| LDCM0313 | AC50 | HEK-293T | C29(0.87) | LDD1552 | [5] |
| LDCM0314 | AC51 | HEK-293T | C29(0.92) | LDD1553 | [5] |
| LDCM0315 | AC52 | HEK-293T | C29(1.01) | LDD1554 | [5] |
| LDCM0316 | AC53 | HEK-293T | C29(1.02) | LDD1555 | [5] |
| LDCM0317 | AC54 | HEK-293T | C29(1.04) | LDD1556 | [5] |
| LDCM0318 | AC55 | HEK-293T | C29(0.93) | LDD1557 | [5] |
| LDCM0319 | AC56 | HEK-293T | C29(0.99) | LDD1558 | [5] |
| LDCM0320 | AC57 | HEK-293T | C29(0.83) | LDD1559 | [5] |
| LDCM0321 | AC58 | HEK-293T | C29(0.83) | LDD1560 | [5] |
| LDCM0322 | AC59 | HEK-293T | C29(0.86) | LDD1561 | [5] |
| LDCM0323 | AC6 | HEK-293T | C29(0.82) | LDD1562 | [5] |
| LDCM0324 | AC60 | HEK-293T | C29(0.88) | LDD1563 | [5] |
| LDCM0325 | AC61 | HEK-293T | C29(0.86) | LDD1564 | [5] |
| LDCM0326 | AC62 | HEK-293T | C29(0.89) | LDD1565 | [5] |
| LDCM0327 | AC63 | HEK-293T | C29(0.80) | LDD1566 | [5] |
| LDCM0328 | AC64 | HEK-293T | C29(0.78) | LDD1567 | [5] |
| LDCM0334 | AC7 | HEK-293T | C29(0.84) | LDD1568 | [5] |
| LDCM0345 | AC8 | HEK-293T | C29(0.85) | LDD1569 | [5] |
| LDCM0248 | AKOS034007472 | HEK-293T | C29(1.09) | LDD1511 | [5] |
| LDCM0356 | AKOS034007680 | HEK-293T | C29(0.88) | LDD1570 | [5] |
| LDCM0275 | AKOS034007705 | HEK-293T | C29(0.93) | LDD1514 | [5] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [11] |
| LDCM0367 | CL1 | HEK-293T | C29(0.94) | LDD1571 | [5] |
| LDCM0368 | CL10 | HEK-293T | C29(0.76) | LDD1572 | [5] |
| LDCM0369 | CL100 | HEK-293T | C29(0.87) | LDD1573 | [5] |
| LDCM0370 | CL101 | HEK-293T | C29(0.95) | LDD1574 | [5] |
| LDCM0371 | CL102 | HEK-293T | C29(0.80) | LDD1575 | [5] |
| LDCM0372 | CL103 | HEK-293T | C29(1.00) | LDD1576 | [5] |
| LDCM0373 | CL104 | HEK-293T | C29(0.86) | LDD1577 | [5] |
| LDCM0374 | CL105 | HEK-293T | C29(0.90) | LDD1578 | [5] |
| LDCM0375 | CL106 | HEK-293T | C29(0.89) | LDD1579 | [5] |
| LDCM0376 | CL107 | HEK-293T | C29(0.93) | LDD1580 | [5] |
| LDCM0377 | CL108 | HEK-293T | C29(0.95) | LDD1581 | [5] |
| LDCM0378 | CL109 | HEK-293T | C29(0.85) | LDD1582 | [5] |
| LDCM0379 | CL11 | HEK-293T | C29(0.87) | LDD1583 | [5] |
| LDCM0380 | CL110 | HEK-293T | C29(0.80) | LDD1584 | [5] |
| LDCM0381 | CL111 | HEK-293T | C29(0.80) | LDD1585 | [5] |
| LDCM0382 | CL112 | HEK-293T | C29(0.97) | LDD1586 | [5] |
| LDCM0383 | CL113 | HEK-293T | C29(0.94) | LDD1587 | [5] |
| LDCM0384 | CL114 | HEK-293T | C29(0.81) | LDD1588 | [5] |
| LDCM0385 | CL115 | HEK-293T | C29(0.89) | LDD1589 | [5] |
| LDCM0386 | CL116 | HEK-293T | C29(0.99) | LDD1590 | [5] |
| LDCM0387 | CL117 | HEK-293T | C29(0.96) | LDD1591 | [5] |
| LDCM0388 | CL118 | HEK-293T | C29(0.85) | LDD1592 | [5] |
| LDCM0389 | CL119 | HEK-293T | C29(0.90) | LDD1593 | [5] |
| LDCM0390 | CL12 | HEK-293T | C29(1.21) | LDD1594 | [5] |
| LDCM0391 | CL120 | HEK-293T | C29(0.95) | LDD1595 | [5] |
| LDCM0392 | CL121 | HEK-293T | C29(0.97) | LDD1596 | [5] |
| LDCM0393 | CL122 | HEK-293T | C29(0.89) | LDD1597 | [5] |
| LDCM0394 | CL123 | HEK-293T | C29(0.86) | LDD1598 | [5] |
| LDCM0395 | CL124 | HEK-293T | C29(0.89) | LDD1599 | [5] |
| LDCM0396 | CL125 | HEK-293T | C29(0.85) | LDD1600 | [5] |
| LDCM0397 | CL126 | HEK-293T | C29(0.92) | LDD1601 | [5] |
| LDCM0398 | CL127 | HEK-293T | C29(0.89) | LDD1602 | [5] |
| LDCM0399 | CL128 | HEK-293T | C29(0.89) | LDD1603 | [5] |
| LDCM0400 | CL13 | HEK-293T | C29(1.02) | LDD1604 | [5] |
| LDCM0401 | CL14 | HEK-293T | C29(0.86) | LDD1605 | [5] |
| LDCM0402 | CL15 | HEK-293T | C29(0.91) | LDD1606 | [5] |
| LDCM0403 | CL16 | HEK-293T | C29(0.89) | LDD1607 | [5] |
| LDCM0404 | CL17 | HEK-293T | C29(0.83) | LDD1608 | [5] |
| LDCM0405 | CL18 | HEK-293T | C29(0.93) | LDD1609 | [5] |
| LDCM0406 | CL19 | HEK-293T | C29(0.99) | LDD1610 | [5] |
| LDCM0407 | CL2 | HEK-293T | C29(0.99) | LDD1611 | [5] |
| LDCM0408 | CL20 | HEK-293T | C29(1.01) | LDD1612 | [5] |
| LDCM0409 | CL21 | HEK-293T | C29(0.94) | LDD1613 | [5] |
| LDCM0410 | CL22 | HEK-293T | C29(1.26) | LDD1614 | [5] |
| LDCM0411 | CL23 | HEK-293T | C29(0.94) | LDD1615 | [5] |
| LDCM0412 | CL24 | HEK-293T | C29(1.13) | LDD1616 | [5] |
| LDCM0413 | CL25 | HEK-293T | C29(0.89) | LDD1617 | [5] |
| LDCM0414 | CL26 | HEK-293T | C29(1.13) | LDD1618 | [5] |
| LDCM0415 | CL27 | HEK-293T | C29(0.91) | LDD1619 | [5] |
| LDCM0416 | CL28 | HEK-293T | C29(0.90) | LDD1620 | [5] |
| LDCM0417 | CL29 | HEK-293T | C29(0.97) | LDD1621 | [5] |
| LDCM0418 | CL3 | HEK-293T | C29(0.86) | LDD1622 | [5] |
| LDCM0419 | CL30 | HEK-293T | C29(1.00) | LDD1623 | [5] |
| LDCM0420 | CL31 | HEK-293T | C29(0.93) | LDD1624 | [5] |
| LDCM0421 | CL32 | HEK-293T | C29(0.97) | LDD1625 | [5] |
| LDCM0422 | CL33 | HEK-293T | C29(0.88) | LDD1626 | [5] |
| LDCM0423 | CL34 | HEK-293T | C29(1.00) | LDD1627 | [5] |
| LDCM0424 | CL35 | HEK-293T | C29(0.81) | LDD1628 | [5] |
| LDCM0425 | CL36 | HEK-293T | C29(0.88) | LDD1629 | [5] |
| LDCM0426 | CL37 | HEK-293T | C29(0.99) | LDD1630 | [5] |
| LDCM0428 | CL39 | HEK-293T | C29(0.90) | LDD1632 | [5] |
| LDCM0429 | CL4 | HEK-293T | C29(0.93) | LDD1633 | [5] |
| LDCM0430 | CL40 | HEK-293T | C29(0.99) | LDD1634 | [5] |
| LDCM0431 | CL41 | HEK-293T | C29(0.83) | LDD1635 | [5] |
| LDCM0432 | CL42 | HEK-293T | C29(1.01) | LDD1636 | [5] |
| LDCM0433 | CL43 | HEK-293T | C29(0.95) | LDD1637 | [5] |
| LDCM0434 | CL44 | HEK-293T | C29(0.95) | LDD1638 | [5] |
| LDCM0435 | CL45 | HEK-293T | C29(1.00) | LDD1639 | [5] |
| LDCM0436 | CL46 | HEK-293T | C29(0.86) | LDD1640 | [5] |
| LDCM0437 | CL47 | HEK-293T | C29(0.96) | LDD1641 | [5] |
| LDCM0438 | CL48 | HEK-293T | C29(0.99) | LDD1642 | [5] |
| LDCM0439 | CL49 | HEK-293T | C29(0.97) | LDD1643 | [5] |
| LDCM0440 | CL5 | HEK-293T | C29(0.92) | LDD1644 | [5] |
| LDCM0441 | CL50 | HEK-293T | C29(1.02) | LDD1645 | [5] |
| LDCM0443 | CL52 | HEK-293T | C29(0.92) | LDD1646 | [5] |
| LDCM0444 | CL53 | HEK-293T | C29(0.92) | LDD1647 | [5] |
| LDCM0445 | CL54 | HEK-293T | C29(0.85) | LDD1648 | [5] |
| LDCM0446 | CL55 | HEK-293T | C29(0.91) | LDD1649 | [5] |
| LDCM0447 | CL56 | HEK-293T | C29(0.95) | LDD1650 | [5] |
| LDCM0448 | CL57 | HEK-293T | C29(0.89) | LDD1651 | [5] |
| LDCM0449 | CL58 | HEK-293T | C29(0.91) | LDD1652 | [5] |
| LDCM0450 | CL59 | HEK-293T | C29(0.97) | LDD1653 | [5] |
| LDCM0451 | CL6 | HEK-293T | C29(0.89) | LDD1654 | [5] |
| LDCM0452 | CL60 | HEK-293T | C29(1.17) | LDD1655 | [5] |
| LDCM0453 | CL61 | HEK-293T | C29(0.98) | LDD1656 | [5] |
| LDCM0454 | CL62 | HEK-293T | C29(0.98) | LDD1657 | [5] |
| LDCM0455 | CL63 | HEK-293T | C29(0.94) | LDD1658 | [5] |
| LDCM0456 | CL64 | HEK-293T | C29(0.88) | LDD1659 | [5] |
| LDCM0457 | CL65 | HEK-293T | C29(0.88) | LDD1660 | [5] |
| LDCM0458 | CL66 | HEK-293T | C29(0.99) | LDD1661 | [5] |
| LDCM0459 | CL67 | HEK-293T | C29(0.91) | LDD1662 | [5] |
| LDCM0460 | CL68 | HEK-293T | C29(0.98) | LDD1663 | [5] |
| LDCM0461 | CL69 | HEK-293T | C29(1.02) | LDD1664 | [5] |
| LDCM0462 | CL7 | HEK-293T | C29(0.94) | LDD1665 | [5] |
| LDCM0463 | CL70 | HEK-293T | C29(0.90) | LDD1666 | [5] |
| LDCM0464 | CL71 | HEK-293T | C29(0.87) | LDD1667 | [5] |
| LDCM0465 | CL72 | HEK-293T | C29(1.11) | LDD1668 | [5] |
| LDCM0466 | CL73 | HEK-293T | C29(0.90) | LDD1669 | [5] |
| LDCM0467 | CL74 | HEK-293T | C29(0.90) | LDD1670 | [5] |
| LDCM0469 | CL76 | HEK-293T | C29(0.95) | LDD1672 | [5] |
| LDCM0470 | CL77 | HEK-293T | C29(0.71) | LDD1673 | [5] |
| LDCM0471 | CL78 | HEK-293T | C29(0.88) | LDD1674 | [5] |
| LDCM0472 | CL79 | HEK-293T | C29(1.15) | LDD1675 | [5] |
| LDCM0473 | CL8 | HEK-293T | C29(0.54) | LDD1676 | [5] |
| LDCM0474 | CL80 | HEK-293T | C29(1.00) | LDD1677 | [5] |
| LDCM0475 | CL81 | HEK-293T | C29(1.20) | LDD1678 | [5] |
| LDCM0476 | CL82 | HEK-293T | C29(0.96) | LDD1679 | [5] |
| LDCM0477 | CL83 | HEK-293T | C29(0.90) | LDD1680 | [5] |
| LDCM0478 | CL84 | HEK-293T | C29(0.84) | LDD1681 | [5] |
| LDCM0479 | CL85 | HEK-293T | C29(1.00) | LDD1682 | [5] |
| LDCM0480 | CL86 | HEK-293T | C29(0.85) | LDD1683 | [5] |
| LDCM0481 | CL87 | HEK-293T | C29(0.82) | LDD1684 | [5] |
| LDCM0482 | CL88 | HEK-293T | C29(0.83) | LDD1685 | [5] |
| LDCM0483 | CL89 | HEK-293T | C29(0.89) | LDD1686 | [5] |
| LDCM0484 | CL9 | HEK-293T | C29(0.97) | LDD1687 | [5] |
| LDCM0485 | CL90 | HEK-293T | C29(0.83) | LDD1688 | [5] |
| LDCM0486 | CL91 | HEK-293T | C29(0.93) | LDD1689 | [5] |
| LDCM0487 | CL92 | HEK-293T | C29(0.95) | LDD1690 | [5] |
| LDCM0488 | CL93 | HEK-293T | C29(1.02) | LDD1691 | [5] |
| LDCM0489 | CL94 | HEK-293T | C29(0.96) | LDD1692 | [5] |
| LDCM0490 | CL95 | HEK-293T | C29(0.84) | LDD1693 | [5] |
| LDCM0491 | CL96 | HEK-293T | C29(0.94) | LDD1694 | [5] |
| LDCM0492 | CL97 | HEK-293T | C29(0.84) | LDD1695 | [5] |
| LDCM0493 | CL98 | HEK-293T | C29(0.84) | LDD1696 | [5] |
| LDCM0494 | CL99 | HEK-293T | C29(0.91) | LDD1697 | [5] |
| LDCM0095 | DC-LC3IN-D5 | HeLa | 2.26 | LDD0055 | [4] |
| LDCM0495 | E2913 | HEK-293T | C29(1.01) | LDD1698 | [5] |
| LDCM0468 | Fragment33 | HEK-293T | C29(0.94) | LDD1671 | [5] |
| LDCM0427 | Fragment51 | HEK-293T | C29(0.98) | LDD1631 | [5] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [11] |
| LDCM0022 | KB02 | HEK-293T | C29(1.02) | LDD1492 | [5] |
| LDCM0023 | KB03 | HEK-293T | C29(0.98) | LDD1497 | [5] |
| LDCM0024 | KB05 | HEK-293T | C29(0.97) | LDD1502 | [5] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0224 | [11] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| B-cell antigen receptor complex-associated protein alpha chain (CD79A) | . | P11912 | |||
Other
References
