Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
OPA-S-S-alkyne Probe Info |
 |
K381(1.61); K662(1.83); K180(1.97); K281(1.99) | LDD3494 | [1] | |
|
IMP-1710 Probe Info |
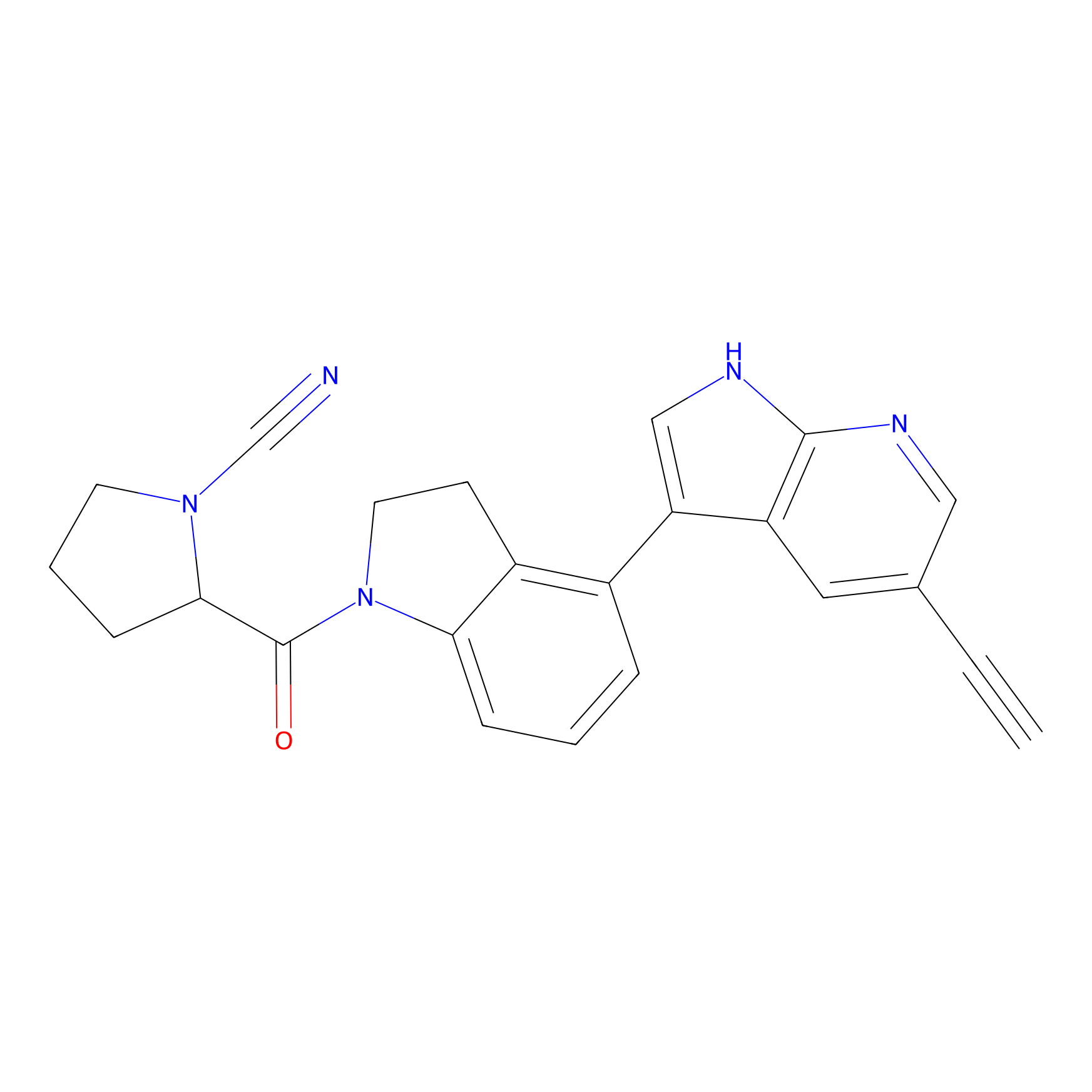 |
2.41 | LDD0040 | [2] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0165 | [3] | |
|
IPM Probe Info |
 |
C814(0.00); C677(0.00); C792(0.00) | LDD2156 | [4] | |
|
AOyne Probe Info |
 |
14.10 | LDD0443 | [5] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C073 Probe Info |
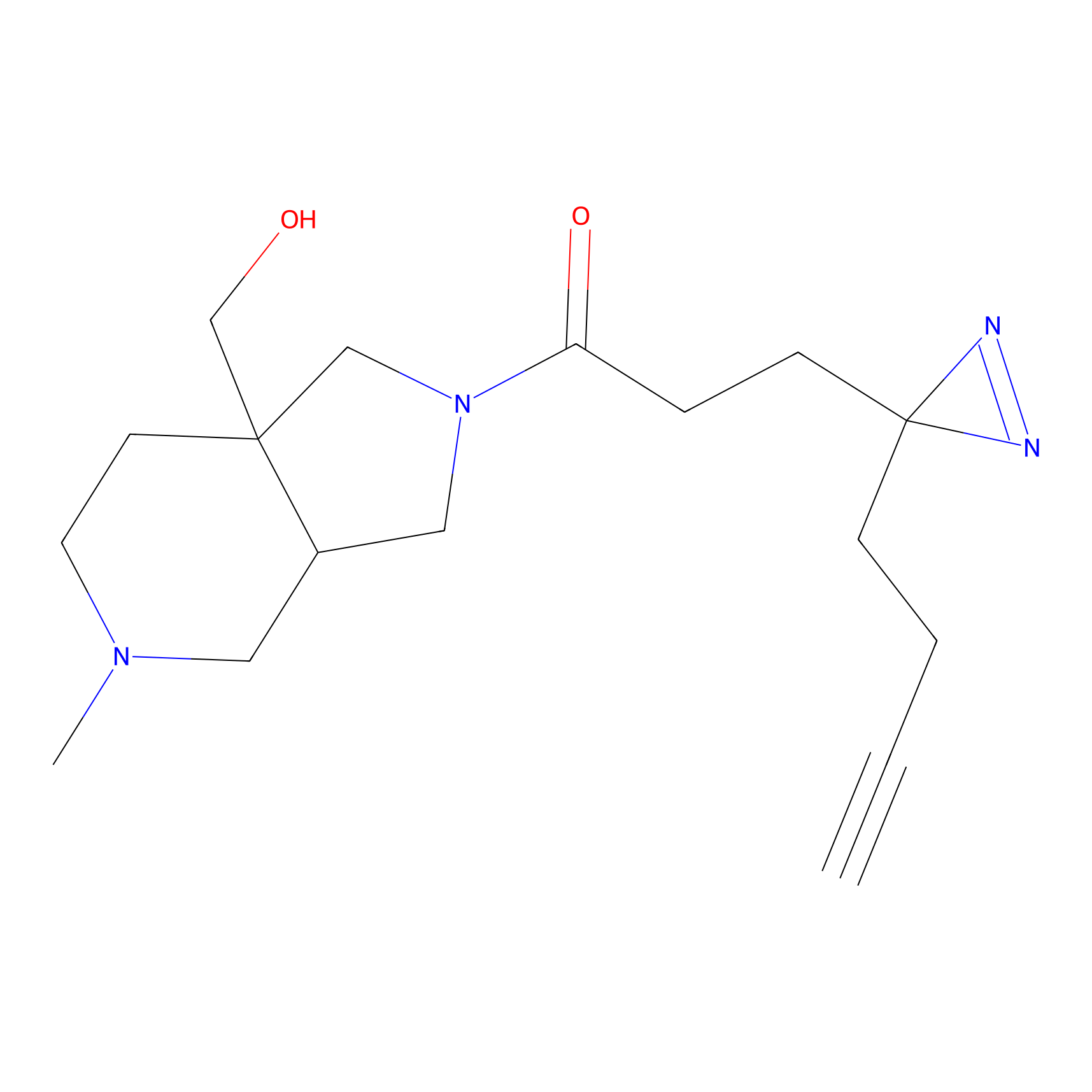 |
5.03 | LDD1769 | [6] | |
|
C115 Probe Info |
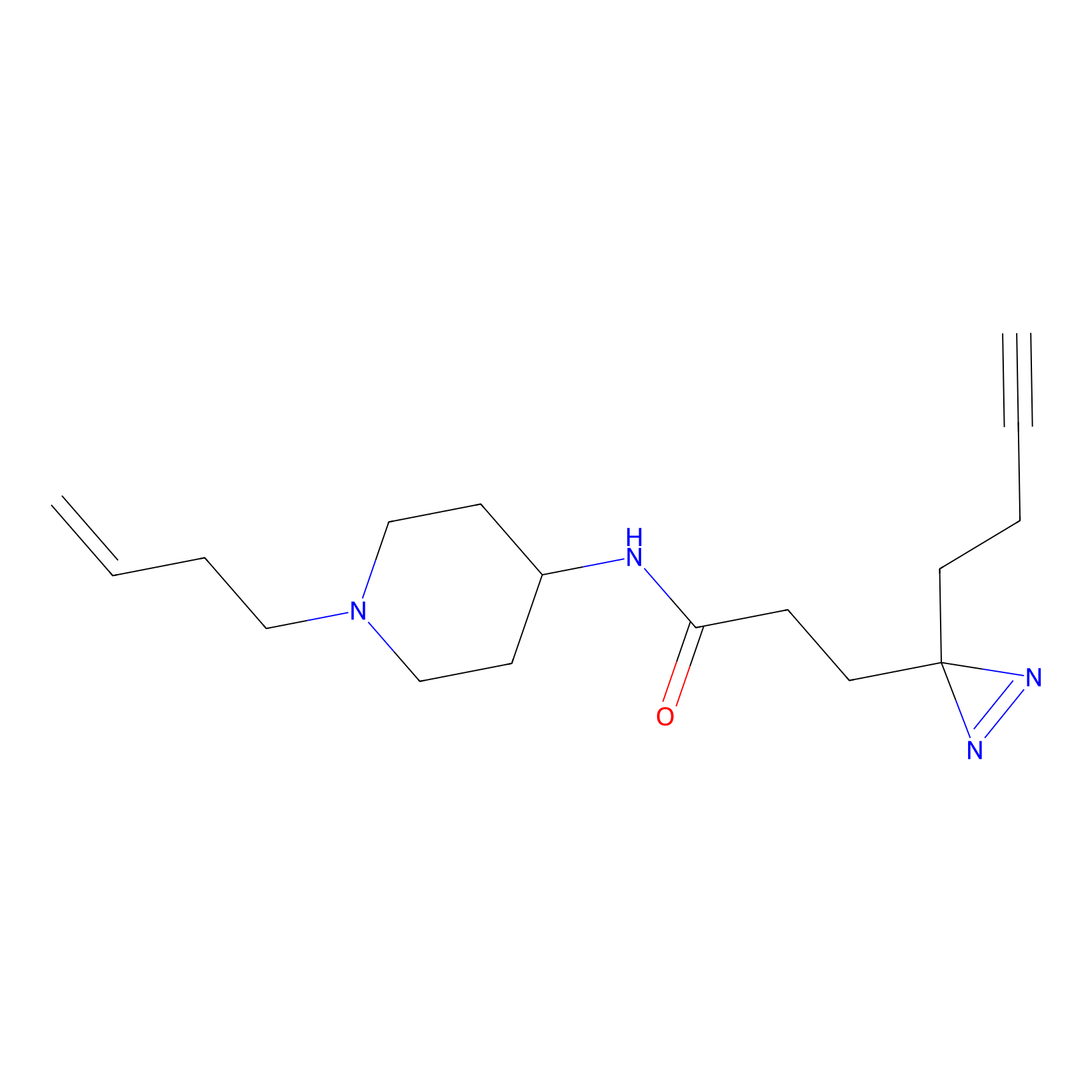 |
4.99 | LDD1802 | [6] | |
|
C149 Probe Info |
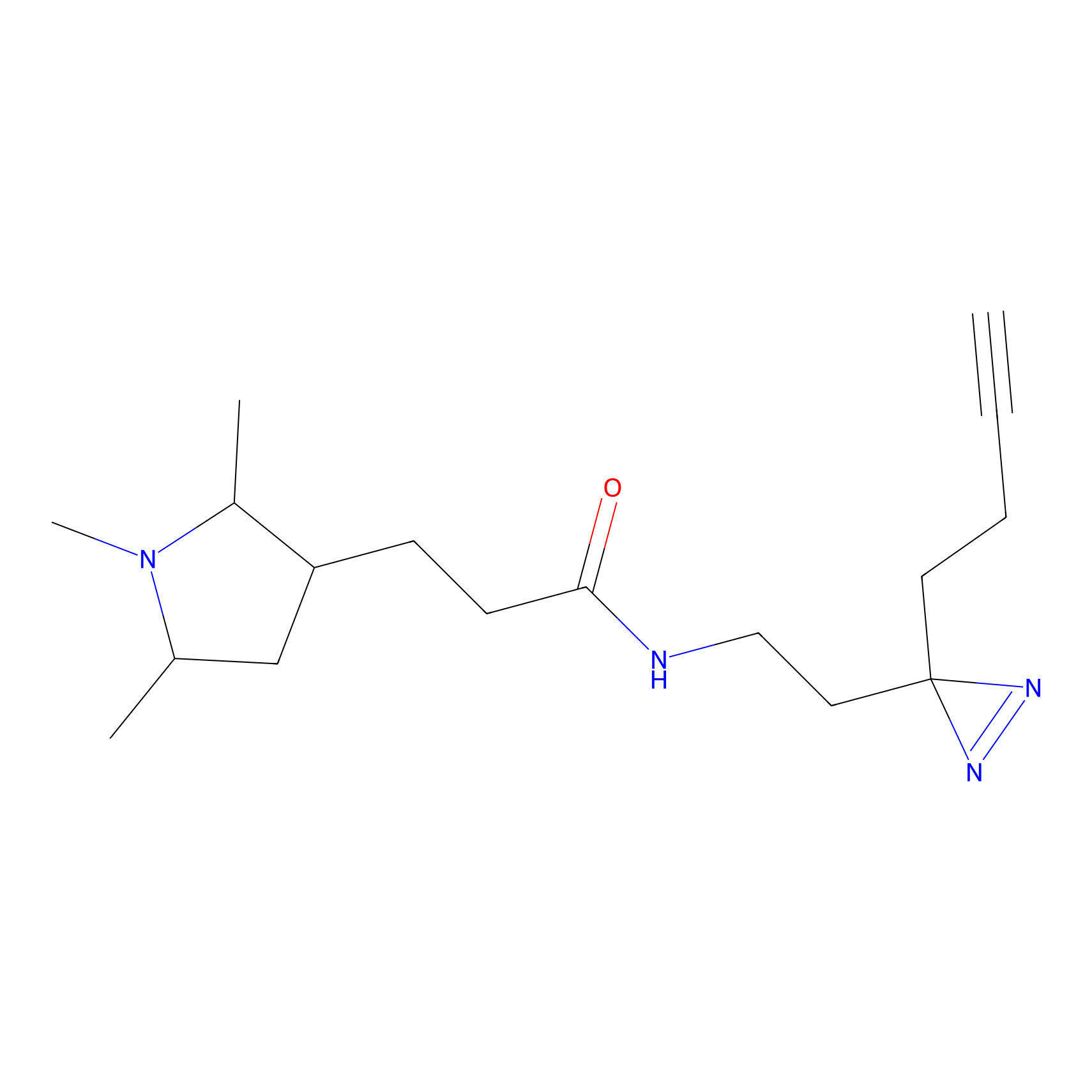 |
7.06 | LDD1830 | [6] | |
|
C159 Probe Info |
 |
7.84 | LDD1839 | [6] | |
|
C170 Probe Info |
 |
23.75 | LDD1850 | [6] | |
|
C171 Probe Info |
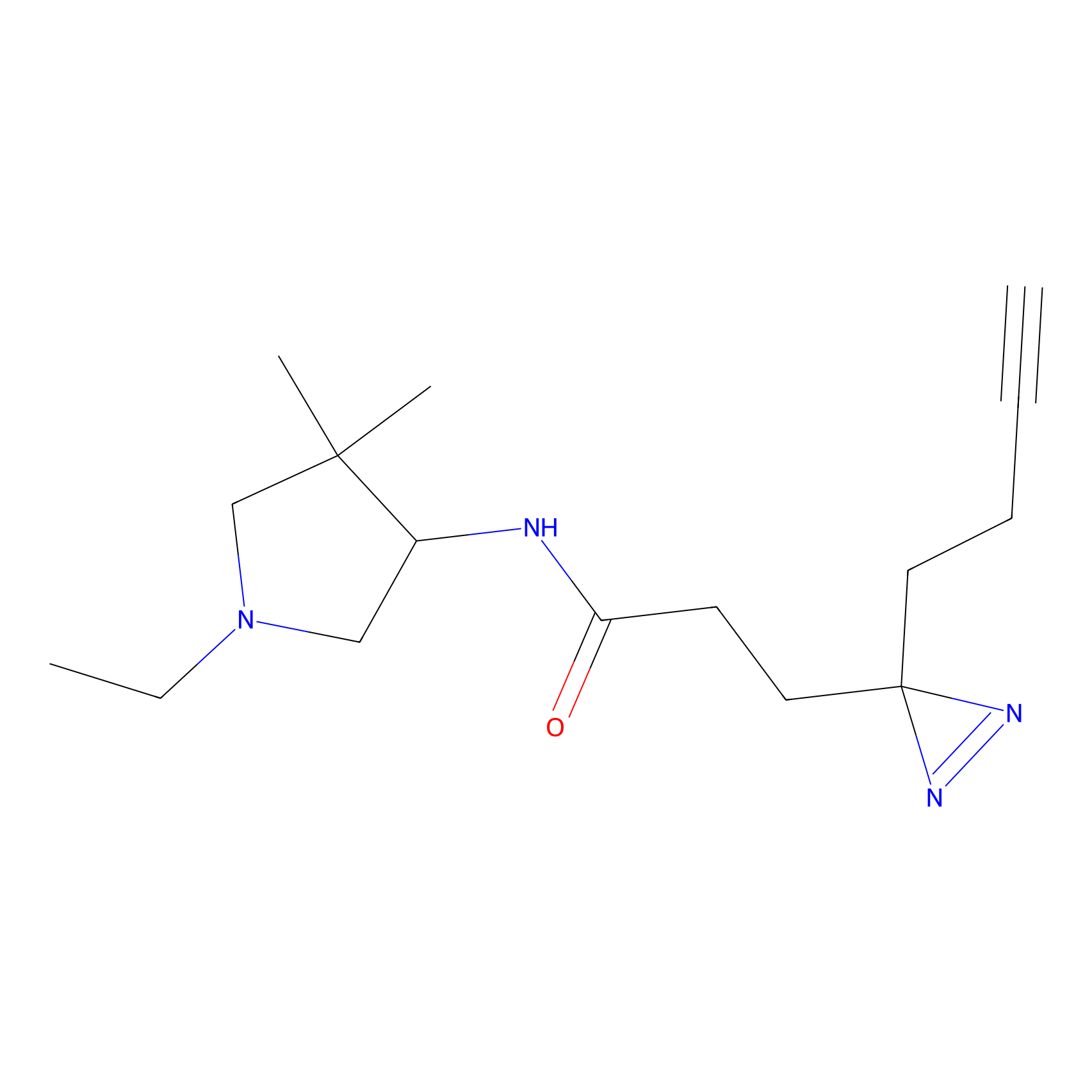 |
5.17 | LDD1851 | [6] | |
|
C197 Probe Info |
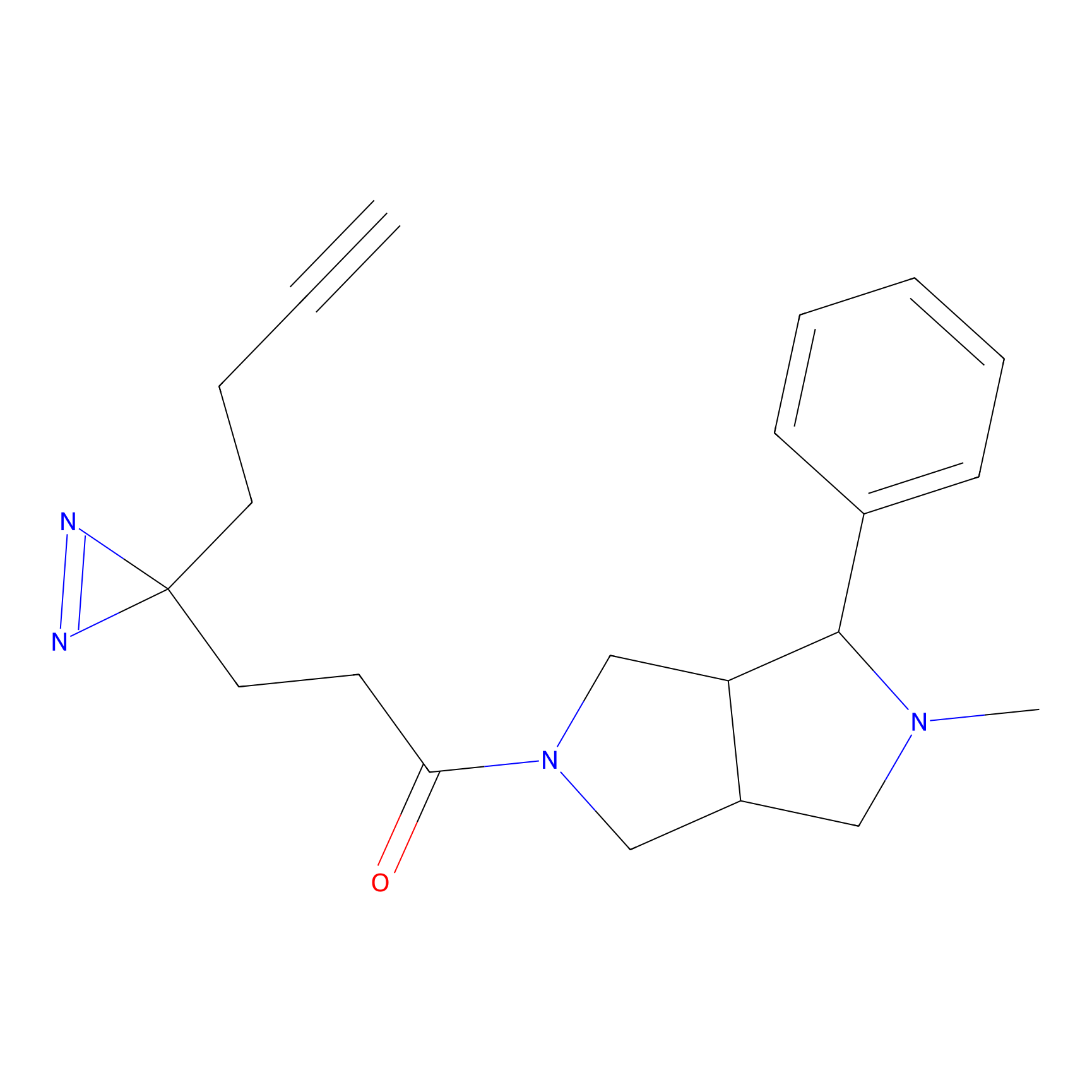 |
7.46 | LDD1873 | [6] | |
|
C237 Probe Info |
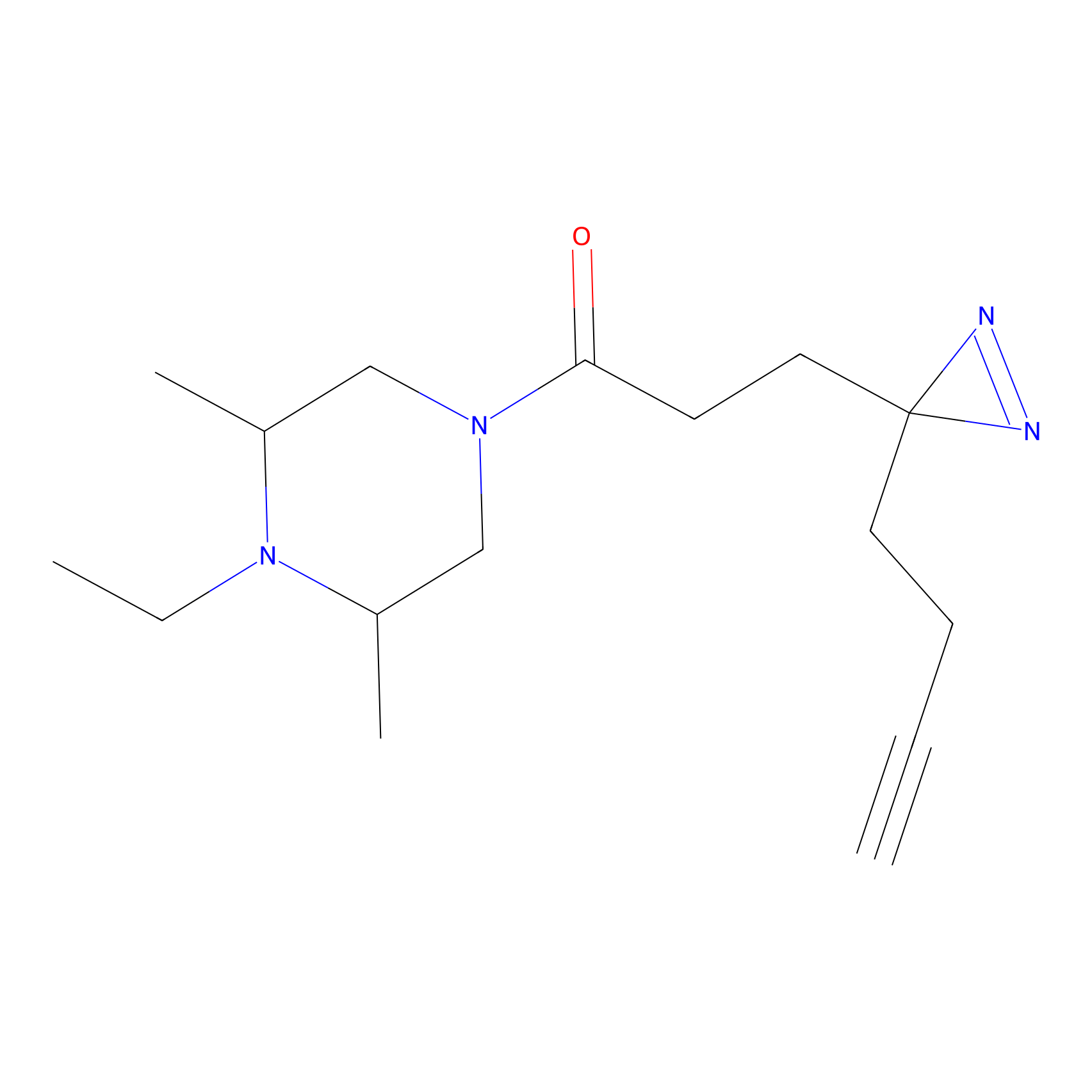 |
7.73 | LDD1910 | [6] | |
|
C281 Probe Info |
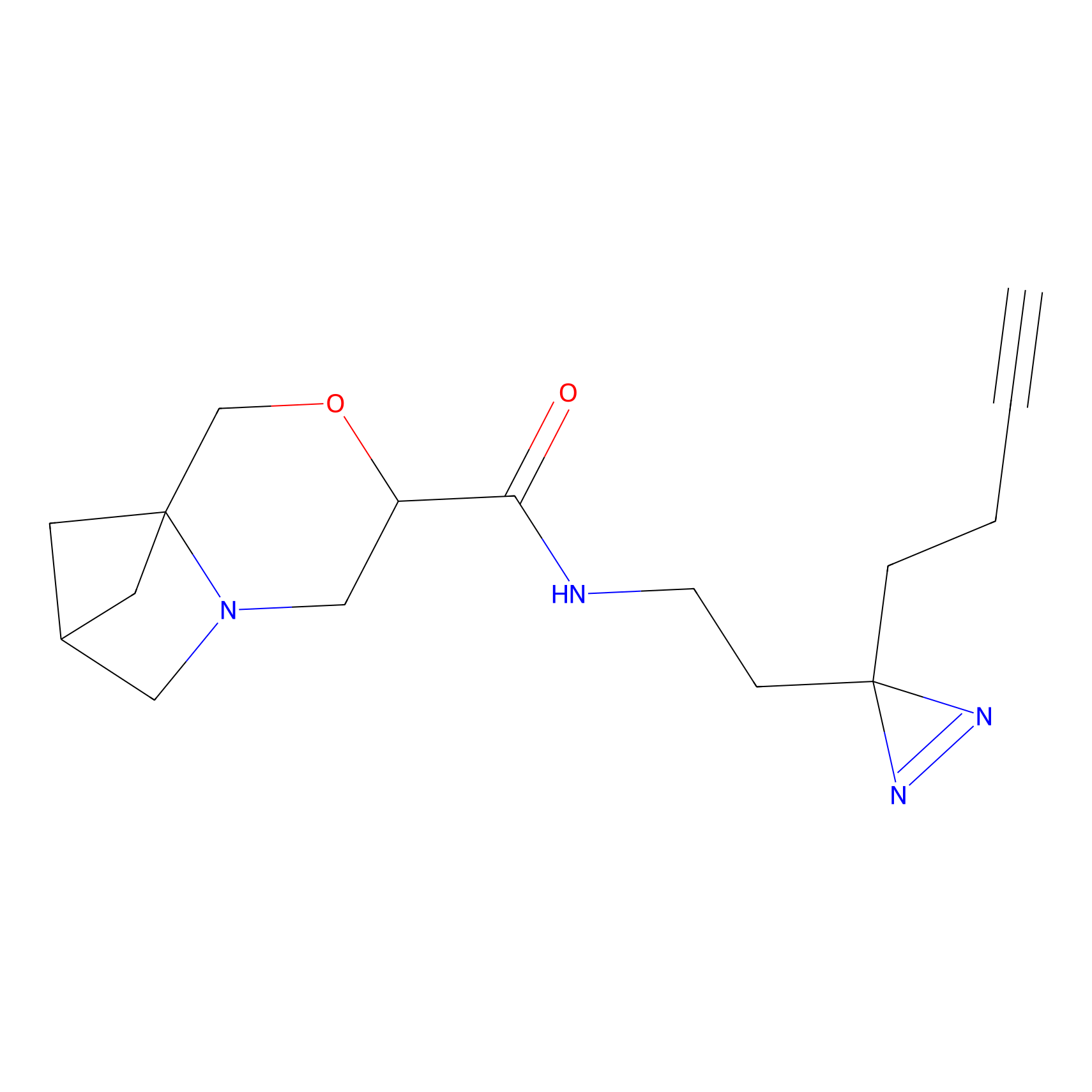 |
9.32 | LDD1951 | [6] | |
|
C310 Probe Info |
 |
12.21 | LDD1977 | [6] | |
|
C344 Probe Info |
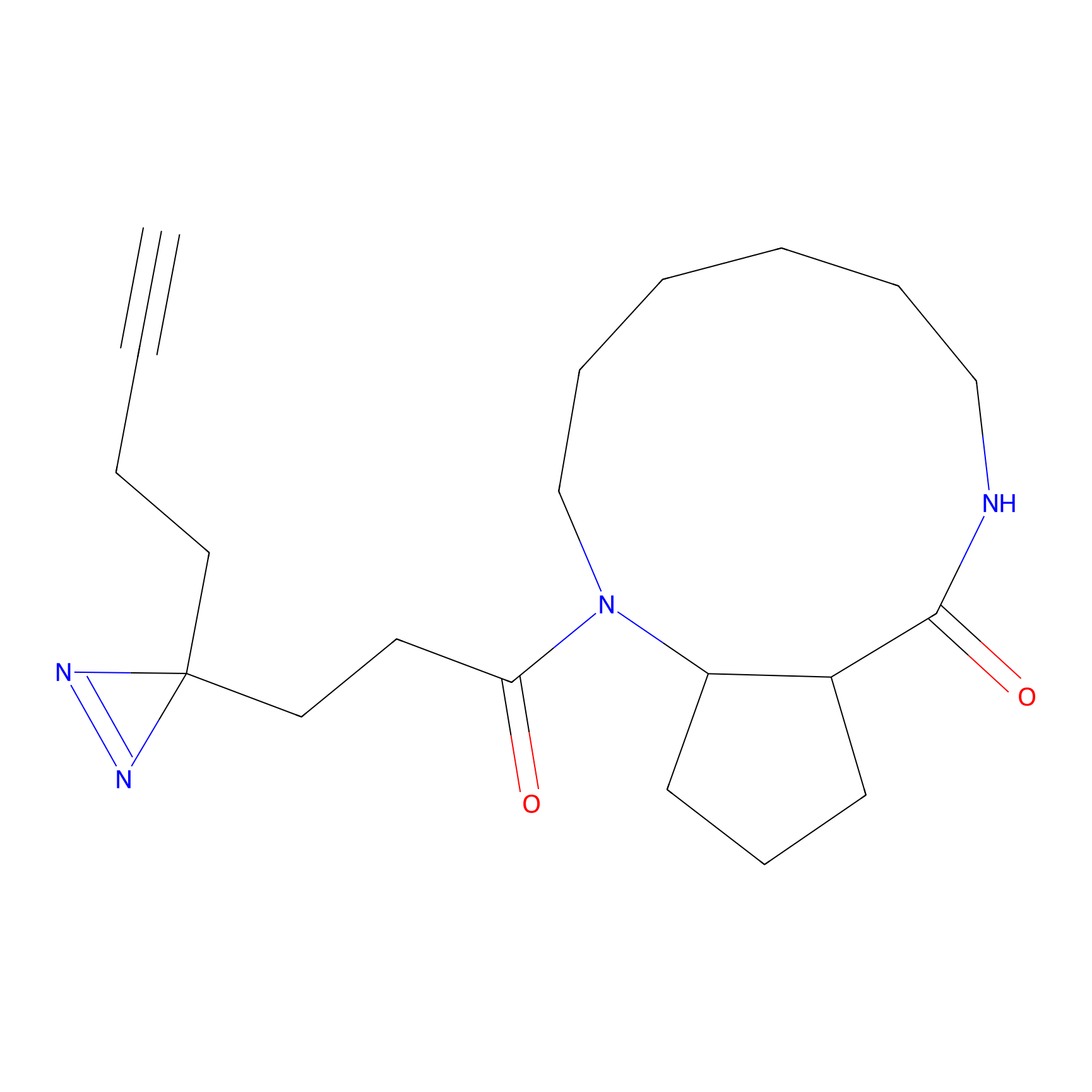 |
5.31 | LDD2006 | [6] | |
|
C355 Probe Info |
 |
48.50 | LDD2016 | [6] | |
|
C405 Probe Info |
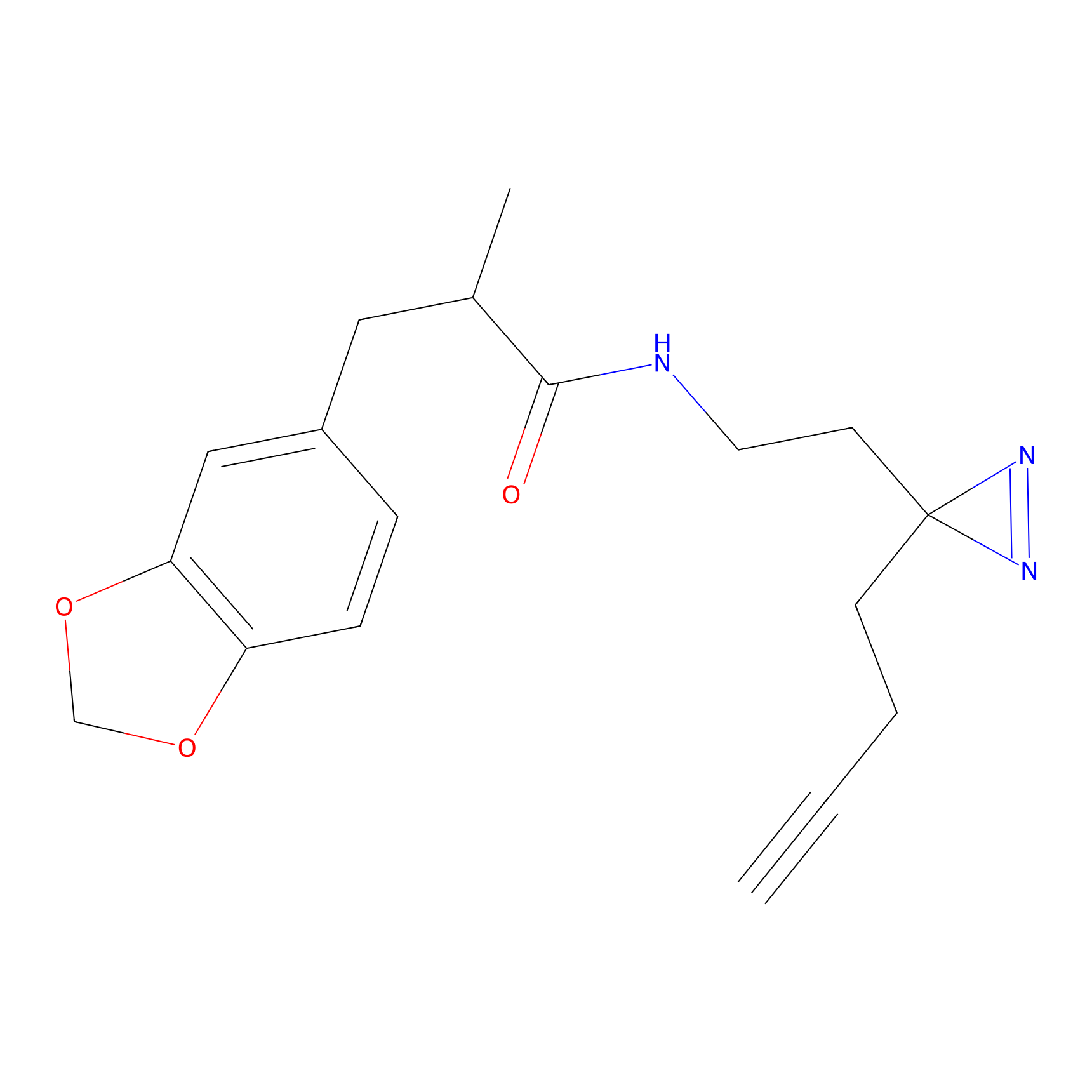 |
5.13 | LDD2063 | [6] | |
|
C407 Probe Info |
 |
11.55 | LDD2064 | [6] | |
|
C423 Probe Info |
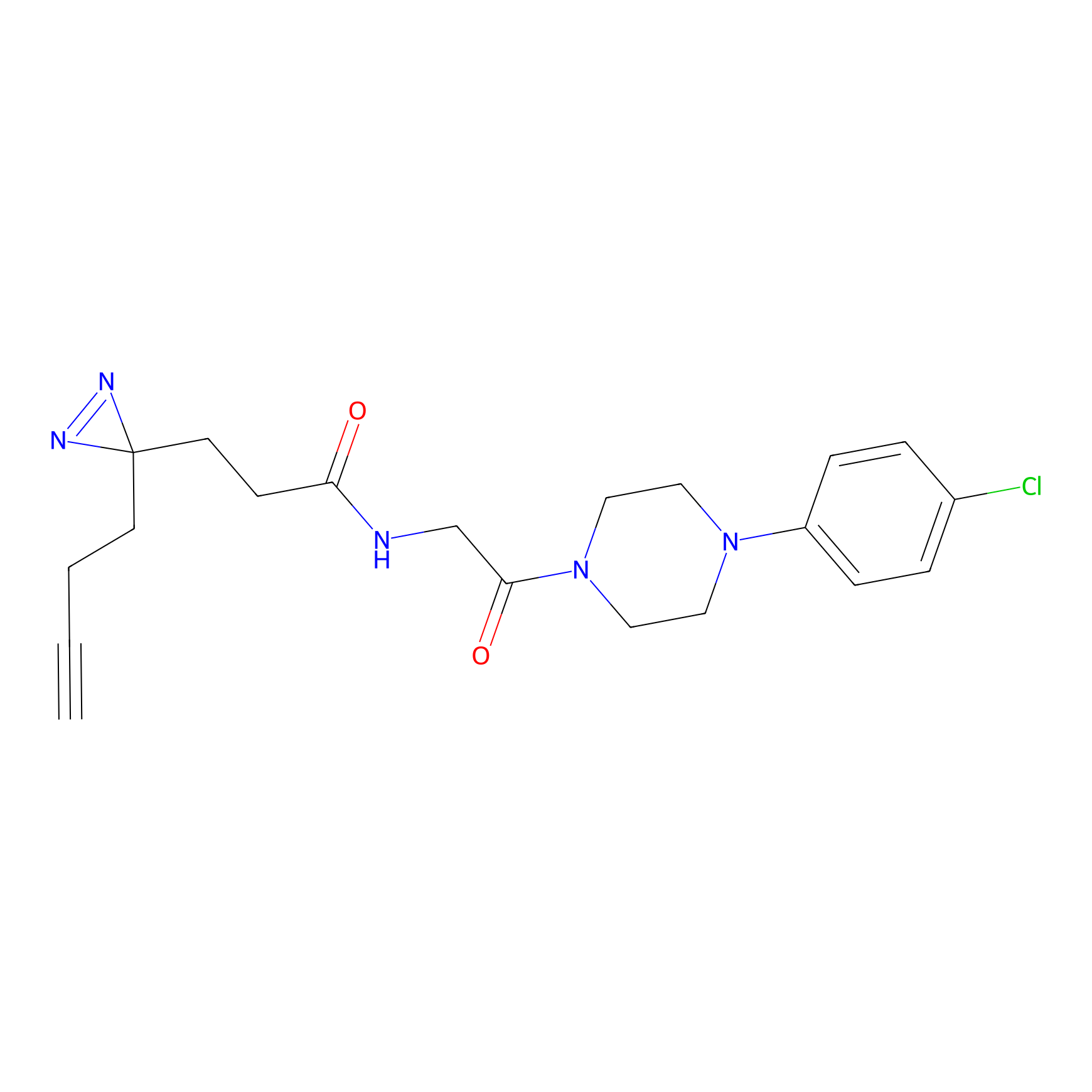 |
17.03 | LDD2078 | [6] | |
Competitor(s) Related to This Target
References
