Details of the Target
General Information of Target
| Target ID | LDTP05038 | |||||
|---|---|---|---|---|---|---|
| Target Name | AP-2 complex subunit beta (AP2B1) | |||||
| Gene Name | AP2B1 | |||||
| Gene ID | 163 | |||||
| Synonyms |
ADTB2; CLAPB1; AP-2 complex subunit beta; AP105B; Adaptor protein complex AP-2 subunit beta; Adaptor-related protein complex 2 subunit beta; Beta-2-adaptin; Beta-adaptin; Clathrin assembly protein complex 2 beta large chain; Plasma membrane adaptor HA2/AP2 adaptin beta subunit
|
|||||
| 3D Structure | ||||||
| Sequence |
MTDSKYFTTNKKGEIFELKAELNNEKKEKRKEAVKKVIAAMTVGKDVSSLFPDVVNCMQT
DNLELKKLVYLYLMNYAKSQPDMAIMAVNSFVKDCEDPNPLIRALAVRTMGCIRVDKITE YLCEPLRKCLKDEDPYVRKTAAVCVAKLHDINAQMVEDQGFLDSLRDLIADSNPMVVANA VAALSEISESHPNSNLLDLNPQNINKLLTALNECTEWGQIFILDCLSNYNPKDDREAQSI CERVTPRLSHANSAVVLSAVKVLMKFLELLPKDSDYYNMLLKKLAPPLVTLLSGEPEVQY VALRNINLIVQKRPEILKQEIKVFFVKYNDPIYVKLEKLDIMIRLASQANIAQVLAELKE YATEVDVDFVRKAVRAIGRCAIKVEQSAERCVSTLLDLIQTKVNYVVQEAIVVIRDIFRK YPNKYESIIATLCENLDSLDEPDARAAMIWIVGEYAERIDNADELLESFLEGFHDESTQV QLTLLTAIVKLFLKKPSETQELVQQVLSLATQDSDNPDLRDRGYIYWRLLSTDPVTAKEV VLSEKPLISEETDLIEPTLLDELICHIGSLASVYHKPPNAFVEGSHGIHRKHLPIHHGST DAGDSPVGTTTATNLEQPQVIPSQGDLLGDLLNLDLGPPVNVPQVSSMQMGAVDLLGGGL DSLVGQSFIPSSVPATFAPSPTPAVVSSGLNDLFELSTGIGMAPGGYVAPKAVWLPAVKA KGLEISGTFTHRQGHIYMEMNFTNKALQHMTDFAIQFNKNSFGVIPSTPLAIHTPLMPNQ SIDVSLPLNTLGPVMKMEPLNNLQVAVKNNIDVFYFSCLIPLNVLFVEDGKMERQVFLAT WKDIPNENELQFQIKECHLNADTVSSKLQNNNVYTIAKRNVEGQDMLYQSLKLTNGIWIL AELRIQPGNPNYTLSLKCRAPEVSQYIYQVYDSILKN |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
Adaptor complexes large subunit family
|
|||||
| Subcellular location |
Cell membrane
|
|||||
| Function |
Component of the adaptor protein complex 2 (AP-2). Adaptor protein complexes function in protein transport via transport vesicles in different membrane traffic pathways. Adaptor protein complexes are vesicle coat components and appear to be involved in cargo selection and vesicle formation. AP-2 is involved in clathrin-dependent endocytosis in which cargo proteins are incorporated into vesicles surrounded by clathrin (clathrin-coated vesicles, CCVs) which are destined for fusion with the early endosome. The clathrin lattice serves as a mechanical scaffold but is itself unable to bind directly to membrane components. Clathrin-associated adaptor protein (AP) complexes which can bind directly to both the clathrin lattice and to the lipid and protein components of membranes are considered to be the major clathrin adaptors contributing the CCV formation. AP-2 also serves as a cargo receptor to selectively sort the membrane proteins involved in receptor-mediated endocytosis. AP-2 seems to play a role in the recycling of synaptic vesicle membranes from the presynaptic surface. AP-2 recognizes Y-X-X-[FILMV] (Y-X-X-Phi) and [ED]-X-X-X-L-[LI] endocytosis signal motifs within the cytosolic tails of transmembrane cargo molecules. AP-2 may also play a role in maintaining normal post-endocytic trafficking through the ARF6-regulated, non-clathrin pathway. During long-term potentiation in hippocampal neurons, AP-2 is responsible for the endocytosis of ADAM10. The AP-2 beta subunit acts via its C-terminal appendage domain as a scaffolding platform for endocytic accessory proteins; at least some clathrin-associated sorting proteins (CLASPs) are recognized by their [DE]-X(1,2)-F-X-X-[FL]-X-X-X-R motif. The AP-2 beta subunit binds to clathrin heavy chain, promoting clathrin lattice assembly; clathrin displaces at least some CLASPs from AP2B1 which probably then can be positioned for further coat assembly.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
8.77 | LDD0402 | [1] | |
|
A-EBA Probe Info |
 |
3.80 | LDD0215 | [2] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [3] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [3] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [3] | |
|
TH211 Probe Info |
 |
Y121(16.45) | LDD0260 | [4] | |
|
C-Sul Probe Info |
 |
2.72 | LDD0066 | [5] | |
|
TH214 Probe Info |
 |
Y121(17.16) | LDD0258 | [4] | |
|
TH216 Probe Info |
 |
Y121(8.60) | LDD0259 | [4] | |
|
STPyne Probe Info |
 |
K11(5.29); K19(1.62); K31(7.21); K719(10.00) | LDD0277 | [6] | |
|
Probe 1 Probe Info |
 |
Y136(91.27); Y333(63.65); Y361(17.40); Y524(16.39) | LDD3495 | [7] | |
|
DBIA Probe Info |
 |
C391(8.01); C241(3.67); C66(3.04); C55(4.63) | LDD3310 | [8] | |
|
AHL-Pu-1 Probe Info |
 |
C123(10.39) | LDD0168 | [9] | |
|
HHS-475 Probe Info |
 |
Y6(0.61); Y737(0.83); Y136(1.11); Y121(1.26) | LDD0264 | [10] | |
|
HHS-465 Probe Info |
 |
Y121(7.66); Y136(10.00) | LDD2237 | [11] | |
|
5E-2FA Probe Info |
 |
H749(0.00); H858(0.00); H773(0.00) | LDD2235 | [12] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [13] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C129(0.00); C918(0.00); C391(0.00); C95(0.00) | LDD0038 | [14] | |
|
IA-alkyne Probe Info |
 |
C123(0.00); C57(0.00); C380(0.00); C129(0.00) | LDD0032 | [15] | |
|
IPIAA_H Probe Info |
 |
C433(0.00); C241(0.00) | LDD0030 | [16] | |
|
IPIAA_L Probe Info |
 |
C123(0.00); C241(0.00); C565(0.00); C95(0.00) | LDD0031 | [16] | |
|
Lodoacetamide azide Probe Info |
 |
C380(0.00); C129(0.00); C918(0.00); C391(0.00) | LDD0037 | [14] | |
|
BTD Probe Info |
 |
N.A. | LDD0004 | [17] | |
|
JW-RF-010 Probe Info |
 |
C857(0.00); C380(0.00) | LDD0026 | [18] | |
|
NAIA_4 Probe Info |
 |
C57(0.00); C95(0.00); C123(0.00); C129(0.00) | LDD2226 | [19] | |
|
WYneN Probe Info |
 |
C857(0.00); C123(0.00); C391(0.00) | LDD0021 | [17] | |
|
WYneO Probe Info |
 |
C857(0.00); C95(0.00); C123(0.00) | LDD0022 | [17] | |
|
ENE Probe Info |
 |
C95(0.00); C241(0.00); C112(0.00) | LDD0006 | [17] | |
|
IPM Probe Info |
 |
C241(0.00); C144(0.00); C95(0.00); C857(0.00) | LDD0147 | [18] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0017 | [20] | |
|
TFBX Probe Info |
 |
C57(0.00); C380(0.00); C144(0.00); C95(0.00) | LDD0148 | [18] | |
|
VSF Probe Info |
 |
C123(0.00); C95(0.00); C391(0.00); C857(0.00) | LDD0007 | [17] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [21] | |
|
Acrolein Probe Info |
 |
C857(0.00); C123(0.00); C391(0.00); C144(0.00) | LDD0217 | [22] | |
|
Cinnamaldehyde Probe Info |
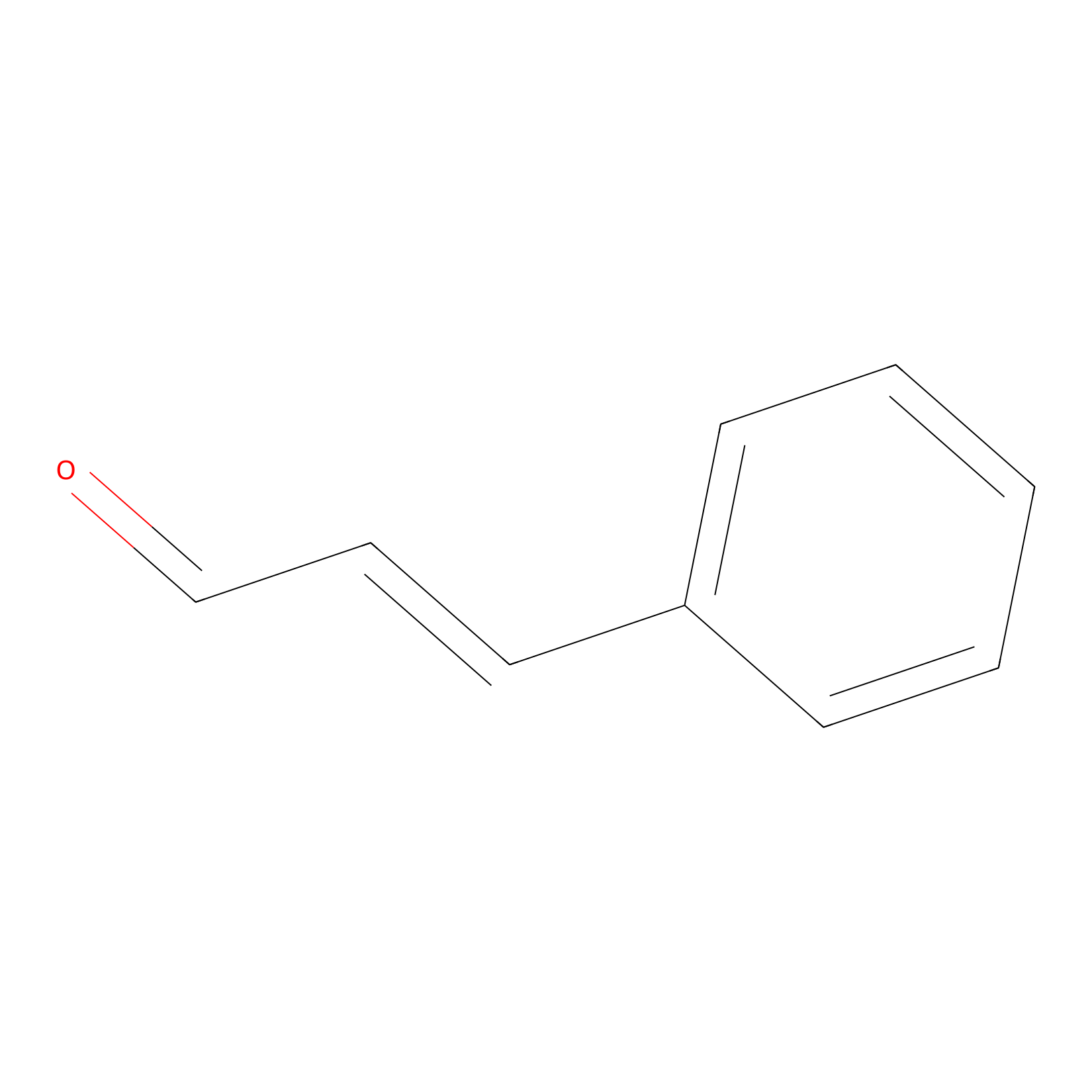 |
C857(0.00); C241(0.00) | LDD0220 | [22] | |
|
Crotonaldehyde Probe Info |
 |
C857(0.00); C241(0.00); C112(0.00); C123(0.00) | LDD0219 | [22] | |
|
Methacrolein Probe Info |
 |
C144(0.00); C123(0.00); C95(0.00); C241(0.00) | LDD0218 | [22] | |
|
W1 Probe Info |
 |
C241(0.00); C123(0.00); C857(0.00) | LDD0236 | [23] | |
|
AOyne Probe Info |
 |
7.30 | LDD0443 | [24] | |
|
NAIA_5 Probe Info |
 |
C380(0.00); C129(0.00); C918(0.00); C391(0.00) | LDD2223 | [19] | |
|
HHS-482 Probe Info |
 |
Y121(0.98); Y328(0.79); Y737(1.02) | LDD2239 | [11] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe11 Probe Info |
 |
6.04 | LDD0471 | [25] | |
|
FFF probe13 Probe Info |
 |
12.53 | LDD0475 | [25] | |
|
FFF probe14 Probe Info |
 |
12.04 | LDD0477 | [25] | |
|
FFF probe3 Probe Info |
 |
7.08 | LDD0464 | [25] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0136 | [26] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [26] | |
|
Alk-rapa Probe Info |
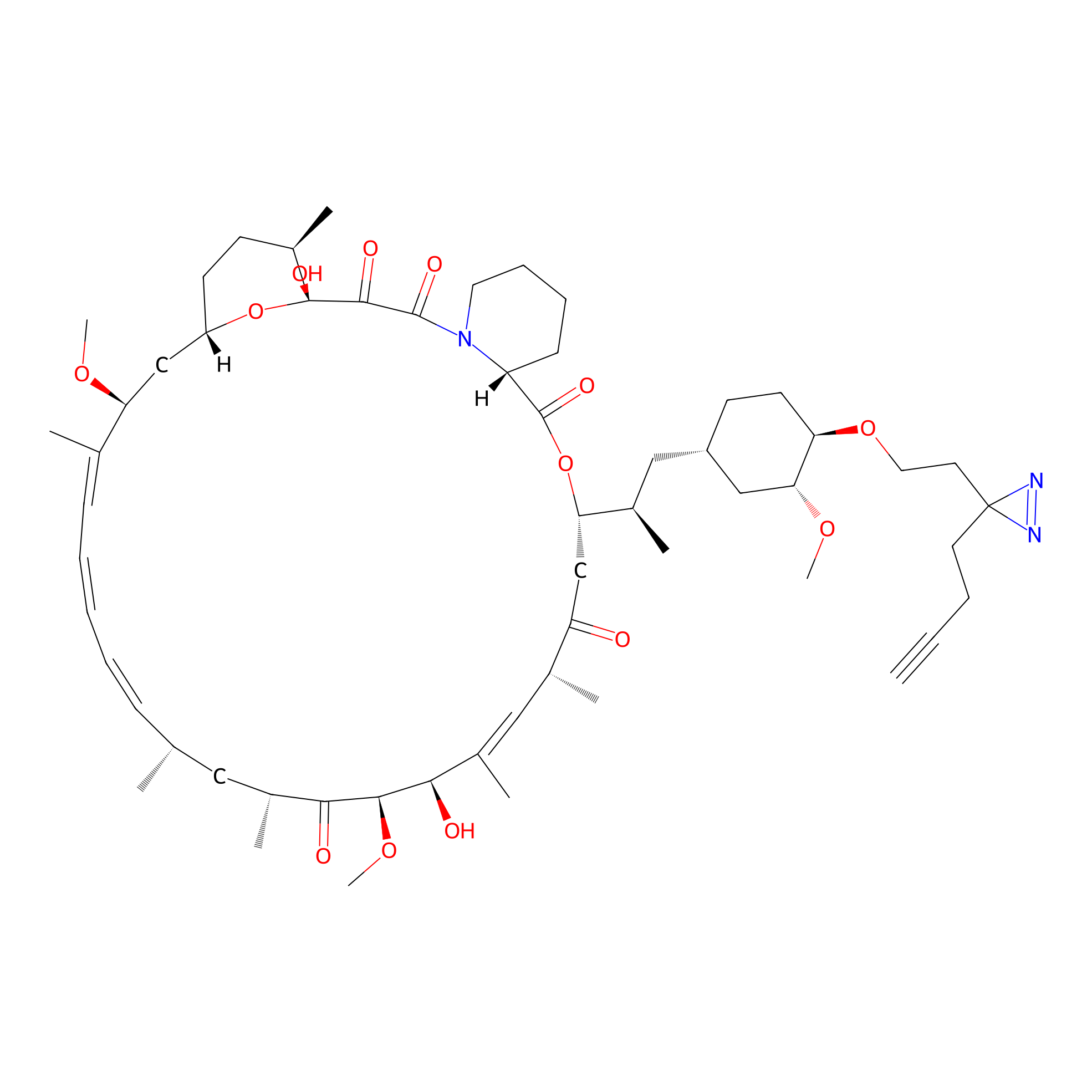 |
4.33 | LDD0213 | [27] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0071 | [28] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C857(0.53) | LDD2142 | [29] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C857(0.86) | LDD2112 | [29] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C857(0.91); C123(0.83); C112(0.52) | LDD2095 | [29] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C857(0.80) | LDD2130 | [29] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C857(1.03); C123(1.35); C112(1.07); C144(0.89) | LDD2117 | [29] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C857(2.20); C112(1.42); C241(1.27); C144(0.97) | LDD2152 | [29] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C123(0.83); C241(1.31) | LDD2103 | [29] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C857(0.98); C112(0.35); C95(0.63); C241(1.01) | LDD2132 | [29] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C857(0.45); C112(0.73); C95(0.66) | LDD2131 | [29] |
| LDCM0025 | 4SU-RNA | HEK-293T | C123(10.39) | LDD0168 | [9] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C123(10.60); C112(3.13) | LDD0169 | [9] |
| LDCM0214 | AC1 | HEK-293T | C57(1.02); C433(1.03); C857(0.98); C225(1.03) | LDD1507 | [30] |
| LDCM0215 | AC10 | HEK-293T | C57(1.07); C433(1.33); C857(1.05); C225(0.97) | LDD1508 | [30] |
| LDCM0226 | AC11 | HEK-293T | C57(1.09); C433(0.85); C857(0.87) | LDD1509 | [30] |
| LDCM0237 | AC12 | HEK-293T | C57(1.01); C433(0.80); C857(1.14) | LDD1510 | [30] |
| LDCM0259 | AC14 | HEK-293T | C57(0.98); C433(0.98); C857(1.12); C225(1.11) | LDD1512 | [30] |
| LDCM0270 | AC15 | HEK-293T | C57(0.95); C433(1.06); C857(1.06) | LDD1513 | [30] |
| LDCM0276 | AC17 | HEK-293T | C57(1.01); C433(0.86); C857(1.01); C225(0.76) | LDD1515 | [30] |
| LDCM0277 | AC18 | HEK-293T | C57(1.03); C433(1.33); C857(1.03); C225(1.02) | LDD1516 | [30] |
| LDCM0278 | AC19 | HEK-293T | C57(1.07); C433(0.74); C857(0.80) | LDD1517 | [30] |
| LDCM0279 | AC2 | HEK-293T | C57(1.13); C433(1.55); C857(1.05); C225(0.99) | LDD1518 | [30] |
| LDCM0280 | AC20 | HEK-293T | C57(1.06); C433(0.96); C857(1.26) | LDD1519 | [30] |
| LDCM0281 | AC21 | HEK-293T | C57(0.99); C433(0.95); C857(1.03) | LDD1520 | [30] |
| LDCM0282 | AC22 | HEK-293T | C57(0.98); C433(0.96); C857(1.07); C225(1.09) | LDD1521 | [30] |
| LDCM0283 | AC23 | HEK-293T | C57(0.89); C433(0.80); C857(1.00) | LDD1522 | [30] |
| LDCM0284 | AC24 | HEK-293T | C57(0.96); C433(0.98); C857(1.09) | LDD1523 | [30] |
| LDCM0285 | AC25 | HEK-293T | C57(1.04); C433(0.87); C857(1.00); C225(0.86) | LDD1524 | [30] |
| LDCM0286 | AC26 | HEK-293T | C57(1.06); C433(1.14); C857(1.08); C225(1.18) | LDD1525 | [30] |
| LDCM0287 | AC27 | HEK-293T | C57(1.00); C433(0.84); C857(0.94) | LDD1526 | [30] |
| LDCM0288 | AC28 | HEK-293T | C57(1.04); C433(1.11); C857(1.18) | LDD1527 | [30] |
| LDCM0289 | AC29 | HEK-293T | C57(1.00); C433(0.99); C857(1.05) | LDD1528 | [30] |
| LDCM0290 | AC3 | HEK-293T | C57(1.03); C433(0.92); C857(1.12) | LDD1529 | [30] |
| LDCM0291 | AC30 | HEK-293T | C57(0.97); C433(0.95); C857(1.11); C225(1.14) | LDD1530 | [30] |
| LDCM0292 | AC31 | HEK-293T | C57(0.92); C433(0.93); C857(1.09) | LDD1531 | [30] |
| LDCM0293 | AC32 | HEK-293T | C57(0.96); C433(0.88); C857(1.03) | LDD1532 | [30] |
| LDCM0294 | AC33 | HEK-293T | C57(1.05); C433(1.01); C857(1.00); C225(1.10) | LDD1533 | [30] |
| LDCM0295 | AC34 | HEK-293T | C57(1.10); C433(1.28); C857(1.00); C225(0.95) | LDD1534 | [30] |
| LDCM0296 | AC35 | HEK-293T | C57(1.03); C433(0.83); C857(0.95) | LDD1535 | [30] |
| LDCM0297 | AC36 | HEK-293T | C57(1.04); C433(0.88); C857(1.07) | LDD1536 | [30] |
| LDCM0298 | AC37 | HEK-293T | C57(1.00); C433(1.02); C857(0.98) | LDD1537 | [30] |
| LDCM0299 | AC38 | HEK-293T | C57(1.05); C433(1.01); C857(1.09); C225(1.23) | LDD1538 | [30] |
| LDCM0300 | AC39 | HEK-293T | C57(0.91); C433(1.18); C857(1.03) | LDD1539 | [30] |
| LDCM0301 | AC4 | HEK-293T | C57(1.10); C433(1.11); C857(1.55) | LDD1540 | [30] |
| LDCM0302 | AC40 | HEK-293T | C57(1.01); C433(0.99); C857(1.07) | LDD1541 | [30] |
| LDCM0303 | AC41 | HEK-293T | C57(1.01); C433(0.88); C857(0.99); C225(1.09) | LDD1542 | [30] |
| LDCM0304 | AC42 | HEK-293T | C57(1.11); C433(1.21); C857(1.04); C225(0.98) | LDD1543 | [30] |
| LDCM0305 | AC43 | HEK-293T | C57(1.04); C433(1.06); C857(0.93) | LDD1544 | [30] |
| LDCM0306 | AC44 | HEK-293T | C57(1.02); C433(0.96); C857(1.00) | LDD1545 | [30] |
| LDCM0307 | AC45 | HEK-293T | C57(0.99); C433(0.97); C857(1.01) | LDD1546 | [30] |
| LDCM0308 | AC46 | HEK-293T | C57(1.00); C433(0.93); C857(1.15); C225(1.17) | LDD1547 | [30] |
| LDCM0309 | AC47 | HEK-293T | C57(0.90); C433(0.95); C857(1.06) | LDD1548 | [30] |
| LDCM0310 | AC48 | HEK-293T | C57(0.97); C433(1.03); C857(1.01) | LDD1549 | [30] |
| LDCM0311 | AC49 | HEK-293T | C57(1.05); C433(0.72); C857(0.96); C225(0.96) | LDD1550 | [30] |
| LDCM0312 | AC5 | HEK-293T | C57(1.06); C433(1.21); C857(1.03) | LDD1551 | [30] |
| LDCM0313 | AC50 | HEK-293T | C57(1.10); C433(1.27); C857(0.98); C225(0.87) | LDD1552 | [30] |
| LDCM0314 | AC51 | HEK-293T | C57(1.12); C433(0.87); C857(0.88) | LDD1553 | [30] |
| LDCM0315 | AC52 | HEK-293T | C57(1.03); C433(0.97); C857(0.85) | LDD1554 | [30] |
| LDCM0316 | AC53 | HEK-293T | C57(1.05); C433(0.95); C857(1.00) | LDD1555 | [30] |
| LDCM0317 | AC54 | HEK-293T | C57(0.99); C433(1.01); C857(1.07); C225(0.88) | LDD1556 | [30] |
| LDCM0318 | AC55 | HEK-293T | C57(0.93); C433(0.96); C857(0.96) | LDD1557 | [30] |
| LDCM0319 | AC56 | HEK-293T | C57(1.01); C433(0.92); C857(1.05) | LDD1558 | [30] |
| LDCM0320 | AC57 | HEK-293T | C57(1.06); C433(0.88); C857(1.05); C225(1.13) | LDD1559 | [30] |
| LDCM0321 | AC58 | HEK-293T | C57(1.17); C433(1.39); C857(1.09); C225(0.93) | LDD1560 | [30] |
| LDCM0322 | AC59 | HEK-293T | C57(1.13); C433(0.76); C857(1.01) | LDD1561 | [30] |
| LDCM0323 | AC6 | HEK-293T | C57(1.00); C433(0.89); C857(1.06); C225(1.00) | LDD1562 | [30] |
| LDCM0324 | AC60 | HEK-293T | C57(1.13); C433(1.04); C857(1.20) | LDD1563 | [30] |
| LDCM0325 | AC61 | HEK-293T | C57(1.03); C433(1.22); C857(1.07) | LDD1564 | [30] |
| LDCM0326 | AC62 | HEK-293T | C57(1.05); C433(0.99); C857(1.07); C225(1.24) | LDD1565 | [30] |
| LDCM0327 | AC63 | HEK-293T | C57(1.02); C433(0.79); C857(1.11) | LDD1566 | [30] |
| LDCM0328 | AC64 | HEK-293T | C57(1.01); C433(0.93); C857(1.14) | LDD1567 | [30] |
| LDCM0334 | AC7 | HEK-293T | C57(1.01); C433(1.08); C857(1.07) | LDD1568 | [30] |
| LDCM0345 | AC8 | HEK-293T | C57(1.04); C433(0.89); C857(1.05) | LDD1569 | [30] |
| LDCM0545 | Acetamide | MDA-MB-231 | C857(0.55) | LDD2138 | [29] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C857(0.78); C123(0.59); C112(0.89); C95(0.93) | LDD2113 | [29] |
| LDCM0248 | AKOS034007472 | HEK-293T | C57(1.01); C433(1.04); C857(1.04) | LDD1511 | [30] |
| LDCM0356 | AKOS034007680 | HEK-293T | C57(1.01); C433(0.91); C857(0.92); C225(1.10) | LDD1570 | [30] |
| LDCM0275 | AKOS034007705 | HEK-293T | C57(1.04); C433(1.00); C857(1.10) | LDD1514 | [30] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C857(0.64); C123(1.01); C112(0.81); C241(0.66) | LDD2091 | [29] |
| LDCM0108 | Chloroacetamide | HeLa | C123(0.00); C57(0.00); C144(0.00); C857(0.00) | LDD0222 | [22] |
| LDCM0367 | CL1 | HEK-293T | C57(1.07); C433(0.93); C857(1.00); C225(0.94) | LDD1571 | [30] |
| LDCM0368 | CL10 | HEK-293T | C57(0.88); C433(0.86); C857(1.06); C225(1.17) | LDD1572 | [30] |
| LDCM0369 | CL100 | HEK-293T | C57(1.11); C433(0.94); C857(0.95); C214(0.75) | LDD1573 | [30] |
| LDCM0370 | CL101 | HEK-293T | C57(1.06); C433(1.27); C857(1.05); C225(0.84) | LDD1574 | [30] |
| LDCM0371 | CL102 | HEK-293T | C57(1.10); C433(0.72); C857(1.05) | LDD1575 | [30] |
| LDCM0372 | CL103 | HEK-293T | C57(0.98); C433(0.90); C857(1.17); C225(0.94) | LDD1576 | [30] |
| LDCM0373 | CL104 | HEK-293T | C57(1.05); C433(1.41); C857(0.99); C214(0.76) | LDD1577 | [30] |
| LDCM0374 | CL105 | HEK-293T | C57(1.00); C433(1.20); C857(0.97); C225(0.80) | LDD1578 | [30] |
| LDCM0375 | CL106 | HEK-293T | C57(0.99); C433(0.98); C857(1.00) | LDD1579 | [30] |
| LDCM0376 | CL107 | HEK-293T | C57(0.97); C433(1.36); C857(1.00); C225(1.02) | LDD1580 | [30] |
| LDCM0377 | CL108 | HEK-293T | C57(1.00); C433(0.97); C857(1.00); C214(0.90) | LDD1581 | [30] |
| LDCM0378 | CL109 | HEK-293T | C57(1.02); C433(1.37); C857(0.94); C225(0.87) | LDD1582 | [30] |
| LDCM0379 | CL11 | HEK-293T | C57(0.73); C433(1.23); C857(1.19) | LDD1583 | [30] |
| LDCM0380 | CL110 | HEK-293T | C57(1.10); C433(1.00); C857(1.00) | LDD1584 | [30] |
| LDCM0381 | CL111 | HEK-293T | C57(0.99); C433(1.01); C857(1.07); C225(0.98) | LDD1585 | [30] |
| LDCM0382 | CL112 | HEK-293T | C57(1.06); C433(0.96); C857(0.99); C214(0.92) | LDD1586 | [30] |
| LDCM0383 | CL113 | HEK-293T | C57(1.00); C433(1.16); C857(1.00); C225(1.05) | LDD1587 | [30] |
| LDCM0384 | CL114 | HEK-293T | C57(1.25); C433(0.87); C857(1.00) | LDD1588 | [30] |
| LDCM0385 | CL115 | HEK-293T | C57(0.93); C433(1.10); C857(1.14); C225(1.04) | LDD1589 | [30] |
| LDCM0386 | CL116 | HEK-293T | C57(1.07); C433(0.96); C857(1.01); C214(0.97) | LDD1590 | [30] |
| LDCM0387 | CL117 | HEK-293T | C57(1.06); C433(1.25); C857(1.01); C225(0.88) | LDD1591 | [30] |
| LDCM0388 | CL118 | HEK-293T | C57(1.07); C433(1.06); C857(1.04) | LDD1592 | [30] |
| LDCM0389 | CL119 | HEK-293T | C57(0.96); C433(0.69); C857(1.05); C225(0.97) | LDD1593 | [30] |
| LDCM0390 | CL12 | HEK-293T | C57(0.89); C433(1.32); C857(1.17) | LDD1594 | [30] |
| LDCM0391 | CL120 | HEK-293T | C57(1.03); C433(0.95); C857(1.01); C214(0.90) | LDD1595 | [30] |
| LDCM0392 | CL121 | HEK-293T | C57(1.08); C433(1.22); C857(1.04); C225(0.84) | LDD1596 | [30] |
| LDCM0393 | CL122 | HEK-293T | C57(1.10); C433(0.87); C857(0.95) | LDD1597 | [30] |
| LDCM0394 | CL123 | HEK-293T | C57(1.05); C433(0.67); C857(1.08); C225(0.91) | LDD1598 | [30] |
| LDCM0395 | CL124 | HEK-293T | C57(1.05); C433(1.01); C857(0.95); C214(0.73) | LDD1599 | [30] |
| LDCM0396 | CL125 | HEK-293T | C57(1.38); C433(1.39); C857(1.04); C225(0.97) | LDD1600 | [30] |
| LDCM0397 | CL126 | HEK-293T | C57(1.34); C433(0.82); C857(1.03) | LDD1601 | [30] |
| LDCM0398 | CL127 | HEK-293T | C57(1.13); C433(0.72); C857(1.05); C225(1.06) | LDD1602 | [30] |
| LDCM0399 | CL128 | HEK-293T | C57(1.18); C433(0.99); C857(1.01); C214(0.96) | LDD1603 | [30] |
| LDCM0400 | CL13 | HEK-293T | C57(1.06); C433(1.42); C857(0.97); C225(0.88) | LDD1604 | [30] |
| LDCM0401 | CL14 | HEK-293T | C57(0.92); C433(0.92); C857(1.02) | LDD1605 | [30] |
| LDCM0402 | CL15 | HEK-293T | C57(1.02); C433(0.55); C857(1.11); C225(1.16) | LDD1606 | [30] |
| LDCM0403 | CL16 | HEK-293T | C57(0.97); C433(1.07); C857(1.07); C214(0.92) | LDD1607 | [30] |
| LDCM0404 | CL17 | HEK-293T | C57(1.02); C433(0.92); C857(0.86); C225(1.16) | LDD1608 | [30] |
| LDCM0405 | CL18 | HEK-293T | C57(0.87); C433(1.52); C857(1.05); C225(1.06) | LDD1609 | [30] |
| LDCM0406 | CL19 | HEK-293T | C57(0.86); C433(0.93); C857(0.79) | LDD1610 | [30] |
| LDCM0407 | CL2 | HEK-293T | C57(1.06); C433(1.12); C857(0.89) | LDD1611 | [30] |
| LDCM0408 | CL20 | HEK-293T | C57(0.81); C433(0.94); C857(1.11) | LDD1612 | [30] |
| LDCM0409 | CL21 | HEK-293T | C57(0.93); C433(0.93); C857(1.05) | LDD1613 | [30] |
| LDCM0410 | CL22 | HEK-293T | C57(0.82); C433(1.13); C857(1.10); C225(1.04) | LDD1614 | [30] |
| LDCM0411 | CL23 | HEK-293T | C57(0.67); C433(1.01); C857(1.13) | LDD1615 | [30] |
| LDCM0412 | CL24 | HEK-293T | C57(0.87); C433(1.16); C857(1.06) | LDD1616 | [30] |
| LDCM0413 | CL25 | HEK-293T | C57(1.21); C433(1.34); C857(0.95); C225(0.90) | LDD1617 | [30] |
| LDCM0414 | CL26 | HEK-293T | C57(0.92); C433(0.89); C857(1.08) | LDD1618 | [30] |
| LDCM0415 | CL27 | HEK-293T | C57(0.94); C433(0.76); C857(1.11); C225(0.93) | LDD1619 | [30] |
| LDCM0416 | CL28 | HEK-293T | C57(0.92); C433(0.80); C857(0.94); C214(0.78) | LDD1620 | [30] |
| LDCM0417 | CL29 | HEK-293T | C57(0.93); C433(0.84); C857(1.02); C225(0.93) | LDD1621 | [30] |
| LDCM0418 | CL3 | HEK-293T | C57(1.03); C433(0.74); C857(1.03); C225(1.08) | LDD1622 | [30] |
| LDCM0419 | CL30 | HEK-293T | C57(0.85); C433(1.33); C857(1.10); C225(1.12) | LDD1623 | [30] |
| LDCM0420 | CL31 | HEK-293T | C57(0.84); C433(0.87); C857(0.86) | LDD1624 | [30] |
| LDCM0421 | CL32 | HEK-293T | C57(0.79); C433(1.03); C857(1.26) | LDD1625 | [30] |
| LDCM0422 | CL33 | HEK-293T | C57(0.87); C433(0.79); C857(1.12) | LDD1626 | [30] |
| LDCM0423 | CL34 | HEK-293T | C57(0.84); C433(1.03); C857(1.06); C225(0.94) | LDD1627 | [30] |
| LDCM0424 | CL35 | HEK-293T | C57(0.66); C433(1.67); C857(1.12) | LDD1628 | [30] |
| LDCM0425 | CL36 | HEK-293T | C57(0.86); C433(1.21); C857(1.15) | LDD1629 | [30] |
| LDCM0426 | CL37 | HEK-293T | C57(1.10); C433(1.37); C857(1.01); C225(1.01) | LDD1630 | [30] |
| LDCM0428 | CL39 | HEK-293T | C57(0.91); C433(0.93); C857(0.96); C225(0.94) | LDD1632 | [30] |
| LDCM0429 | CL4 | HEK-293T | C57(1.10); C433(1.11); C857(0.97); C214(0.94) | LDD1633 | [30] |
| LDCM0430 | CL40 | HEK-293T | C57(0.97); C433(0.82); C857(0.99); C214(0.79) | LDD1634 | [30] |
| LDCM0431 | CL41 | HEK-293T | C57(0.92); C433(0.93); C857(1.03); C225(0.93) | LDD1635 | [30] |
| LDCM0432 | CL42 | HEK-293T | C57(0.77); C433(1.51); C857(1.11); C225(0.85) | LDD1636 | [30] |
| LDCM0433 | CL43 | HEK-293T | C57(0.83); C433(0.95); C857(1.03) | LDD1637 | [30] |
| LDCM0434 | CL44 | HEK-293T | C57(0.81); C433(1.08); C857(1.20) | LDD1638 | [30] |
| LDCM0435 | CL45 | HEK-293T | C57(0.86); C433(1.49); C857(1.07) | LDD1639 | [30] |
| LDCM0436 | CL46 | HEK-293T | C57(0.79); C433(1.08); C857(1.14); C225(0.82) | LDD1640 | [30] |
| LDCM0437 | CL47 | HEK-293T | C57(0.64); C433(1.18); C857(1.16) | LDD1641 | [30] |
| LDCM0438 | CL48 | HEK-293T | C57(0.82); C433(1.01); C857(1.17) | LDD1642 | [30] |
| LDCM0439 | CL49 | HEK-293T | C57(1.23); C433(1.03); C857(1.00); C225(1.01) | LDD1643 | [30] |
| LDCM0440 | CL5 | HEK-293T | C57(1.00); C433(0.84); C857(0.91); C225(1.06) | LDD1644 | [30] |
| LDCM0441 | CL50 | HEK-293T | C57(0.99); C433(1.01); C857(0.99) | LDD1645 | [30] |
| LDCM0443 | CL52 | HEK-293T | C57(0.91); C433(1.12); C857(0.97); C214(0.90) | LDD1646 | [30] |
| LDCM0444 | CL53 | HEK-293T | C57(0.96); C433(1.00); C857(0.93); C225(1.04) | LDD1647 | [30] |
| LDCM0445 | CL54 | HEK-293T | C57(1.00); C433(1.25); C857(0.93); C225(1.13) | LDD1648 | [30] |
| LDCM0446 | CL55 | HEK-293T | C57(0.79); C433(0.94); C857(1.11) | LDD1649 | [30] |
| LDCM0447 | CL56 | HEK-293T | C57(0.77); C433(1.05); C857(1.15) | LDD1650 | [30] |
| LDCM0448 | CL57 | HEK-293T | C57(0.89); C433(1.24); C857(1.08) | LDD1651 | [30] |
| LDCM0449 | CL58 | HEK-293T | C57(0.79); C433(1.09); C857(1.18); C225(1.00) | LDD1652 | [30] |
| LDCM0450 | CL59 | HEK-293T | C57(0.63); C433(1.09); C857(1.15) | LDD1653 | [30] |
| LDCM0451 | CL6 | HEK-293T | C57(1.03); C433(1.40); C857(0.95); C225(0.88) | LDD1654 | [30] |
| LDCM0452 | CL60 | HEK-293T | C57(0.84); C433(1.18); C857(1.13) | LDD1655 | [30] |
| LDCM0453 | CL61 | HEK-293T | C57(1.28); C433(1.27); C857(1.02); C225(0.98) | LDD1656 | [30] |
| LDCM0454 | CL62 | HEK-293T | C57(1.03); C433(0.85); C857(1.01) | LDD1657 | [30] |
| LDCM0455 | CL63 | HEK-293T | C57(0.94); C433(0.96); C857(1.09); C225(1.16) | LDD1658 | [30] |
| LDCM0456 | CL64 | HEK-293T | C57(1.00); C433(1.00); C857(0.94); C214(0.95) | LDD1659 | [30] |
| LDCM0457 | CL65 | HEK-293T | C57(0.94); C433(0.71); C857(0.96); C225(1.04) | LDD1660 | [30] |
| LDCM0458 | CL66 | HEK-293T | C57(0.88); C433(1.38); C857(1.05); C225(1.03) | LDD1661 | [30] |
| LDCM0459 | CL67 | HEK-293T | C57(0.89); C433(0.82); C857(0.98) | LDD1662 | [30] |
| LDCM0460 | CL68 | HEK-293T | C57(0.80); C433(1.09); C857(0.93) | LDD1663 | [30] |
| LDCM0461 | CL69 | HEK-293T | C57(0.85); C433(0.77); C857(1.03) | LDD1664 | [30] |
| LDCM0462 | CL7 | HEK-293T | C57(0.96); C433(1.03); C857(1.14) | LDD1665 | [30] |
| LDCM0463 | CL70 | HEK-293T | C57(0.80); C433(1.13); C857(1.15); C225(1.05) | LDD1666 | [30] |
| LDCM0464 | CL71 | HEK-293T | C57(0.65); C433(1.15); C857(1.08) | LDD1667 | [30] |
| LDCM0465 | CL72 | HEK-293T | C57(0.84); C433(1.21); C857(1.06) | LDD1668 | [30] |
| LDCM0466 | CL73 | HEK-293T | C57(1.21); C433(1.23); C857(1.01); C225(1.09) | LDD1669 | [30] |
| LDCM0467 | CL74 | HEK-293T | C57(1.14); C433(1.20); C857(1.09) | LDD1670 | [30] |
| LDCM0469 | CL76 | HEK-293T | C57(1.01); C433(1.22); C857(1.03); C214(0.79) | LDD1672 | [30] |
| LDCM0470 | CL77 | HEK-293T | C57(0.98); C433(0.69); C857(0.97); C225(1.04) | LDD1673 | [30] |
| LDCM0471 | CL78 | HEK-293T | C57(0.92); C433(1.29); C857(1.00); C225(1.11) | LDD1674 | [30] |
| LDCM0472 | CL79 | HEK-293T | C57(0.90); C433(0.81); C857(0.90) | LDD1675 | [30] |
| LDCM0473 | CL8 | HEK-293T | C57(1.59); C433(1.07); C857(1.64) | LDD1676 | [30] |
| LDCM0474 | CL80 | HEK-293T | C57(0.81); C433(1.02); C857(1.13) | LDD1677 | [30] |
| LDCM0475 | CL81 | HEK-293T | C57(0.91); C433(1.27); C857(1.01) | LDD1678 | [30] |
| LDCM0476 | CL82 | HEK-293T | C57(0.83); C433(1.01); C857(1.14); C225(0.98) | LDD1679 | [30] |
| LDCM0477 | CL83 | HEK-293T | C57(0.71); C433(1.24); C857(1.10) | LDD1680 | [30] |
| LDCM0478 | CL84 | HEK-293T | C57(0.92); C433(1.09); C857(1.12) | LDD1681 | [30] |
| LDCM0479 | CL85 | HEK-293T | C57(1.49); C433(1.96); C857(1.03); C225(0.94) | LDD1682 | [30] |
| LDCM0480 | CL86 | HEK-293T | C57(1.70); C433(0.94); C857(1.04) | LDD1683 | [30] |
| LDCM0481 | CL87 | HEK-293T | C57(1.08); C433(0.84); C857(0.97); C225(0.98) | LDD1684 | [30] |
| LDCM0482 | CL88 | HEK-293T | C57(1.23); C433(1.27); C857(1.02); C214(0.89) | LDD1685 | [30] |
| LDCM0483 | CL89 | HEK-293T | C57(1.09); C433(0.90); C857(0.96); C225(1.09) | LDD1686 | [30] |
| LDCM0484 | CL9 | HEK-293T | C57(0.86); C433(0.96); C857(0.96) | LDD1687 | [30] |
| LDCM0485 | CL90 | HEK-293T | C57(1.37); C433(1.34); C857(1.02); C225(1.18) | LDD1688 | [30] |
| LDCM0486 | CL91 | HEK-293T | C57(0.89); C433(0.92); C857(1.15) | LDD1689 | [30] |
| LDCM0487 | CL92 | HEK-293T | C57(0.96); C433(1.02); C857(1.08) | LDD1690 | [30] |
| LDCM0488 | CL93 | HEK-293T | C57(1.06); C433(1.19); C857(1.00) | LDD1691 | [30] |
| LDCM0489 | CL94 | HEK-293T | C57(0.88); C433(0.91); C857(1.10); C225(1.09) | LDD1692 | [30] |
| LDCM0490 | CL95 | HEK-293T | C57(1.11); C433(1.24); C857(1.07) | LDD1693 | [30] |
| LDCM0491 | CL96 | HEK-293T | C57(1.07); C433(1.14); C857(1.06) | LDD1694 | [30] |
| LDCM0492 | CL97 | HEK-293T | C57(1.19); C433(1.29); C857(0.97); C225(0.86) | LDD1695 | [30] |
| LDCM0493 | CL98 | HEK-293T | C57(1.12); C433(0.85); C857(0.94) | LDD1696 | [30] |
| LDCM0494 | CL99 | HEK-293T | C57(0.98); C433(0.78); C857(1.17); C225(1.01) | LDD1697 | [30] |
| LDCM0495 | E2913 | HEK-293T | C57(0.88); C433(0.69); C857(1.05); C225(1.04) | LDD1698 | [30] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C123(1.16) | LDD1702 | [29] |
| LDCM0468 | Fragment33 | HEK-293T | C57(0.99); C433(0.77); C857(1.06); C225(0.96) | LDD1671 | [30] |
| LDCM0427 | Fragment51 | HEK-293T | C57(0.94); C433(0.87); C857(1.06) | LDD1631 | [30] |
| LDCM0116 | HHS-0101 | DM93 | Y6(0.61); Y737(0.83); Y136(1.11); Y121(1.26) | LDD0264 | [10] |
| LDCM0117 | HHS-0201 | DM93 | Y737(0.69); Y6(0.74); Y136(0.85); Y121(1.08) | LDD0265 | [10] |
| LDCM0118 | HHS-0301 | DM93 | Y6(0.64); Y737(0.71); Y121(0.79); Y136(0.95) | LDD0266 | [10] |
| LDCM0119 | HHS-0401 | DM93 | Y737(0.81); Y121(0.94); Y136(0.99); Y6(1.05) | LDD0267 | [10] |
| LDCM0120 | HHS-0701 | DM93 | Y6(0.20); Y121(0.78); Y136(1.06); Y737(1.40) | LDD0268 | [10] |
| LDCM0107 | IAA | HeLa | C123(0.00); C144(0.00); C112(0.00); H858(0.00) | LDD0221 | [22] |
| LDCM0022 | KB02 | HEK-293T | C857(0.96); C57(0.91) | LDD1492 | [30] |
| LDCM0023 | KB03 | HEK-293T | C857(1.02); C57(1.03) | LDD1497 | [30] |
| LDCM0024 | KB05 | COLO792 | C391(8.01); C241(3.67); C66(3.04); C55(4.63) | LDD3310 | [8] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C857(0.70); C123(1.30); C144(0.55) | LDD2121 | [29] |
| LDCM0109 | NEM | HeLa | H149(0.00); H250(0.00); H858(0.00) | LDD0223 | [22] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C857(0.64) | LDD2089 | [29] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C857(0.94); C112(1.33) | LDD2093 | [29] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C857(1.49); C123(1.38) | LDD2094 | [29] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C857(0.83); C123(0.60); C144(0.13); C129(0.34) | LDD2096 | [29] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C112(0.76); C241(0.65); C144(0.79) | LDD2097 | [29] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C857(0.76); C123(0.95) | LDD2098 | [29] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C857(1.38) | LDD2099 | [29] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C857(0.54); C241(0.69) | LDD2100 | [29] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C857(0.59); C123(0.88); C112(1.46); C241(1.56) | LDD2101 | [29] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C857(0.63); C241(0.97) | LDD2104 | [29] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C857(1.07); C123(1.93); C241(1.52) | LDD2105 | [29] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C857(0.90); C123(0.99); C112(0.87); C95(1.18) | LDD2107 | [29] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C857(0.68); C123(1.04) | LDD2108 | [29] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C123(0.86); C112(0.64); C95(0.96); C241(0.93) | LDD2109 | [29] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C112(1.04); C241(1.12); C129(1.03) | LDD2111 | [29] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C857(0.49); C123(0.33); C112(0.56); C95(0.60) | LDD2115 | [29] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C241(3.06); C144(0.52); C129(0.64) | LDD2116 | [29] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C857(1.34); C144(0.52); C129(0.55); C380(1.36) | LDD2118 | [29] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C857(1.08); C112(1.91); C241(2.23) | LDD2119 | [29] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C857(0.80); C123(0.96); C112(0.95); C241(1.05) | LDD2120 | [29] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C857(0.82); C144(0.22); C129(0.31) | LDD2122 | [29] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C857(1.47); C123(1.26); C112(1.01); C241(0.91) | LDD2123 | [29] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C857(0.94); C241(0.93); C144(0.24) | LDD2124 | [29] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C857(0.88); C123(1.13); C112(0.93); C241(0.85) | LDD2125 | [29] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C857(1.00); C241(1.48); C144(0.15); C129(0.41) | LDD2126 | [29] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C857(1.02); C123(1.51); C112(1.13); C241(0.89) | LDD2127 | [29] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C857(0.98); C123(0.94) | LDD2128 | [29] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C857(0.60); C123(0.40); C112(0.66) | LDD2133 | [29] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C857(0.33); C123(0.58); C112(0.46); C95(0.63) | LDD2134 | [29] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C857(2.34); C123(1.88); C112(1.66); C95(2.24) | LDD2135 | [29] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C857(1.04); C123(1.36); C112(1.29); C95(1.96) | LDD2136 | [29] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C857(1.11); C123(1.41); C112(1.05); C241(1.01) | LDD2137 | [29] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C857(2.70) | LDD1700 | [29] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C857(0.95) | LDD2140 | [29] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C857(0.72) | LDD2141 | [29] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C857(0.90); C123(0.69); C112(0.85); C144(1.09) | LDD2143 | [29] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C857(2.11); C123(1.59); C112(2.19); C95(2.59) | LDD2144 | [29] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C123(1.10); C241(0.87); C144(0.85) | LDD2146 | [29] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C857(3.56); C112(1.47); C241(3.67) | LDD2147 | [29] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C857(0.42); C112(0.40); C144(0.38) | LDD2148 | [29] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C857(1.21); C144(0.28); C129(0.55) | LDD2149 | [29] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C857(0.54); C112(0.54); C241(1.14); C144(0.34) | LDD2150 | [29] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C144(0.99) | LDD2206 | [31] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C380(1.89); C391(0.22) | LDD2207 | [31] |
| LDCM0090 | Rapamycin | JHH-7 | 4.33 | LDD0213 | [27] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| AF4/FMR2 family member 4 (AFF4) | AF4 family | Q9UHB7 | |||
| THAP domain-containing protein 1 (THAP1) | THAP1 family | Q9NVV9 | |||
Other
References
