Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
TG42 Probe Info |
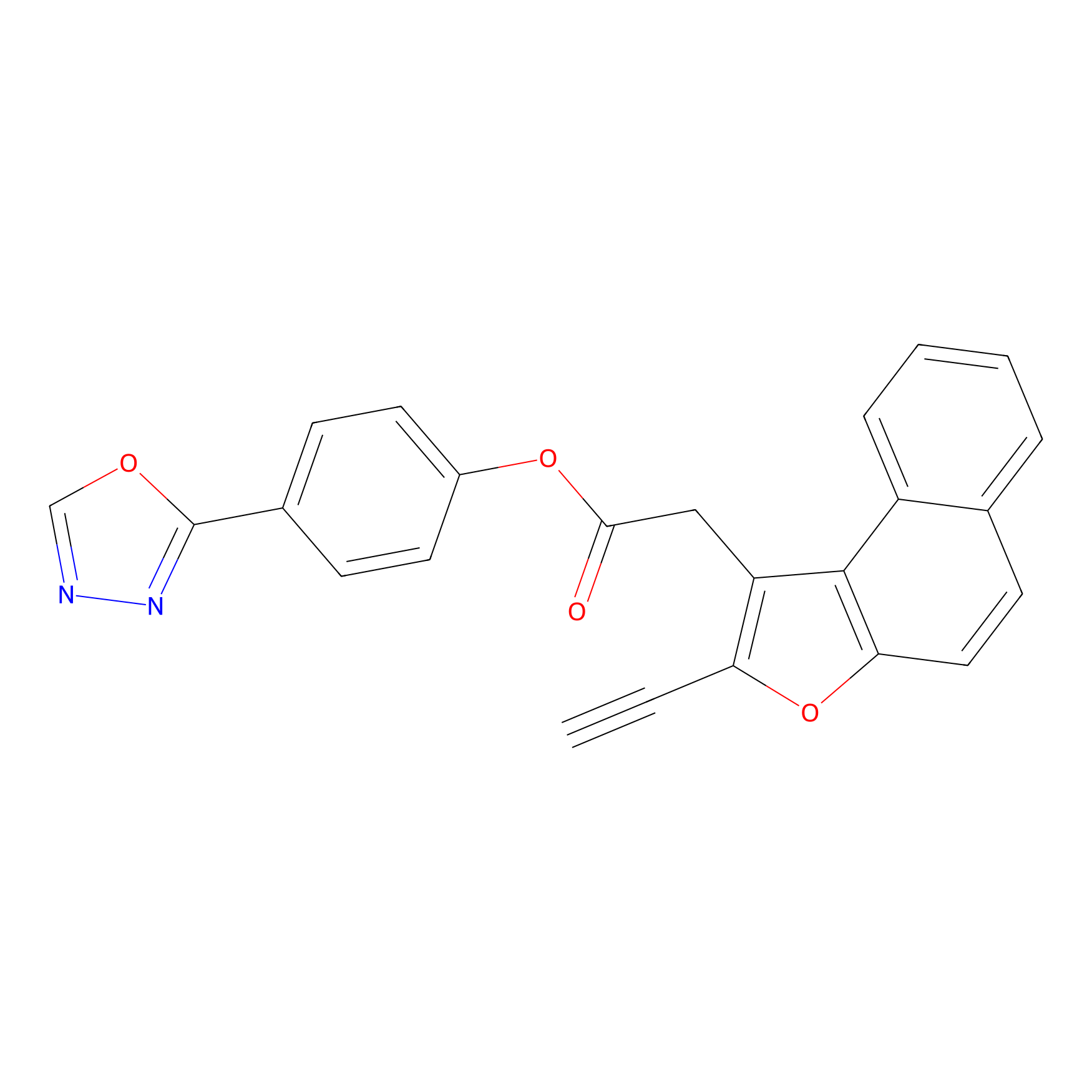 |
89.63 | LDD0326 | [1] | |
|
C-Sul Probe Info |
 |
5.08 | LDD0066 | [2] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [3] | |
|
STPyne Probe Info |
 |
K209(9.17) | LDD0277 | [4] | |
|
AZ-9 Probe Info |
 |
E206(1.01); E246(0.96) | LDD2208 | [5] | |
|
Probe 1 Probe Info |
 |
Y76(7.65) | LDD3495 | [6] | |
|
HHS-482 Probe Info |
 |
Y129(1.28); Y68(0.90); Y76(0.96) | LDD0285 | [7] | |
|
HHS-475 Probe Info |
 |
Y76(0.91); Y127(0.98); Y129(1.09) | LDD0264 | [8] | |
|
W1 Probe Info |
 |
W208(1.14) | LDD0237 | [9] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [10] | |
|
Acrolein Probe Info |
 |
H80(0.00); H12(0.00); H259(0.00); H258(0.00) | LDD0217 | [11] | |
|
Crotonaldehyde Probe Info |
 |
H12(0.00); H259(0.00) | LDD0219 | [11] | |
|
Methacrolein Probe Info |
 |
H80(0.00); H12(0.00) | LDD0218 | [11] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [12] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [12] | |
|
HHS-465 Probe Info |
 |
Y129(0.00); K135(0.00); Y76(0.00); K77(0.00) | LDD2240 | [13] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe11 Probe Info |
 |
7.73 | LDD0472 | [14] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [14] | |
|
FFF probe6 Probe Info |
 |
13.39 | LDD0468 | [14] | |
|
Staurosporine capture compound Probe Info |
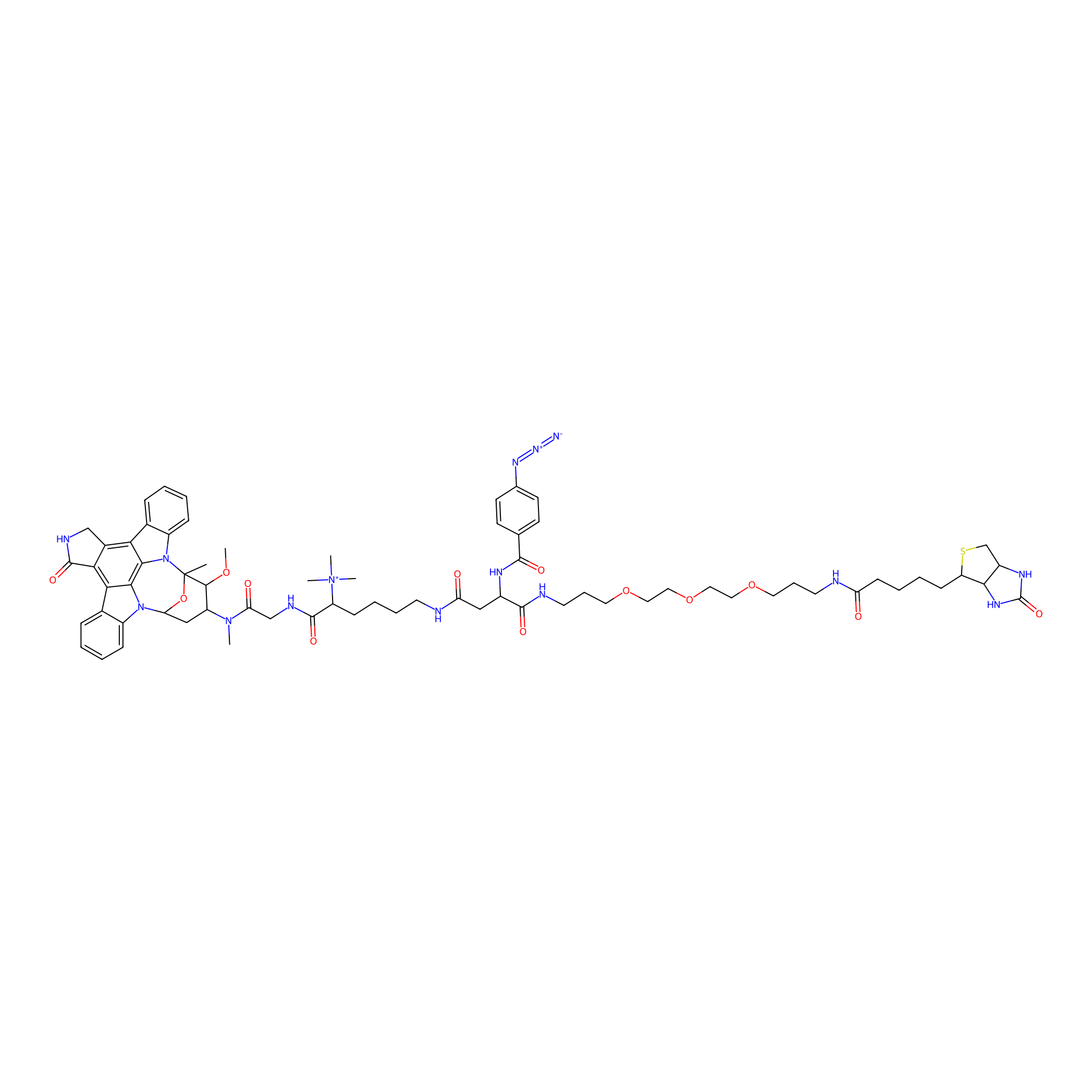 |
12.00 | LDD0083 | [15] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0072 | [16] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | H80(0.00); H12(0.00) | LDD0222 | [11] |
| LDCM0116 | HHS-0101 | DM93 | Y76(0.91); Y127(0.98); Y129(1.09) | LDD0264 | [8] |
| LDCM0117 | HHS-0201 | DM93 | Y76(0.82); Y129(1.04); Y127(1.29) | LDD0265 | [8] |
| LDCM0118 | HHS-0301 | DM93 | Y76(0.93); Y129(1.36); Y127(1.75) | LDD0266 | [8] |
| LDCM0119 | HHS-0401 | DM93 | Y129(0.91); Y127(0.96); Y76(6.13) | LDD0267 | [8] |
| LDCM0120 | HHS-0701 | DM93 | Y127(3.13); Y129(3.47); Y76(8.14) | LDD0268 | [8] |
| LDCM0107 | IAA | HeLa | H80(0.00); H12(0.00); H258(0.00); H259(0.00) | LDD0221 | [11] |
| LDCM0123 | JWB131 | DM93 | Y129(1.28); Y68(0.90); Y76(0.96) | LDD0285 | [7] |
| LDCM0124 | JWB142 | DM93 | Y129(0.72); Y43(1.50); Y68(0.62); Y76(1.06) | LDD0286 | [7] |
| LDCM0125 | JWB146 | DM93 | Y129(1.17); Y68(0.81); Y76(1.28) | LDD0287 | [7] |
| LDCM0126 | JWB150 | DM93 | Y129(3.64); Y43(16.47); Y68(6.44); Y76(4.81) | LDD0288 | [7] |
| LDCM0127 | JWB152 | DM93 | Y43(15.12); Y68(2.95); Y76(4.05) | LDD0289 | [7] |
| LDCM0128 | JWB198 | DM93 | Y129(0.78); Y68(0.96); Y76(1.30) | LDD0290 | [7] |
| LDCM0129 | JWB202 | DM93 | Y68(0.83); Y76(1.03) | LDD0291 | [7] |
| LDCM0130 | JWB211 | DM93 | Y43(5.24); Y68(1.20); Y76(2.07) | LDD0292 | [7] |
| LDCM0109 | NEM | HeLa | H80(0.00); H12(0.00) | LDD0223 | [11] |
| LDCM0019 | Staurosporine | Hep-G2 | 12.00 | LDD0083 | [15] |
| LDCM0110 | W12 | Hep-G2 | W208(1.14) | LDD0237 | [9] |
| LDCM0112 | W16 | Hep-G2 | W208(1.16) | LDD0239 | [9] |
| LDCM0113 | W17 | Hep-G2 | K210(0.91); W208(0.91) | LDD0240 | [9] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Inhibitor of growth protein 1 (ING1) | ING family | Q9UK53 | |||
The Drug(s) Related To This Target
Approved
Phase 2
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Arq 761 | . | D0DR0P | |||
| Bioe-743 | . | D0RI4I | |||
| Coenzyme Q10 Analog | . | D0MM6A | |||
Investigative
References
