Details of the Target
General Information of Target
| Target ID | LDTP05875 | |||||
|---|---|---|---|---|---|---|
| Target Name | Transmembrane emp24 domain-containing protein 1 (TMED1) | |||||
| Gene Name | TMED1 | |||||
| Gene ID | 11018 | |||||
| Synonyms |
IL1RL1L; IL1RL1LG; Transmembrane emp24 domain-containing protein 1; Interleukin-1 receptor-like 1 ligand; Putative T1/ST2 receptor-binding protein; p24 family protein gamma-1; Tp24; p24gamma1 |
|||||
| 3D Structure | ||||||
| Sequence |
MMAAGAALALALWLLMPPVEVGGAGPPPIQDGEFTFLLPAGRKQCFYQSAPANASLETEY
QVIGGAGLDVDFTLESPQGVLLVSESRKADGVHTVEPTEAGDYKLCFDNSFSTISEKLVF FELIFDSLQDDEEVEGWAEAVEPEEMLDVKMEDIKESIETMRTRLERSIQMLTLLRAFEA RDRNLQEGNLERVNFWSAVNVAVLLLVAVLQVCTLKRFFQDKRPVPT |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
EMP24/GP25L family
|
|||||
| Subcellular location |
Cell membrane
|
|||||
| Function |
Potential role in vesicular protein trafficking, mainly in the early secretory pathway. May act as a cargo receptor at the lumenal side for incorporation of secretory cargo molecules into transport vesicles and may be involved in vesicle coat formation at the cytoplasmic side. Plays a positive role in IL-33-mediated IL-8 and IL-6 production by interacting with interleukin-33 receptor IL1RL1. Also plays a role in the modulation of innate immune signaling through the cGAS-STING pathway by interacting with RNF26.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FBPP2 Probe Info |
 |
2.67 | LDD0318 | [1] | |
|
CHEMBL5175495 Probe Info |
 |
9.98 | LDD0196 | [2] | |
|
TG42 Probe Info |
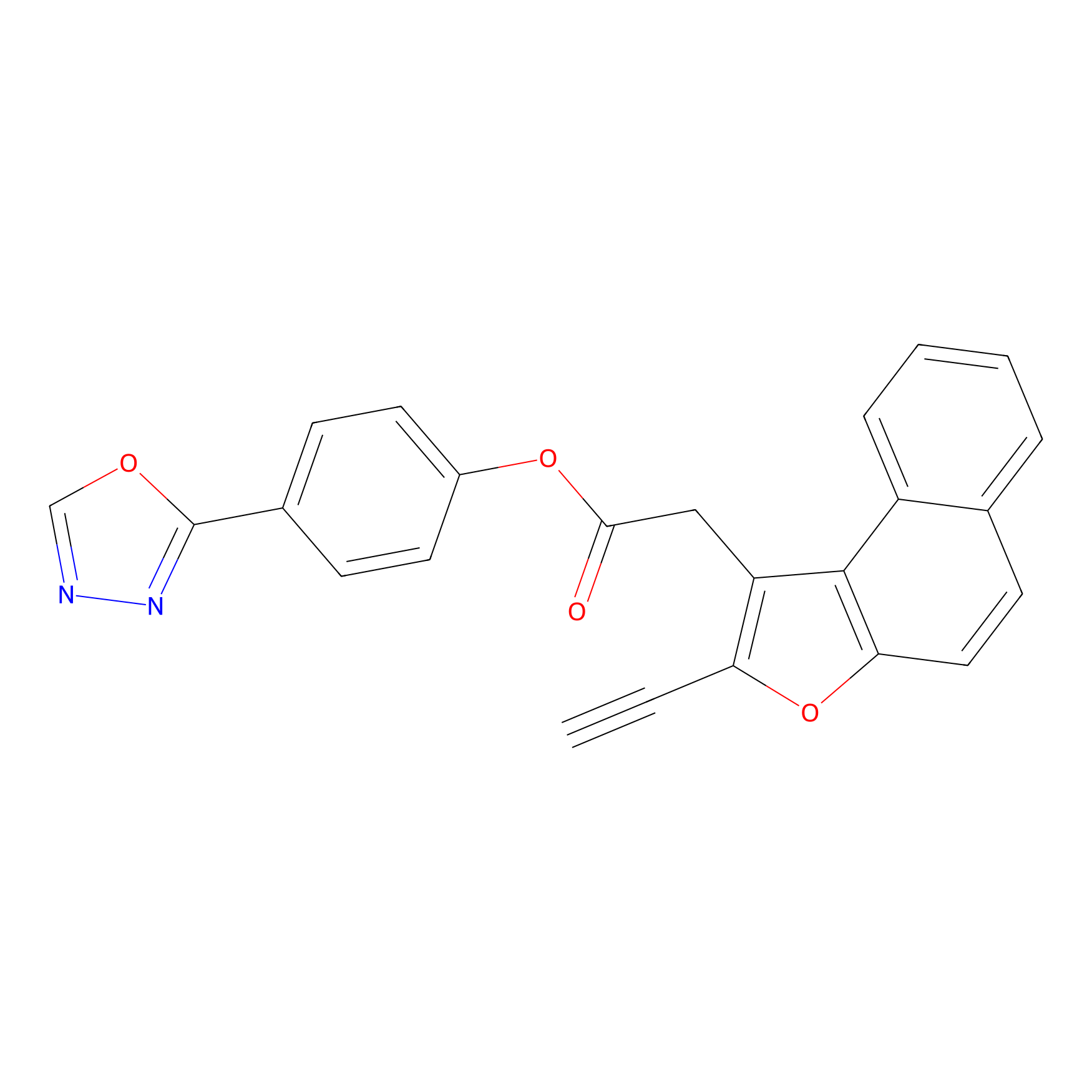 |
24.47 | LDD0326 | [3] | |
|
C-Sul Probe Info |
 |
3.27 | LDD0066 | [4] | |
|
FBP2 Probe Info |
 |
3.53 | LDD0317 | [1] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [5] | |
|
DBIA Probe Info |
 |
C106(1.68) | LDD3312 | [6] | |
|
Curcusone 37 Probe Info |
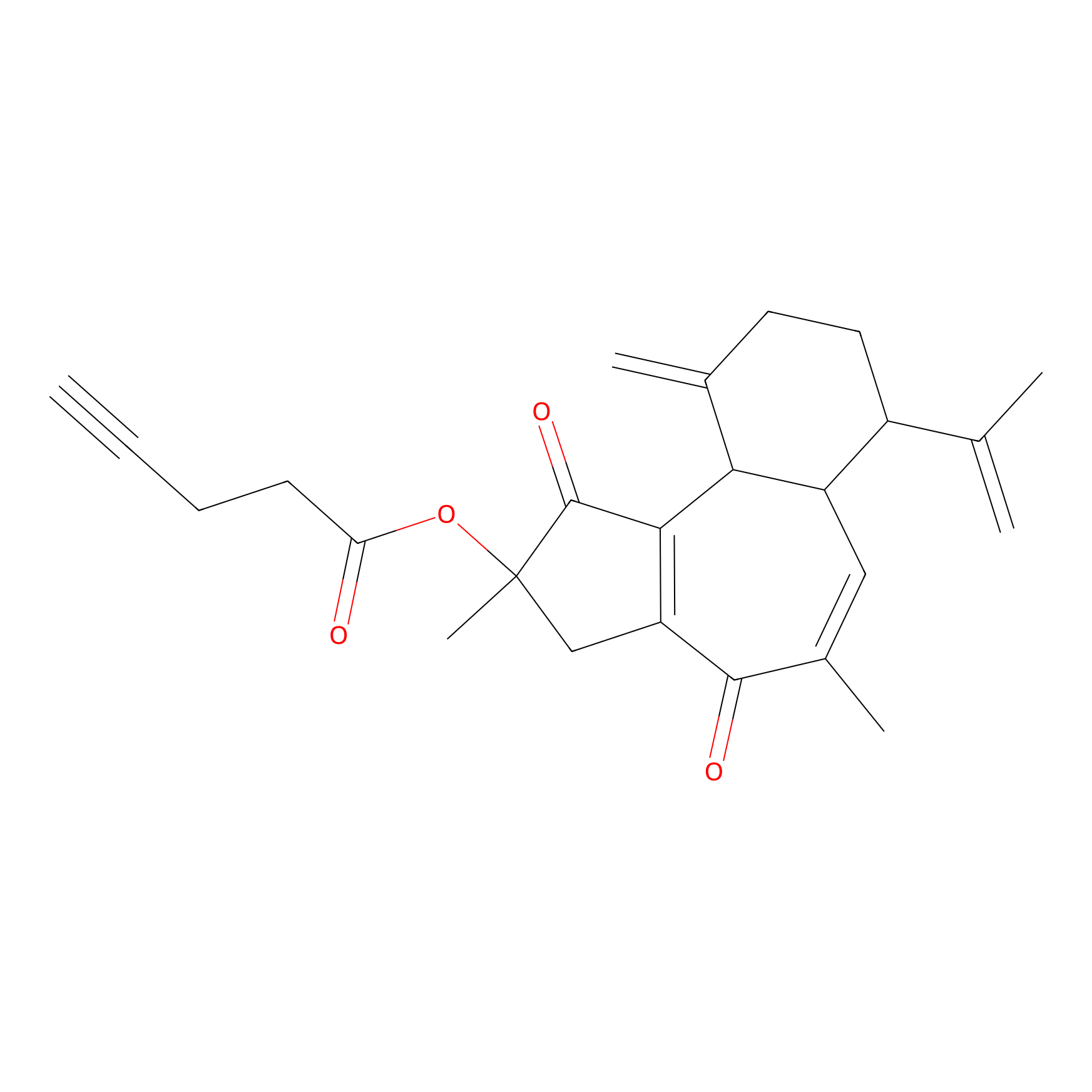 |
5.44 | LDD0188 | [7] | |
|
Johansson_61 Probe Info |
 |
_(11.65) | LDD1485 | [8] | |
|
HPAP Probe Info |
 |
4.68 | LDD0062 | [9] | |
|
Alkyne-RA190 Probe Info |
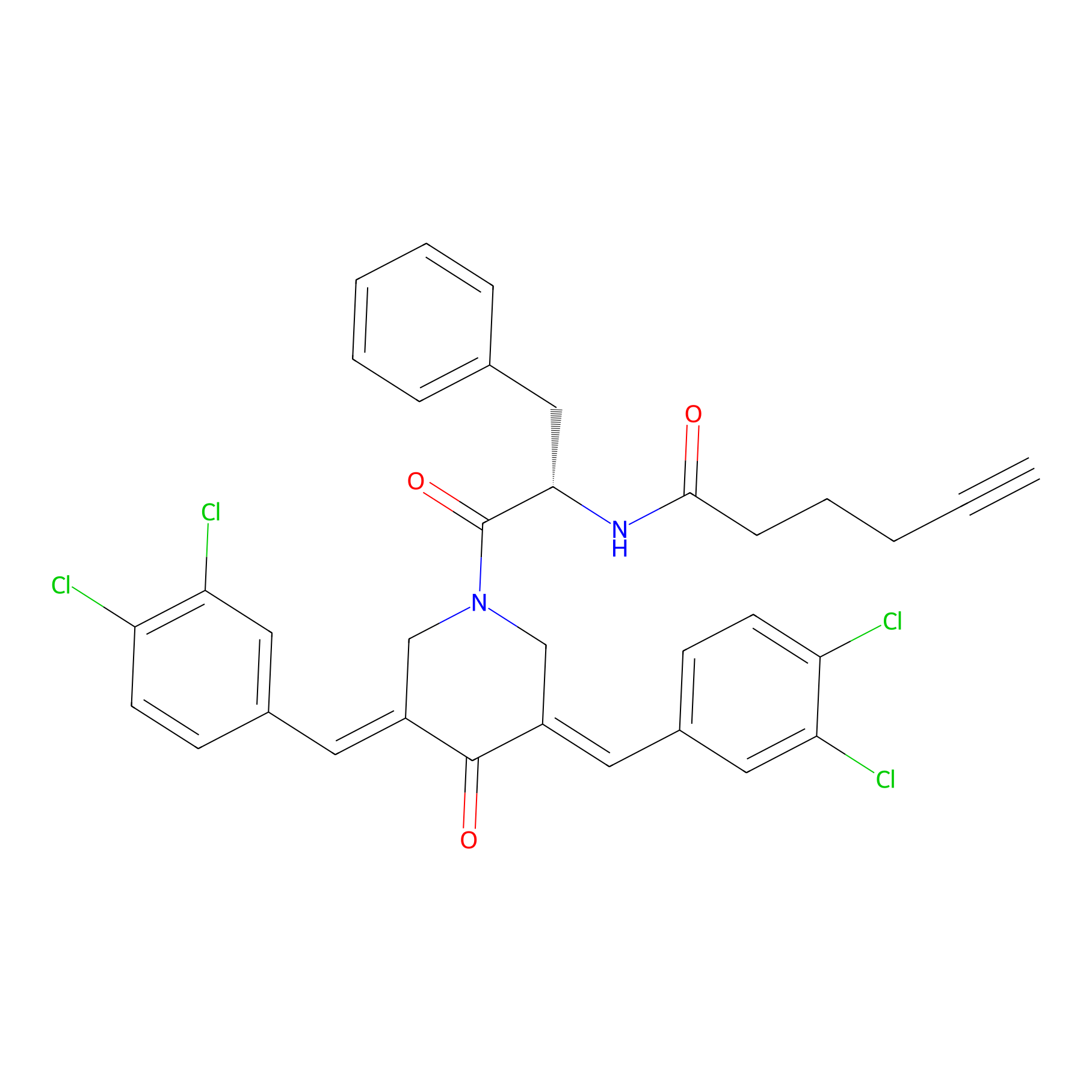 |
2.56 | LDD0303 | [10] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [11] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0162 | [12] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [11] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [13] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C299 Probe Info |
 |
17.27 | LDD1968 | [14] | |
|
C305 Probe Info |
 |
11.55 | LDD1974 | [14] | |
|
C362 Probe Info |
 |
34.30 | LDD2023 | [14] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0472 | [15] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0465 | [15] | |
|
OEA-DA Probe Info |
 |
6.69 | LDD0046 | [16] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0033 | Curcusone1d | MCF-7 | 5.44 | LDD0188 | [7] |
| LDCM0617 | Fragment63-S | Jurkat | _(20.00) | LDD1490 | [8] |
| LDCM0569 | Fragment7 | Jurkat | _(11.65) | LDD1485 | [8] |
| LDCM0022 | KB02 | HEK-293T | C106(0.97) | LDD1492 | [17] |
| LDCM0023 | KB03 | HEK-293T | C106(0.91) | LDD1497 | [17] |
| LDCM0024 | KB05 | HMCB | C106(1.68) | LDD3312 | [6] |
| LDCM0014 | Panhematin | HEK-293T | 4.68 | LDD0062 | [9] |
| LDCM0131 | RA190 | SK-MEL-5 | 2.56 | LDD0303 | [10] |
References
