Details of the Target
General Information of Target
| Target ID | LDTP00361 | |||||
|---|---|---|---|---|---|---|
| Target Name | Nuclear envelope integral membrane protein 1 (NEMP1) | |||||
| Gene Name | NEMP1 | |||||
| Gene ID | 23306 | |||||
| Synonyms |
KIAA0286; TMEM194; TMEM194A; Nuclear envelope integral membrane protein 1 |
|||||
| 3D Structure | ||||||
| Sequence |
MAGGMKVAVSPAVGPGPWGSGVGGGGTVRLLLILSGCLVYGTAETDVNVVMLQESQVCEK
RASQQFCYTNVLIPKWHDIWTRIQIRVNSSRLVRVTQVENEEKLKELEQFSIWNFFSSFL KEKLNDTYVNVGLYSTKTCLKVEIIEKDTKYSVIVIRRFDPKLFLVFLLGLMLFFCGDLL SRSQIFYYSTGMTVGIVASLLIIIFILSKFMPKKSPIYVILVGGWSFSLYLIQLVFKNLQ EIWRCYWQYLLSYVLTVGFMSFAVCYKYGPLENERSINLLTWTLQLMGLCFMYSGIQIPH IALAIIIIALCTKNLEHPIQWLYITCRKVCKGAEKPVPPRLLTEEEYRIQGEVETRKALE ELREFCNSPDCSAWKTVSRIQSPKRFADFVEGSSHLTPNEVSVHEQEYGLGSIIAQDEIY EEASSEEEDSYSRCPAITQNNFLT |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
NEMP family
|
|||||
| Subcellular location |
Nucleus inner membrane
|
|||||
| Function |
Together with EMD, contributes to nuclear envelope stiffness in germ cells. Required for female fertility. Essential for normal erythropoiesis. Required for efficient nuclear envelope opening and enucleation during the late stages of erythroblast maturation.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| MOLT4 | SNV: p.G3V | . | |||
| NCIH23 | SNV: p.V98L | . | |||
| SNGM | SNV: p.T397M | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
10.76 | LDD0402 | [1] | |
|
HDSF-alk Probe Info |
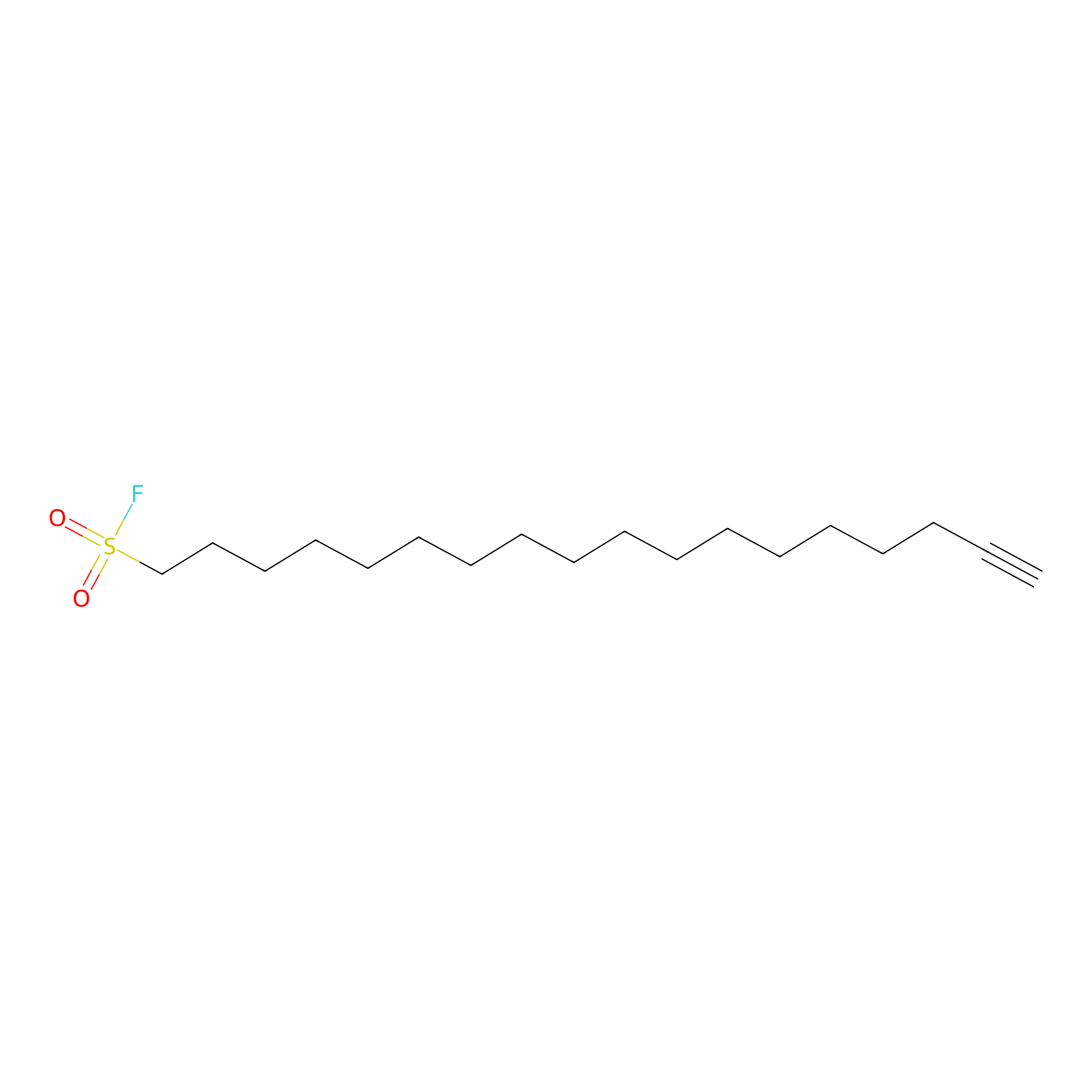 |
2.95 | LDD0197 | [2] | |
|
DBIA Probe Info |
 |
C366(1.05); C371(1.05) | LDD0078 | [3] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C290(0.00); C311(0.00) | LDD0038 | [4] | |
|
IA-alkyne Probe Info |
 |
C67(0.00); C326(0.00) | LDD0165 | [5] | |
|
TFBX Probe Info |
 |
C434(0.00); C330(0.00) | LDD0027 | [6] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [7] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [8] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DR-1 Probe Info |
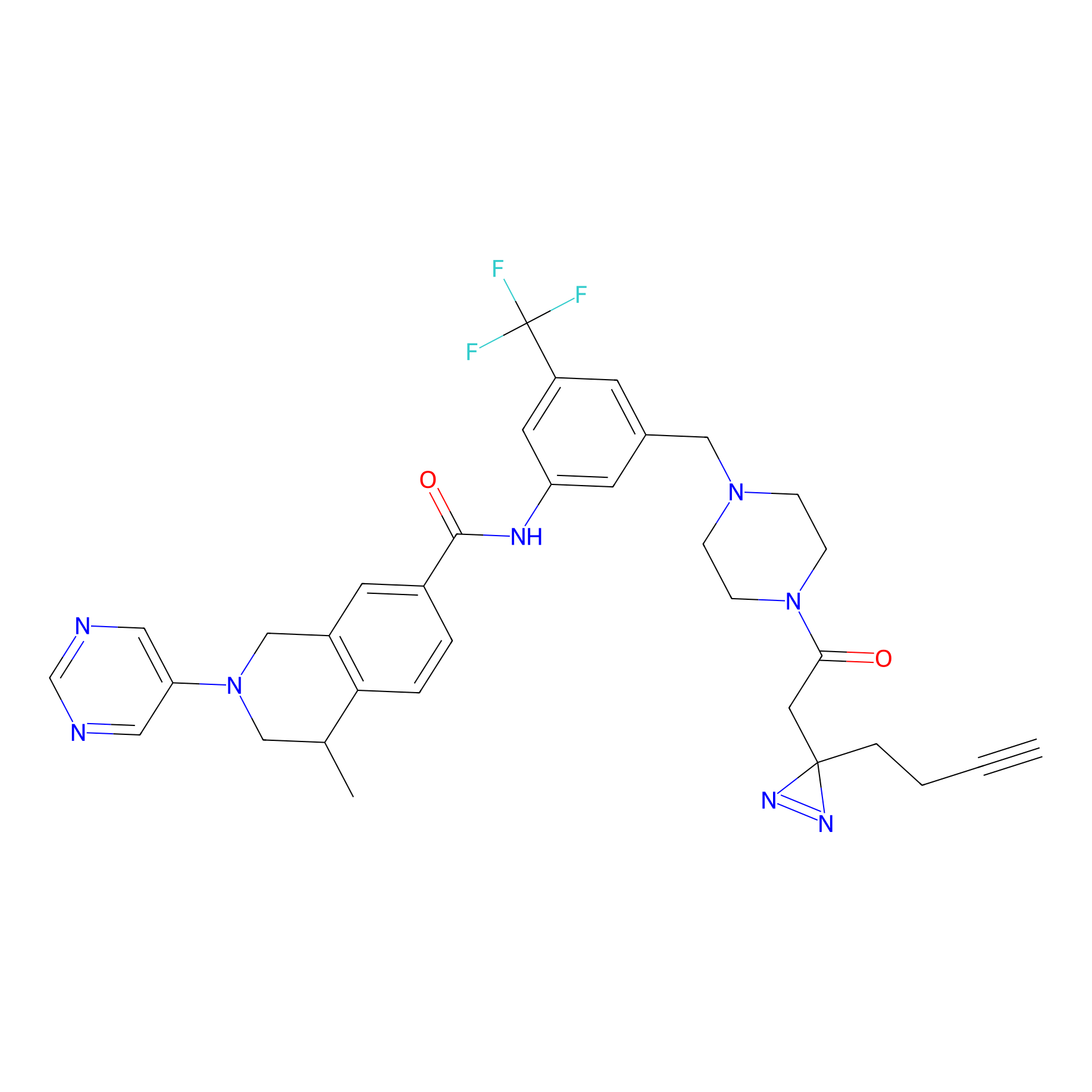 |
2.51 | LDD0398 | [9] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [10] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [10] | |
|
FFF probe14 Probe Info |
 |
20.00 | LDD0477 | [10] | |
|
OEA-DA Probe Info |
 |
10.38 | LDD0046 | [11] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HCT 116 | C434(0.96) | LDD0531 | [3] |
| LDCM0215 | AC10 | HCT 116 | C371(0.94); C434(1.15) | LDD0532 | [3] |
| LDCM0216 | AC100 | HCT 116 | C366(1.00); C371(0.92); C434(0.77) | LDD0533 | [3] |
| LDCM0217 | AC101 | HCT 116 | C366(0.97); C371(1.07); C434(1.02) | LDD0534 | [3] |
| LDCM0218 | AC102 | HCT 116 | C366(1.06); C371(1.01); C434(1.05) | LDD0535 | [3] |
| LDCM0219 | AC103 | HCT 116 | C366(1.16); C371(1.14); C434(1.11) | LDD0536 | [3] |
| LDCM0220 | AC104 | HCT 116 | C366(1.13); C371(1.17); C434(0.91) | LDD0537 | [3] |
| LDCM0221 | AC105 | HCT 116 | C366(1.14); C371(1.08); C434(0.94) | LDD0538 | [3] |
| LDCM0222 | AC106 | HCT 116 | C366(1.19); C371(1.25); C434(1.00) | LDD0539 | [3] |
| LDCM0223 | AC107 | HCT 116 | C366(1.27); C371(1.33); C434(0.82) | LDD0540 | [3] |
| LDCM0224 | AC108 | HCT 116 | C366(0.91); C371(0.99); C434(0.97) | LDD0541 | [3] |
| LDCM0225 | AC109 | HCT 116 | C366(0.90); C371(1.10); C434(0.86) | LDD0542 | [3] |
| LDCM0226 | AC11 | HCT 116 | C371(0.93); C434(0.99) | LDD0543 | [3] |
| LDCM0227 | AC110 | HCT 116 | C366(0.88); C371(0.99); C434(0.85) | LDD0544 | [3] |
| LDCM0228 | AC111 | HCT 116 | C366(1.03); C371(1.40); C434(1.16) | LDD0545 | [3] |
| LDCM0229 | AC112 | HCT 116 | C366(1.12); C371(1.37); C434(0.92) | LDD0546 | [3] |
| LDCM0230 | AC113 | HCT 116 | C371(1.12); C434(0.89) | LDD0547 | [3] |
| LDCM0231 | AC114 | HCT 116 | C371(1.26); C434(0.92) | LDD0548 | [3] |
| LDCM0232 | AC115 | HCT 116 | C371(1.19); C434(1.16) | LDD0549 | [3] |
| LDCM0233 | AC116 | HCT 116 | C371(1.10); C434(1.02) | LDD0550 | [3] |
| LDCM0234 | AC117 | HCT 116 | C371(0.97); C434(0.96) | LDD0551 | [3] |
| LDCM0235 | AC118 | HCT 116 | C371(1.13); C434(0.87) | LDD0552 | [3] |
| LDCM0236 | AC119 | HCT 116 | C371(1.16); C434(0.89) | LDD0553 | [3] |
| LDCM0237 | AC12 | HCT 116 | C371(0.91); C434(0.97) | LDD0554 | [3] |
| LDCM0238 | AC120 | HCT 116 | C371(1.26); C434(1.26) | LDD0555 | [3] |
| LDCM0239 | AC121 | HCT 116 | C371(0.87); C434(1.02) | LDD0556 | [3] |
| LDCM0240 | AC122 | HCT 116 | C371(1.23); C434(1.01) | LDD0557 | [3] |
| LDCM0241 | AC123 | HCT 116 | C371(0.93); C434(0.72) | LDD0558 | [3] |
| LDCM0242 | AC124 | HCT 116 | C371(1.06); C434(1.00) | LDD0559 | [3] |
| LDCM0243 | AC125 | HCT 116 | C371(1.18); C434(1.14) | LDD0560 | [3] |
| LDCM0244 | AC126 | HCT 116 | C371(1.11); C434(1.06) | LDD0561 | [3] |
| LDCM0245 | AC127 | HCT 116 | C371(1.07); C434(0.87) | LDD0562 | [3] |
| LDCM0246 | AC128 | HCT 116 | C434(1.04) | LDD0563 | [3] |
| LDCM0247 | AC129 | HCT 116 | C434(1.14) | LDD0564 | [3] |
| LDCM0249 | AC130 | HCT 116 | C434(1.23) | LDD0566 | [3] |
| LDCM0250 | AC131 | HCT 116 | C434(1.14) | LDD0567 | [3] |
| LDCM0251 | AC132 | HCT 116 | C434(1.20) | LDD0568 | [3] |
| LDCM0252 | AC133 | HCT 116 | C434(1.14) | LDD0569 | [3] |
| LDCM0253 | AC134 | HCT 116 | C434(1.23) | LDD0570 | [3] |
| LDCM0254 | AC135 | HCT 116 | C434(1.23) | LDD0571 | [3] |
| LDCM0255 | AC136 | HCT 116 | C434(1.27) | LDD0572 | [3] |
| LDCM0256 | AC137 | HCT 116 | C434(1.09) | LDD0573 | [3] |
| LDCM0257 | AC138 | HCT 116 | C434(1.23) | LDD0574 | [3] |
| LDCM0258 | AC139 | HCT 116 | C434(1.67) | LDD0575 | [3] |
| LDCM0259 | AC14 | HCT 116 | C371(0.79); C434(1.00) | LDD0576 | [3] |
| LDCM0260 | AC140 | HCT 116 | C434(1.50) | LDD0577 | [3] |
| LDCM0261 | AC141 | HCT 116 | C434(1.48) | LDD0578 | [3] |
| LDCM0262 | AC142 | HCT 116 | C434(1.27) | LDD0579 | [3] |
| LDCM0263 | AC143 | HCT 116 | C371(1.19) | LDD0580 | [3] |
| LDCM0264 | AC144 | HCT 116 | C371(1.26) | LDD0581 | [3] |
| LDCM0265 | AC145 | HCT 116 | C371(1.03) | LDD0582 | [3] |
| LDCM0266 | AC146 | HCT 116 | C371(1.15) | LDD0583 | [3] |
| LDCM0267 | AC147 | HCT 116 | C371(1.03) | LDD0584 | [3] |
| LDCM0268 | AC148 | HCT 116 | C371(1.07) | LDD0585 | [3] |
| LDCM0269 | AC149 | HCT 116 | C371(1.22) | LDD0586 | [3] |
| LDCM0270 | AC15 | HCT 116 | C371(0.84); C434(1.03) | LDD0587 | [3] |
| LDCM0271 | AC150 | HCT 116 | C371(1.20) | LDD0588 | [3] |
| LDCM0272 | AC151 | HCT 116 | C371(1.22) | LDD0589 | [3] |
| LDCM0273 | AC152 | HCT 116 | C371(1.17) | LDD0590 | [3] |
| LDCM0274 | AC153 | HCT 116 | C371(1.41) | LDD0591 | [3] |
| LDCM0621 | AC154 | HCT 116 | C371(1.24) | LDD2158 | [3] |
| LDCM0622 | AC155 | HCT 116 | C371(1.23) | LDD2159 | [3] |
| LDCM0623 | AC156 | HCT 116 | C371(1.26) | LDD2160 | [3] |
| LDCM0624 | AC157 | HCT 116 | C371(1.29) | LDD2161 | [3] |
| LDCM0276 | AC17 | HCT 116 | C434(1.12); C67(1.13) | LDD0593 | [3] |
| LDCM0277 | AC18 | HCT 116 | C434(0.80); C67(1.14) | LDD0594 | [3] |
| LDCM0278 | AC19 | HCT 116 | C434(0.84); C67(1.27) | LDD0595 | [3] |
| LDCM0279 | AC2 | HCT 116 | C434(1.15) | LDD0596 | [3] |
| LDCM0280 | AC20 | HCT 116 | C67(0.97); C434(1.08) | LDD0597 | [3] |
| LDCM0281 | AC21 | HCT 116 | C434(0.95); C67(1.02) | LDD0598 | [3] |
| LDCM0282 | AC22 | HCT 116 | C434(0.93); C67(0.98) | LDD0599 | [3] |
| LDCM0283 | AC23 | HCT 116 | C434(0.94); C67(1.26) | LDD0600 | [3] |
| LDCM0284 | AC24 | HCT 116 | C434(1.01); C67(1.14) | LDD0601 | [3] |
| LDCM0285 | AC25 | HCT 116 | C434(0.99) | LDD0602 | [3] |
| LDCM0286 | AC26 | HCT 116 | C434(1.52) | LDD0603 | [3] |
| LDCM0287 | AC27 | HCT 116 | C434(1.35) | LDD0604 | [3] |
| LDCM0288 | AC28 | HCT 116 | C434(1.46) | LDD0605 | [3] |
| LDCM0289 | AC29 | HCT 116 | C434(1.12) | LDD0606 | [3] |
| LDCM0290 | AC3 | HCT 116 | C434(1.07) | LDD0607 | [3] |
| LDCM0291 | AC30 | HCT 116 | C434(1.22) | LDD0608 | [3] |
| LDCM0292 | AC31 | HCT 116 | C434(1.41) | LDD0609 | [3] |
| LDCM0293 | AC32 | HCT 116 | C434(1.83) | LDD0610 | [3] |
| LDCM0294 | AC33 | HCT 116 | C434(1.37) | LDD0611 | [3] |
| LDCM0295 | AC34 | HCT 116 | C434(1.40) | LDD0612 | [3] |
| LDCM0296 | AC35 | HCT 116 | C434(0.79) | LDD0613 | [3] |
| LDCM0297 | AC36 | HCT 116 | C434(0.87) | LDD0614 | [3] |
| LDCM0298 | AC37 | HCT 116 | C434(0.99) | LDD0615 | [3] |
| LDCM0299 | AC38 | HCT 116 | C434(1.01) | LDD0616 | [3] |
| LDCM0300 | AC39 | HCT 116 | C434(1.16) | LDD0617 | [3] |
| LDCM0301 | AC4 | HCT 116 | C434(1.06) | LDD0618 | [3] |
| LDCM0302 | AC40 | HCT 116 | C434(1.08) | LDD0619 | [3] |
| LDCM0303 | AC41 | HCT 116 | C434(1.00) | LDD0620 | [3] |
| LDCM0304 | AC42 | HCT 116 | C434(1.13) | LDD0621 | [3] |
| LDCM0305 | AC43 | HCT 116 | C434(1.09) | LDD0622 | [3] |
| LDCM0306 | AC44 | HCT 116 | C434(1.14) | LDD0623 | [3] |
| LDCM0307 | AC45 | HCT 116 | C434(1.24) | LDD0624 | [3] |
| LDCM0308 | AC46 | HCT 116 | C434(1.16) | LDD0625 | [3] |
| LDCM0309 | AC47 | HCT 116 | C434(1.22) | LDD0626 | [3] |
| LDCM0310 | AC48 | HCT 116 | C434(1.25) | LDD0627 | [3] |
| LDCM0311 | AC49 | HCT 116 | C434(1.19) | LDD0628 | [3] |
| LDCM0312 | AC5 | HCT 116 | C434(1.01) | LDD0629 | [3] |
| LDCM0313 | AC50 | HCT 116 | C434(1.30) | LDD0630 | [3] |
| LDCM0314 | AC51 | HCT 116 | C434(1.18) | LDD0631 | [3] |
| LDCM0315 | AC52 | HCT 116 | C434(1.13) | LDD0632 | [3] |
| LDCM0316 | AC53 | HCT 116 | C434(1.03) | LDD0633 | [3] |
| LDCM0317 | AC54 | HCT 116 | C434(1.19) | LDD0634 | [3] |
| LDCM0318 | AC55 | HCT 116 | C434(1.10) | LDD0635 | [3] |
| LDCM0319 | AC56 | HCT 116 | C434(1.10) | LDD0636 | [3] |
| LDCM0320 | AC57 | HCT 116 | C67(0.95); C366(1.01); C371(1.01); C434(1.27) | LDD0637 | [3] |
| LDCM0321 | AC58 | HCT 116 | C366(0.97); C371(0.97); C67(1.10); C434(1.60) | LDD0638 | [3] |
| LDCM0322 | AC59 | HCT 116 | C67(0.82); C366(1.01); C371(1.01); C434(1.33) | LDD0639 | [3] |
| LDCM0323 | AC6 | HCT 116 | C371(0.90); C434(1.21) | LDD0640 | [3] |
| LDCM0324 | AC60 | HCT 116 | C366(0.85); C371(0.85); C67(0.88); C434(1.39) | LDD0641 | [3] |
| LDCM0325 | AC61 | HCT 116 | C366(0.74); C371(0.74); C67(0.99); C434(1.15) | LDD0642 | [3] |
| LDCM0326 | AC62 | HCT 116 | C366(1.01); C371(1.01); C67(1.03); C434(1.45) | LDD0643 | [3] |
| LDCM0327 | AC63 | HCT 116 | C67(0.87); C366(0.89); C371(0.89); C434(1.31) | LDD0644 | [3] |
| LDCM0328 | AC64 | HCT 116 | C366(0.85); C371(0.85); C67(1.07); C434(1.33) | LDD0645 | [3] |
| LDCM0329 | AC65 | HCT 116 | C366(0.63); C371(0.63); C67(1.02); C434(1.32) | LDD0646 | [3] |
| LDCM0330 | AC66 | HCT 116 | C366(0.65); C371(0.65); C67(0.86); C434(1.18) | LDD0647 | [3] |
| LDCM0331 | AC67 | HCT 116 | C366(0.61); C371(0.61); C67(0.74); C434(1.33) | LDD0648 | [3] |
| LDCM0332 | AC68 | HCT 116 | C371(0.79) | LDD0649 | [3] |
| LDCM0333 | AC69 | HCT 116 | C371(0.81) | LDD0650 | [3] |
| LDCM0334 | AC7 | HCT 116 | C371(0.92); C434(1.02) | LDD0651 | [3] |
| LDCM0335 | AC70 | HCT 116 | C371(1.01) | LDD0652 | [3] |
| LDCM0336 | AC71 | HCT 116 | C371(0.84) | LDD0653 | [3] |
| LDCM0337 | AC72 | HCT 116 | C371(1.02) | LDD0654 | [3] |
| LDCM0338 | AC73 | HCT 116 | C371(1.10) | LDD0655 | [3] |
| LDCM0339 | AC74 | HCT 116 | C371(0.82) | LDD0656 | [3] |
| LDCM0340 | AC75 | HCT 116 | C371(1.12) | LDD0657 | [3] |
| LDCM0341 | AC76 | HCT 116 | C371(1.20) | LDD0658 | [3] |
| LDCM0342 | AC77 | HCT 116 | C371(0.80) | LDD0659 | [3] |
| LDCM0343 | AC78 | HCT 116 | C371(0.91) | LDD0660 | [3] |
| LDCM0344 | AC79 | HCT 116 | C371(1.25) | LDD0661 | [3] |
| LDCM0345 | AC8 | HCT 116 | C371(0.93); C434(1.03) | LDD0662 | [3] |
| LDCM0346 | AC80 | HCT 116 | C371(1.05) | LDD0663 | [3] |
| LDCM0347 | AC81 | HCT 116 | C371(0.81) | LDD0664 | [3] |
| LDCM0348 | AC82 | HCT 116 | C371(1.02) | LDD0665 | [3] |
| LDCM0349 | AC83 | HCT 116 | C371(1.56) | LDD0666 | [3] |
| LDCM0350 | AC84 | HCT 116 | C371(1.28) | LDD0667 | [3] |
| LDCM0351 | AC85 | HCT 116 | C371(1.33) | LDD0668 | [3] |
| LDCM0352 | AC86 | HCT 116 | C371(1.32) | LDD0669 | [3] |
| LDCM0353 | AC87 | HCT 116 | C371(1.18) | LDD0670 | [3] |
| LDCM0354 | AC88 | HCT 116 | C371(1.25) | LDD0671 | [3] |
| LDCM0355 | AC89 | HCT 116 | C371(1.10) | LDD0672 | [3] |
| LDCM0357 | AC90 | HCT 116 | C371(1.14) | LDD0674 | [3] |
| LDCM0358 | AC91 | HCT 116 | C371(1.42) | LDD0675 | [3] |
| LDCM0359 | AC92 | HCT 116 | C371(1.39) | LDD0676 | [3] |
| LDCM0360 | AC93 | HCT 116 | C371(1.20) | LDD0677 | [3] |
| LDCM0361 | AC94 | HCT 116 | C371(1.14) | LDD0678 | [3] |
| LDCM0362 | AC95 | HCT 116 | C371(1.35) | LDD0679 | [3] |
| LDCM0363 | AC96 | HCT 116 | C371(1.07) | LDD0680 | [3] |
| LDCM0364 | AC97 | HCT 116 | C371(1.31) | LDD0681 | [3] |
| LDCM0365 | AC98 | HCT 116 | C434(1.00); C371(1.13); C366(1.25) | LDD0682 | [3] |
| LDCM0366 | AC99 | HCT 116 | C371(0.88); C366(0.93); C434(0.99) | LDD0683 | [3] |
| LDCM0248 | AKOS034007472 | HCT 116 | C371(0.92); C434(1.02) | LDD0565 | [3] |
| LDCM0356 | AKOS034007680 | HCT 116 | C434(0.93); C371(1.11) | LDD0673 | [3] |
| LDCM0275 | AKOS034007705 | HCT 116 | C371(0.91); C434(1.60) | LDD0592 | [3] |
| LDCM0156 | Aniline | NCI-H1299 | 13.87 | LDD0403 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C366(1.05); C371(1.05) | LDD0078 | [3] |
| LDCM0367 | CL1 | HCT 116 | C434(0.95); C67(1.03) | LDD0684 | [3] |
| LDCM0368 | CL10 | HCT 116 | C67(0.62); C434(1.12) | LDD0685 | [3] |
| LDCM0369 | CL100 | HCT 116 | C434(1.00) | LDD0686 | [3] |
| LDCM0370 | CL101 | HCT 116 | C371(0.69); C434(1.27) | LDD0687 | [3] |
| LDCM0371 | CL102 | HCT 116 | C371(0.91); C434(0.98) | LDD0688 | [3] |
| LDCM0372 | CL103 | HCT 116 | C371(0.96); C434(1.01) | LDD0689 | [3] |
| LDCM0373 | CL104 | HCT 116 | C371(0.98); C434(1.10) | LDD0690 | [3] |
| LDCM0374 | CL105 | HCT 116 | C434(0.98); C67(1.17) | LDD0691 | [3] |
| LDCM0375 | CL106 | HCT 116 | C434(0.92); C67(0.99) | LDD0692 | [3] |
| LDCM0376 | CL107 | HCT 116 | C434(0.91); C67(1.03) | LDD0693 | [3] |
| LDCM0377 | CL108 | HCT 116 | C434(0.97); C67(1.15) | LDD0694 | [3] |
| LDCM0378 | CL109 | HCT 116 | C434(0.89); C67(1.14) | LDD0695 | [3] |
| LDCM0379 | CL11 | HCT 116 | C434(0.99); C67(1.02) | LDD0696 | [3] |
| LDCM0380 | CL110 | HCT 116 | C434(1.00); C67(1.13) | LDD0697 | [3] |
| LDCM0381 | CL111 | HCT 116 | C67(0.90); C434(1.12) | LDD0698 | [3] |
| LDCM0382 | CL112 | HCT 116 | C434(1.10) | LDD0699 | [3] |
| LDCM0383 | CL113 | HCT 116 | C434(1.32) | LDD0700 | [3] |
| LDCM0384 | CL114 | HCT 116 | C434(1.21) | LDD0701 | [3] |
| LDCM0385 | CL115 | HCT 116 | C434(1.28) | LDD0702 | [3] |
| LDCM0386 | CL116 | HCT 116 | C434(1.45) | LDD0703 | [3] |
| LDCM0387 | CL117 | HCT 116 | C434(1.58) | LDD0704 | [3] |
| LDCM0388 | CL118 | HCT 116 | C434(1.10) | LDD0705 | [3] |
| LDCM0389 | CL119 | HCT 116 | C434(1.14) | LDD0706 | [3] |
| LDCM0390 | CL12 | HCT 116 | C434(1.06); C67(1.19) | LDD0707 | [3] |
| LDCM0391 | CL120 | HCT 116 | C434(1.08) | LDD0708 | [3] |
| LDCM0392 | CL121 | HCT 116 | C434(1.11) | LDD0709 | [3] |
| LDCM0393 | CL122 | HCT 116 | C434(1.14) | LDD0710 | [3] |
| LDCM0394 | CL123 | HCT 116 | C434(1.15) | LDD0711 | [3] |
| LDCM0395 | CL124 | HCT 116 | C434(1.16) | LDD0712 | [3] |
| LDCM0396 | CL125 | HCT 116 | C366(0.82); C371(0.82); C67(0.92); C434(0.98) | LDD0713 | [3] |
| LDCM0397 | CL126 | HCT 116 | C366(0.90); C371(0.90); C67(1.00); C434(1.02) | LDD0714 | [3] |
| LDCM0398 | CL127 | HCT 116 | C366(0.75); C371(0.75); C67(0.93); C434(1.10) | LDD0715 | [3] |
| LDCM0399 | CL128 | HCT 116 | C366(1.07); C371(1.07); C434(1.16); C67(1.25) | LDD0716 | [3] |
| LDCM0400 | CL13 | HCT 116 | C434(0.96); C67(1.10) | LDD0717 | [3] |
| LDCM0401 | CL14 | HCT 116 | C434(1.00); C67(1.12) | LDD0718 | [3] |
| LDCM0402 | CL15 | HCT 116 | C434(0.99); C67(1.22) | LDD0719 | [3] |
| LDCM0403 | CL16 | HCT 116 | C371(1.03); C434(1.21) | LDD0720 | [3] |
| LDCM0404 | CL17 | HCT 116 | C371(0.86) | LDD0721 | [3] |
| LDCM0405 | CL18 | HCT 116 | C371(0.80) | LDD0722 | [3] |
| LDCM0406 | CL19 | HCT 116 | C371(0.74) | LDD0723 | [3] |
| LDCM0407 | CL2 | HCT 116 | C434(1.06); C67(1.36) | LDD0724 | [3] |
| LDCM0408 | CL20 | HCT 116 | C371(0.84) | LDD0725 | [3] |
| LDCM0409 | CL21 | HCT 116 | C371(0.70) | LDD0726 | [3] |
| LDCM0410 | CL22 | HCT 116 | C371(0.94) | LDD0727 | [3] |
| LDCM0411 | CL23 | HCT 116 | C371(0.80) | LDD0728 | [3] |
| LDCM0412 | CL24 | HCT 116 | C371(0.82) | LDD0729 | [3] |
| LDCM0413 | CL25 | HCT 116 | C371(0.82) | LDD0730 | [3] |
| LDCM0414 | CL26 | HCT 116 | C371(1.11) | LDD0731 | [3] |
| LDCM0415 | CL27 | HCT 116 | C371(0.79) | LDD0732 | [3] |
| LDCM0416 | CL28 | HCT 116 | C371(0.88) | LDD0733 | [3] |
| LDCM0417 | CL29 | HCT 116 | C371(0.75) | LDD0734 | [3] |
| LDCM0418 | CL3 | HCT 116 | C434(0.96); C67(1.05) | LDD0735 | [3] |
| LDCM0419 | CL30 | HCT 116 | C371(0.89) | LDD0736 | [3] |
| LDCM0420 | CL31 | HCT 116 | C434(0.79) | LDD0737 | [3] |
| LDCM0421 | CL32 | HCT 116 | C371(1.16); C434(0.51) | LDD0738 | [3] |
| LDCM0422 | CL33 | HCT 116 | C371(0.92); C434(1.30) | LDD0739 | [3] |
| LDCM0423 | CL34 | HCT 116 | C371(0.93); C434(1.41) | LDD0740 | [3] |
| LDCM0424 | CL35 | HCT 116 | C371(1.03); C434(0.93) | LDD0741 | [3] |
| LDCM0425 | CL36 | HCT 116 | C371(0.96); C434(1.31) | LDD0742 | [3] |
| LDCM0426 | CL37 | HCT 116 | C371(1.07); C434(1.29) | LDD0743 | [3] |
| LDCM0428 | CL39 | HCT 116 | C371(1.14); C434(1.12) | LDD0745 | [3] |
| LDCM0429 | CL4 | HCT 116 | C434(1.05); C67(1.18) | LDD0746 | [3] |
| LDCM0430 | CL40 | HCT 116 | C371(0.92); C434(1.04) | LDD0747 | [3] |
| LDCM0431 | CL41 | HCT 116 | C371(0.90); C434(1.29) | LDD0748 | [3] |
| LDCM0432 | CL42 | HCT 116 | C371(0.76); C434(1.56) | LDD0749 | [3] |
| LDCM0433 | CL43 | HCT 116 | C371(0.99); C434(1.22) | LDD0750 | [3] |
| LDCM0434 | CL44 | HCT 116 | C371(0.91); C434(1.21) | LDD0751 | [3] |
| LDCM0435 | CL45 | HCT 116 | C371(0.98); C434(1.30) | LDD0752 | [3] |
| LDCM0436 | CL46 | HCT 116 | C366(0.86); C371(0.91); C434(0.99) | LDD0753 | [3] |
| LDCM0437 | CL47 | HCT 116 | C366(0.91); C371(0.85); C434(0.92) | LDD0754 | [3] |
| LDCM0438 | CL48 | HCT 116 | C366(1.09); C371(1.03); C434(0.96) | LDD0755 | [3] |
| LDCM0439 | CL49 | HCT 116 | C366(1.13); C371(1.08); C434(0.99) | LDD0756 | [3] |
| LDCM0440 | CL5 | HCT 116 | C434(0.94); C67(1.30) | LDD0757 | [3] |
| LDCM0441 | CL50 | HCT 116 | C366(0.92); C371(0.93); C434(1.06) | LDD0758 | [3] |
| LDCM0442 | CL51 | HCT 116 | C366(1.11); C371(1.00); C434(1.03) | LDD0759 | [3] |
| LDCM0443 | CL52 | HCT 116 | C366(1.00); C371(0.98); C434(1.18) | LDD0760 | [3] |
| LDCM0444 | CL53 | HCT 116 | C366(1.09); C371(1.07); C434(0.99) | LDD0761 | [3] |
| LDCM0445 | CL54 | HCT 116 | C366(1.01); C371(0.94); C434(0.93) | LDD0762 | [3] |
| LDCM0446 | CL55 | HCT 116 | C366(1.08); C371(1.01); C434(0.89) | LDD0763 | [3] |
| LDCM0447 | CL56 | HCT 116 | C366(1.02); C371(0.91); C434(1.03) | LDD0764 | [3] |
| LDCM0448 | CL57 | HCT 116 | C366(0.97); C371(0.92); C434(1.17) | LDD0765 | [3] |
| LDCM0449 | CL58 | HCT 116 | C366(1.06); C371(0.97); C434(0.95) | LDD0766 | [3] |
| LDCM0450 | CL59 | HCT 116 | C366(1.09); C371(1.01); C434(1.00) | LDD0767 | [3] |
| LDCM0451 | CL6 | HCT 116 | C434(1.04); C67(1.37) | LDD0768 | [3] |
| LDCM0452 | CL60 | HCT 116 | C366(1.02); C371(0.99); C434(1.09) | LDD0769 | [3] |
| LDCM0453 | CL61 | HCT 116 | C366(0.79); C371(0.79); C434(1.29); C67(0.96) | LDD0770 | [3] |
| LDCM0454 | CL62 | HCT 116 | C366(1.07); C371(1.07); C434(1.05); C67(0.88) | LDD0771 | [3] |
| LDCM0455 | CL63 | HCT 116 | C366(1.00); C371(1.00); C434(1.32); C67(0.85) | LDD0772 | [3] |
| LDCM0456 | CL64 | HCT 116 | C366(0.79); C371(0.79); C434(1.26); C67(0.85) | LDD0773 | [3] |
| LDCM0457 | CL65 | HCT 116 | C366(0.84); C371(0.84); C434(1.08); C67(0.86) | LDD0774 | [3] |
| LDCM0458 | CL66 | HCT 116 | C366(0.92); C371(0.92); C434(1.38); C67(0.86) | LDD0775 | [3] |
| LDCM0459 | CL67 | HCT 116 | C366(0.72); C371(0.72); C434(1.29); C67(0.82) | LDD0776 | [3] |
| LDCM0460 | CL68 | HCT 116 | C366(0.74); C371(0.74); C434(1.27); C67(0.88) | LDD0777 | [3] |
| LDCM0461 | CL69 | HCT 116 | C366(0.96); C371(0.96); C434(1.05); C67(0.97) | LDD0778 | [3] |
| LDCM0462 | CL7 | HCT 116 | C434(1.00); C67(1.49) | LDD0779 | [3] |
| LDCM0463 | CL70 | HCT 116 | C366(0.89); C371(0.89); C434(1.23); C67(0.92) | LDD0780 | [3] |
| LDCM0464 | CL71 | HCT 116 | C366(0.85); C371(0.85); C434(1.00); C67(0.83) | LDD0781 | [3] |
| LDCM0465 | CL72 | HCT 116 | C366(0.71); C371(0.71); C434(1.23); C67(0.92) | LDD0782 | [3] |
| LDCM0466 | CL73 | HCT 116 | C366(0.82); C371(0.82); C434(1.20); C67(1.01) | LDD0783 | [3] |
| LDCM0467 | CL74 | HCT 116 | C366(0.89); C371(0.89); C434(1.25); C67(0.88) | LDD0784 | [3] |
| LDCM0469 | CL76 | HCT 116 | C434(1.19) | LDD0786 | [3] |
| LDCM0470 | CL77 | HCT 116 | C434(1.10) | LDD0787 | [3] |
| LDCM0471 | CL78 | HCT 116 | C434(1.08) | LDD0788 | [3] |
| LDCM0472 | CL79 | HCT 116 | C434(1.35) | LDD0789 | [3] |
| LDCM0473 | CL8 | HCT 116 | C434(1.31); C67(1.30) | LDD0790 | [3] |
| LDCM0474 | CL80 | HCT 116 | C434(1.14) | LDD0791 | [3] |
| LDCM0475 | CL81 | HCT 116 | C434(1.23) | LDD0792 | [3] |
| LDCM0476 | CL82 | HCT 116 | C434(1.54) | LDD0793 | [3] |
| LDCM0477 | CL83 | HCT 116 | C434(1.39) | LDD0794 | [3] |
| LDCM0478 | CL84 | HCT 116 | C434(1.55) | LDD0795 | [3] |
| LDCM0479 | CL85 | HCT 116 | C434(1.18) | LDD0796 | [3] |
| LDCM0480 | CL86 | HCT 116 | C434(0.96) | LDD0797 | [3] |
| LDCM0481 | CL87 | HCT 116 | C434(1.26) | LDD0798 | [3] |
| LDCM0482 | CL88 | HCT 116 | C434(1.37) | LDD0799 | [3] |
| LDCM0483 | CL89 | HCT 116 | C434(1.68) | LDD0800 | [3] |
| LDCM0484 | CL9 | HCT 116 | C434(1.02); C67(1.15) | LDD0801 | [3] |
| LDCM0485 | CL90 | HCT 116 | C434(1.22) | LDD0802 | [3] |
| LDCM0486 | CL91 | HCT 116 | C434(0.92) | LDD0803 | [3] |
| LDCM0487 | CL92 | HCT 116 | C434(0.77) | LDD0804 | [3] |
| LDCM0488 | CL93 | HCT 116 | C434(0.96) | LDD0805 | [3] |
| LDCM0489 | CL94 | HCT 116 | C434(0.99) | LDD0806 | [3] |
| LDCM0490 | CL95 | HCT 116 | C434(1.05) | LDD0807 | [3] |
| LDCM0491 | CL96 | HCT 116 | C434(0.94) | LDD0808 | [3] |
| LDCM0492 | CL97 | HCT 116 | C434(1.02) | LDD0809 | [3] |
| LDCM0493 | CL98 | HCT 116 | C434(1.04) | LDD0810 | [3] |
| LDCM0494 | CL99 | HCT 116 | C434(0.97) | LDD0811 | [3] |
| LDCM0634 | CY-0357 | Hep-G2 | C434(0.57) | LDD2228 | [8] |
| LDCM0495 | E2913 | HEK-293T | C434(1.07); C326(0.85); C371(1.02) | LDD1698 | [12] |
| LDCM0468 | Fragment33 | HCT 116 | C366(0.66); C371(0.66); C434(1.19); C67(0.90) | LDD0785 | [3] |
| LDCM0427 | Fragment51 | HCT 116 | C371(0.96); C434(1.38) | LDD0744 | [3] |
| LDCM0022 | KB02 | HCT 116 | C371(4.00); C366(8.45) | LDD0080 | [3] |
| LDCM0023 | KB03 | HCT 116 | C371(8.95); C366(13.51) | LDD0081 | [3] |
| LDCM0024 | KB05 | HCT 116 | C371(20.00); C366(17.71) | LDD0082 | [3] |
| LDCM0021 | THZ1 | HCT 116 | C366(1.09); C371(1.09) | LDD2173 | [3] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Cytokine and receptor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| C-X-C motif chemokine 9 (CXCL9) | Intercrine alpha (chemokine CxC) family | Q07325 | |||
Other
References
