Details of the Target
General Information of Target
| Target ID | LDTP11545 | |||||
|---|---|---|---|---|---|---|
| Target Name | Ubiquitin-like protein 5 (UBL5) | |||||
| Gene Name | UBL5 | |||||
| Gene ID | 59286 | |||||
| Synonyms |
Ubiquitin-like protein 5 |
|||||
| 3D Structure | ||||||
| Sequence |
MSISSDEVNFLVYRYLQESGFSHSAFTFGIESHISQSNINGALVPPAALISIIQKGLQYV
EAEVSINEDGTLFDGRPIESLSLIDAVMPDVVQTRQQAYRDKLAQQQAAAAAAAAAAASQ QGSAKNGENTANGEENGAHTIANNHTDMMEVDGDVEIPPNKAVVLRGHESEVFICAWNPV SDLLASGSGDSTARIWNLSENSTSGSTQLVLRHCIREGGQDVPSNKDVTSLDWNSEGTLL ATGSYDGFARIWTKDGNLASTLGQHKGPIFALKWNKKGNFILSAGVDKTTIIWDAHTGEA KQQFPFHSAPALDVDWQSNNTFASCSTDMCIHVCKLGQDRPIKTFQGHTNEVNAIKWDPT GNLLASCSDDMTLKIWSMKQDNCVHDLQAHNKEIYTIKWSPTGPGTNNPNANLMLASASF DSTVRLWDVDRGICIHTLTKHQEPVYSVAFSPDGRYLASGSFDKCVHIWNTQTGALVHSY RGTGGIFEVCWNAAGDKVGASASDGSVCVLDLRK |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Ubiquitin-like protein that plays a role in cell proliferation and sister chromatid cohesion by associating with spliceosomal proteins. Participates thereby in pre-mRNA splicing by maintaining spliceosome integrity. Promotes the functional integrity of the Fanconi anemia DNA repair pathway by interacting with FANCI component and subsequently mediating the formation of FANCI homodimers. Plays also a protective role against ER stress-induced apoptosis.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STPyne Probe Info |
 |
K29(1.96) | LDD0277 | [1] | |
|
IPM Probe Info |
 |
C18(0.00); C6(0.00) | LDD0241 | [2] | |
|
BTD Probe Info |
 |
C18(1.36) | LDD1700 | [3] | |
|
DBIA Probe Info |
 |
C18(1.39) | LDD0613 | [4] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C18(0.00); C6(0.00) | LDD0038 | [5] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [6] | |
|
Lodoacetamide azide Probe Info |
 |
C18(0.00); C6(0.00) | LDD0037 | [5] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [7] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [7] | |
|
NPM Probe Info |
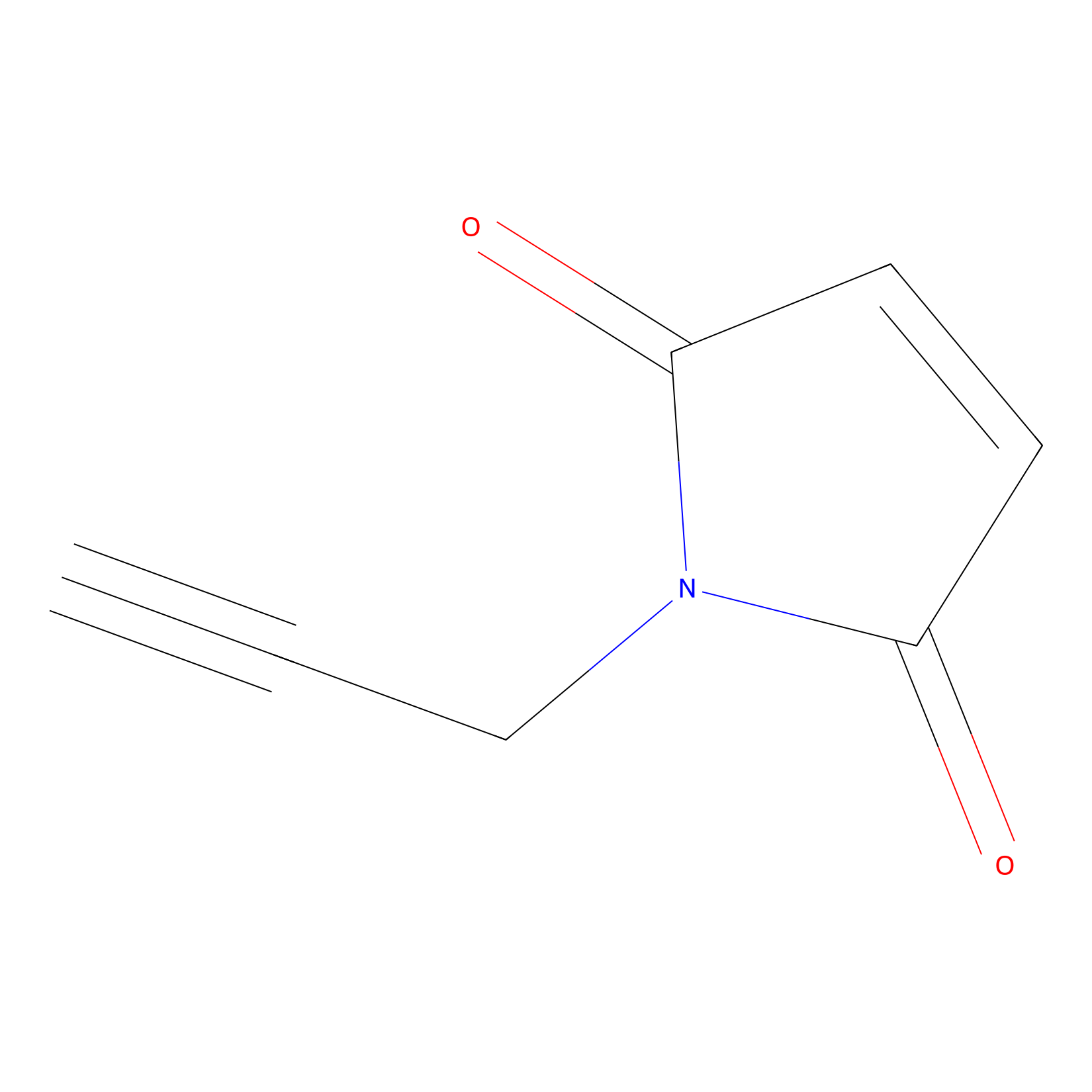 |
N.A. | LDD0016 | [8] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0152 | [9] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [10] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [2] | |
|
NAIA_5 Probe Info |
 |
C18(0.00); C6(0.00) | LDD2223 | [11] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C18(0.42) | LDD2142 | [3] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C18(0.93) | LDD2112 | [3] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C18(0.53) | LDD2095 | [3] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C18(0.97) | LDD2130 | [3] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C18(0.90) | LDD2117 | [3] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C18(1.28) | LDD2152 | [3] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C18(1.08) | LDD2103 | [3] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C18(0.31) | LDD2132 | [3] |
| LDCM0214 | AC1 | HEK-293T | C18(0.85); C6(1.08) | LDD1507 | [12] |
| LDCM0215 | AC10 | HEK-293T | C18(0.96); C6(0.97) | LDD1508 | [12] |
| LDCM0226 | AC11 | HEK-293T | C18(1.01); C6(1.00) | LDD1509 | [12] |
| LDCM0237 | AC12 | HEK-293T | C18(1.22); C6(0.98) | LDD1510 | [12] |
| LDCM0259 | AC14 | HEK-293T | C18(1.04); C6(1.03) | LDD1512 | [12] |
| LDCM0270 | AC15 | HEK-293T | C18(1.11); C6(0.97) | LDD1513 | [12] |
| LDCM0276 | AC17 | HEK-293T | C18(0.96); C6(0.96) | LDD1515 | [12] |
| LDCM0277 | AC18 | HEK-293T | C18(1.00); C6(0.99) | LDD1516 | [12] |
| LDCM0278 | AC19 | HEK-293T | C18(0.97); C6(0.94) | LDD1517 | [12] |
| LDCM0279 | AC2 | HEK-293T | C18(0.82); C6(0.86) | LDD1518 | [12] |
| LDCM0280 | AC20 | HEK-293T | C18(1.04); C6(1.10) | LDD1519 | [12] |
| LDCM0281 | AC21 | HEK-293T | C18(1.10); C6(0.93) | LDD1520 | [12] |
| LDCM0282 | AC22 | HEK-293T | C18(1.05); C6(1.08) | LDD1521 | [12] |
| LDCM0283 | AC23 | HEK-293T | C18(0.99); C6(0.96) | LDD1522 | [12] |
| LDCM0284 | AC24 | HEK-293T | C18(0.99); C6(0.99) | LDD1523 | [12] |
| LDCM0285 | AC25 | HEK-293T | C18(0.93); C6(0.90) | LDD1524 | [12] |
| LDCM0286 | AC26 | HEK-293T | C18(1.00); C6(0.94) | LDD1525 | [12] |
| LDCM0287 | AC27 | HEK-293T | C18(1.02); C6(0.98) | LDD1526 | [12] |
| LDCM0288 | AC28 | HEK-293T | C18(1.09); C6(1.00) | LDD1527 | [12] |
| LDCM0289 | AC29 | HEK-293T | C18(1.07); C6(1.04) | LDD1528 | [12] |
| LDCM0290 | AC3 | HEK-293T | C18(0.92); C6(1.02) | LDD1529 | [12] |
| LDCM0291 | AC30 | HEK-293T | C18(1.13); C6(1.17) | LDD1530 | [12] |
| LDCM0292 | AC31 | HEK-293T | C18(1.04); C6(0.91) | LDD1531 | [12] |
| LDCM0293 | AC32 | HEK-293T | C18(1.04); C6(0.94) | LDD1532 | [12] |
| LDCM0294 | AC33 | HEK-293T | C18(0.99); C6(0.92) | LDD1533 | [12] |
| LDCM0295 | AC34 | HEK-293T | C18(0.94); C6(0.98) | LDD1534 | [12] |
| LDCM0296 | AC35 | HCT 116 | C18(1.39) | LDD0613 | [4] |
| LDCM0297 | AC36 | HCT 116 | C18(1.44) | LDD0614 | [4] |
| LDCM0298 | AC37 | HCT 116 | C18(1.22) | LDD0615 | [4] |
| LDCM0299 | AC38 | HCT 116 | C18(1.48) | LDD0616 | [4] |
| LDCM0300 | AC39 | HCT 116 | C18(1.04) | LDD0617 | [4] |
| LDCM0301 | AC4 | HEK-293T | C18(1.01); C6(1.00) | LDD1540 | [12] |
| LDCM0302 | AC40 | HCT 116 | C18(0.91) | LDD0619 | [4] |
| LDCM0303 | AC41 | HCT 116 | C18(1.13) | LDD0620 | [4] |
| LDCM0304 | AC42 | HCT 116 | C18(1.26) | LDD0621 | [4] |
| LDCM0305 | AC43 | HCT 116 | C18(1.34) | LDD0622 | [4] |
| LDCM0306 | AC44 | HCT 116 | C18(0.96) | LDD0623 | [4] |
| LDCM0307 | AC45 | HCT 116 | C18(0.75) | LDD0624 | [4] |
| LDCM0308 | AC46 | HCT 116 | C18(0.99) | LDD0625 | [4] |
| LDCM0309 | AC47 | HCT 116 | C18(0.82) | LDD0626 | [4] |
| LDCM0310 | AC48 | HCT 116 | C18(1.18) | LDD0627 | [4] |
| LDCM0311 | AC49 | HCT 116 | C18(0.26) | LDD0628 | [4] |
| LDCM0312 | AC5 | HEK-293T | C18(1.03); C6(1.05) | LDD1551 | [12] |
| LDCM0313 | AC50 | HCT 116 | C18(0.34) | LDD0630 | [4] |
| LDCM0314 | AC51 | HCT 116 | C18(2.93) | LDD0631 | [4] |
| LDCM0315 | AC52 | HCT 116 | C18(0.79) | LDD0632 | [4] |
| LDCM0316 | AC53 | HCT 116 | C18(0.50) | LDD0633 | [4] |
| LDCM0317 | AC54 | HCT 116 | C18(0.62) | LDD0634 | [4] |
| LDCM0318 | AC55 | HCT 116 | C18(0.36) | LDD0635 | [4] |
| LDCM0319 | AC56 | HCT 116 | C18(0.17) | LDD0636 | [4] |
| LDCM0320 | AC57 | HEK-293T | C18(0.90); C6(0.91) | LDD1559 | [12] |
| LDCM0321 | AC58 | HEK-293T | C18(0.95); C6(0.97) | LDD1560 | [12] |
| LDCM0322 | AC59 | HEK-293T | C18(0.92); C6(0.99) | LDD1561 | [12] |
| LDCM0323 | AC6 | HEK-293T | C18(1.03); C6(1.01) | LDD1562 | [12] |
| LDCM0324 | AC60 | HEK-293T | C18(0.91); C6(0.92) | LDD1563 | [12] |
| LDCM0325 | AC61 | HEK-293T | C18(0.98); C6(0.92) | LDD1564 | [12] |
| LDCM0326 | AC62 | HEK-293T | C18(0.95); C6(0.96) | LDD1565 | [12] |
| LDCM0327 | AC63 | HEK-293T | C18(1.01); C6(0.99) | LDD1566 | [12] |
| LDCM0328 | AC64 | HEK-293T | C18(1.06); C6(1.04) | LDD1567 | [12] |
| LDCM0332 | AC68 | HCT 116 | C18(1.02) | LDD0649 | [4] |
| LDCM0333 | AC69 | HCT 116 | C18(0.78) | LDD0650 | [4] |
| LDCM0334 | AC7 | HEK-293T | C18(1.01); C6(0.96) | LDD1568 | [12] |
| LDCM0335 | AC70 | HCT 116 | C18(0.50) | LDD0652 | [4] |
| LDCM0336 | AC71 | HCT 116 | C18(1.21) | LDD0653 | [4] |
| LDCM0337 | AC72 | HCT 116 | C18(0.82) | LDD0654 | [4] |
| LDCM0338 | AC73 | HCT 116 | C18(0.30) | LDD0655 | [4] |
| LDCM0339 | AC74 | HCT 116 | C18(0.36) | LDD0656 | [4] |
| LDCM0340 | AC75 | HCT 116 | C18(0.23) | LDD0657 | [4] |
| LDCM0341 | AC76 | HCT 116 | C18(0.77) | LDD0658 | [4] |
| LDCM0342 | AC77 | HCT 116 | C18(0.60) | LDD0659 | [4] |
| LDCM0343 | AC78 | HCT 116 | C18(1.17) | LDD0660 | [4] |
| LDCM0344 | AC79 | HCT 116 | C18(0.87) | LDD0661 | [4] |
| LDCM0345 | AC8 | HEK-293T | C18(0.99); C6(1.03) | LDD1569 | [12] |
| LDCM0346 | AC80 | HCT 116 | C18(0.66) | LDD0663 | [4] |
| LDCM0347 | AC81 | HCT 116 | C18(0.74) | LDD0664 | [4] |
| LDCM0348 | AC82 | HCT 116 | C18(0.17) | LDD0665 | [4] |
| LDCM0545 | Acetamide | MDA-MB-231 | C18(0.51) | LDD2138 | [3] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C18(0.83) | LDD2113 | [3] |
| LDCM0248 | AKOS034007472 | HEK-293T | C18(1.02); C6(1.06) | LDD1511 | [12] |
| LDCM0356 | AKOS034007680 | HEK-293T | C18(0.96); C6(1.00) | LDD1570 | [12] |
| LDCM0275 | AKOS034007705 | HEK-293T | C18(1.04); C6(1.04) | LDD1514 | [12] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C18(0.46) | LDD2091 | [3] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [10] |
| LDCM0367 | CL1 | HEK-293T | C18(0.94); C6(0.91) | LDD1571 | [12] |
| LDCM0368 | CL10 | HEK-293T | C18(1.07); C6(0.93) | LDD1572 | [12] |
| LDCM0369 | CL100 | HEK-293T | C18(0.97); C6(0.87) | LDD1573 | [12] |
| LDCM0370 | CL101 | HEK-293T | C18(1.18); C6(1.03) | LDD1574 | [12] |
| LDCM0371 | CL102 | HEK-293T | C18(1.05); C6(0.90) | LDD1575 | [12] |
| LDCM0372 | CL103 | HEK-293T | C18(1.04); C6(0.91) | LDD1576 | [12] |
| LDCM0373 | CL104 | HEK-293T | C18(1.00); C6(0.95) | LDD1577 | [12] |
| LDCM0374 | CL105 | HEK-293T | C18(1.14); C6(1.03) | LDD1578 | [12] |
| LDCM0375 | CL106 | HEK-293T | C18(1.16); C6(1.00) | LDD1579 | [12] |
| LDCM0376 | CL107 | HEK-293T | C18(1.12); C6(0.90) | LDD1580 | [12] |
| LDCM0377 | CL108 | HEK-293T | C18(1.07); C6(0.91) | LDD1581 | [12] |
| LDCM0378 | CL109 | HEK-293T | C18(1.05); C6(0.86) | LDD1582 | [12] |
| LDCM0379 | CL11 | HEK-293T | C18(1.09); C6(0.98) | LDD1583 | [12] |
| LDCM0380 | CL110 | HEK-293T | C18(1.07); C6(0.92) | LDD1584 | [12] |
| LDCM0381 | CL111 | HEK-293T | C18(1.04); C6(0.92) | LDD1585 | [12] |
| LDCM0382 | CL112 | HEK-293T | C18(1.04); C6(0.93) | LDD1586 | [12] |
| LDCM0383 | CL113 | HEK-293T | C18(1.19); C6(0.98) | LDD1587 | [12] |
| LDCM0384 | CL114 | HEK-293T | C18(1.04); C6(0.95) | LDD1588 | [12] |
| LDCM0385 | CL115 | HEK-293T | C18(0.83); C6(1.02) | LDD1589 | [12] |
| LDCM0386 | CL116 | HEK-293T | C18(0.97); C6(0.98) | LDD1590 | [12] |
| LDCM0387 | CL117 | HCT 116 | C18(0.36) | LDD0704 | [4] |
| LDCM0388 | CL118 | HCT 116 | C18(1.32) | LDD0705 | [4] |
| LDCM0389 | CL119 | HCT 116 | C18(1.01) | LDD0706 | [4] |
| LDCM0390 | CL12 | HEK-293T | C18(1.14); C6(1.02) | LDD1594 | [12] |
| LDCM0391 | CL120 | HCT 116 | C18(1.50) | LDD0708 | [4] |
| LDCM0392 | CL121 | HCT 116 | C18(0.95) | LDD0709 | [4] |
| LDCM0393 | CL122 | HCT 116 | C18(0.72) | LDD0710 | [4] |
| LDCM0394 | CL123 | HCT 116 | C18(0.60) | LDD0711 | [4] |
| LDCM0395 | CL124 | HCT 116 | C18(0.41) | LDD0712 | [4] |
| LDCM0396 | CL125 | HEK-293T | C18(0.96); C6(0.97) | LDD1600 | [12] |
| LDCM0397 | CL126 | HEK-293T | C18(1.11); C6(1.06) | LDD1601 | [12] |
| LDCM0398 | CL127 | HEK-293T | C18(0.99); C6(0.98) | LDD1602 | [12] |
| LDCM0399 | CL128 | HEK-293T | C18(0.99); C6(0.94) | LDD1603 | [12] |
| LDCM0400 | CL13 | HEK-293T | C18(0.90); C6(1.01) | LDD1604 | [12] |
| LDCM0401 | CL14 | HEK-293T | C18(1.14); C6(1.01) | LDD1605 | [12] |
| LDCM0402 | CL15 | HEK-293T | C18(0.79); C6(0.85) | LDD1606 | [12] |
| LDCM0403 | CL16 | HEK-293T | C18(1.24); C6(0.97) | LDD1607 | [12] |
| LDCM0404 | CL17 | HEK-293T | C18(0.95); C6(0.90) | LDD1608 | [12] |
| LDCM0405 | CL18 | HEK-293T | C18(1.07); C6(0.92) | LDD1609 | [12] |
| LDCM0406 | CL19 | HEK-293T | C18(1.11); C6(0.98) | LDD1610 | [12] |
| LDCM0407 | CL2 | HEK-293T | C18(1.11); C6(0.96) | LDD1611 | [12] |
| LDCM0408 | CL20 | HEK-293T | C18(1.37); C6(1.06) | LDD1612 | [12] |
| LDCM0409 | CL21 | HEK-293T | C18(1.11); C6(0.98) | LDD1613 | [12] |
| LDCM0410 | CL22 | HEK-293T | C18(1.23); C6(1.06) | LDD1614 | [12] |
| LDCM0411 | CL23 | HEK-293T | C18(1.13); C6(1.10) | LDD1615 | [12] |
| LDCM0412 | CL24 | HEK-293T | C18(1.29); C6(1.09) | LDD1616 | [12] |
| LDCM0413 | CL25 | HEK-293T | C18(1.02); C6(0.93) | LDD1617 | [12] |
| LDCM0414 | CL26 | HEK-293T | C18(1.19); C6(1.05) | LDD1618 | [12] |
| LDCM0415 | CL27 | HEK-293T | C18(1.12); C6(0.95) | LDD1619 | [12] |
| LDCM0416 | CL28 | HEK-293T | C18(1.05); C6(0.89) | LDD1620 | [12] |
| LDCM0417 | CL29 | HEK-293T | C18(1.07); C6(0.93) | LDD1621 | [12] |
| LDCM0418 | CL3 | HEK-293T | C18(1.04); C6(0.90) | LDD1622 | [12] |
| LDCM0419 | CL30 | HEK-293T | C18(1.19); C6(0.96) | LDD1623 | [12] |
| LDCM0420 | CL31 | HCT 116 | C18(0.85) | LDD0737 | [4] |
| LDCM0421 | CL32 | HCT 116 | C18(0.85) | LDD0738 | [4] |
| LDCM0422 | CL33 | HCT 116 | C18(0.72) | LDD0739 | [4] |
| LDCM0423 | CL34 | HCT 116 | C18(0.38) | LDD0740 | [4] |
| LDCM0424 | CL35 | HCT 116 | C18(0.44) | LDD0741 | [4] |
| LDCM0425 | CL36 | HCT 116 | C18(0.50) | LDD0742 | [4] |
| LDCM0426 | CL37 | HCT 116 | C18(0.47) | LDD0743 | [4] |
| LDCM0428 | CL39 | HCT 116 | C18(0.47) | LDD0745 | [4] |
| LDCM0429 | CL4 | HEK-293T | C18(1.02); C6(1.01) | LDD1633 | [12] |
| LDCM0430 | CL40 | HCT 116 | C18(0.64) | LDD0747 | [4] |
| LDCM0431 | CL41 | HCT 116 | C18(0.77) | LDD0748 | [4] |
| LDCM0432 | CL42 | HCT 116 | C18(0.21) | LDD0749 | [4] |
| LDCM0433 | CL43 | HCT 116 | C18(0.33) | LDD0750 | [4] |
| LDCM0434 | CL44 | HCT 116 | C18(0.79) | LDD0751 | [4] |
| LDCM0435 | CL45 | HCT 116 | C18(0.52) | LDD0752 | [4] |
| LDCM0436 | CL46 | HEK-293T | C18(1.43); C6(1.14) | LDD1640 | [12] |
| LDCM0437 | CL47 | HEK-293T | C18(1.03); C6(1.05) | LDD1641 | [12] |
| LDCM0438 | CL48 | HEK-293T | C18(1.47); C6(0.99) | LDD1642 | [12] |
| LDCM0439 | CL49 | HEK-293T | C18(1.09); C6(1.18) | LDD1643 | [12] |
| LDCM0440 | CL5 | HEK-293T | C18(0.96); C6(0.98) | LDD1644 | [12] |
| LDCM0441 | CL50 | HEK-293T | C18(1.21); C6(0.98) | LDD1645 | [12] |
| LDCM0443 | CL52 | HEK-293T | C18(1.16); C6(1.05) | LDD1646 | [12] |
| LDCM0444 | CL53 | HEK-293T | C18(1.06); C6(0.90) | LDD1647 | [12] |
| LDCM0445 | CL54 | HEK-293T | C18(1.15); C6(0.83) | LDD1648 | [12] |
| LDCM0446 | CL55 | HEK-293T | C18(1.14); C6(1.12) | LDD1649 | [12] |
| LDCM0447 | CL56 | HEK-293T | C18(1.19); C6(1.07) | LDD1650 | [12] |
| LDCM0448 | CL57 | HEK-293T | C18(1.28); C6(1.14) | LDD1651 | [12] |
| LDCM0449 | CL58 | HEK-293T | C18(1.46); C6(1.09) | LDD1652 | [12] |
| LDCM0450 | CL59 | HEK-293T | C18(1.18); C6(1.10) | LDD1653 | [12] |
| LDCM0451 | CL6 | HEK-293T | C18(0.86); C6(0.88) | LDD1654 | [12] |
| LDCM0452 | CL60 | HEK-293T | C18(1.42); C6(1.00) | LDD1655 | [12] |
| LDCM0453 | CL61 | HEK-293T | C18(0.98); C6(0.99) | LDD1656 | [12] |
| LDCM0454 | CL62 | HEK-293T | C18(1.14); C6(0.98) | LDD1657 | [12] |
| LDCM0455 | CL63 | HEK-293T | C18(1.08); C6(0.96) | LDD1658 | [12] |
| LDCM0456 | CL64 | HEK-293T | C18(1.05); C6(1.05) | LDD1659 | [12] |
| LDCM0457 | CL65 | HEK-293T | C18(0.99); C6(1.05) | LDD1660 | [12] |
| LDCM0458 | CL66 | HEK-293T | C18(1.02); C6(0.96) | LDD1661 | [12] |
| LDCM0459 | CL67 | HEK-293T | C18(1.10); C6(1.09) | LDD1662 | [12] |
| LDCM0460 | CL68 | HEK-293T | C18(1.14); C6(1.09) | LDD1663 | [12] |
| LDCM0461 | CL69 | HEK-293T | C18(1.31); C6(1.19) | LDD1664 | [12] |
| LDCM0462 | CL7 | HEK-293T | C18(0.94); C6(1.04) | LDD1665 | [12] |
| LDCM0463 | CL70 | HEK-293T | C18(1.23); C6(1.08) | LDD1666 | [12] |
| LDCM0464 | CL71 | HEK-293T | C18(1.14); C6(0.96) | LDD1667 | [12] |
| LDCM0465 | CL72 | HEK-293T | C18(1.26); C6(1.11) | LDD1668 | [12] |
| LDCM0466 | CL73 | HEK-293T | C18(1.01); C6(1.01) | LDD1669 | [12] |
| LDCM0467 | CL74 | HEK-293T | C18(1.09); C6(0.92) | LDD1670 | [12] |
| LDCM0469 | CL76 | HEK-293T | C18(1.12); C6(1.02) | LDD1672 | [12] |
| LDCM0470 | CL77 | HEK-293T | C18(0.94); C6(0.96) | LDD1673 | [12] |
| LDCM0471 | CL78 | HEK-293T | C18(1.01); C6(1.00) | LDD1674 | [12] |
| LDCM0472 | CL79 | HEK-293T | C18(1.16); C6(1.14) | LDD1675 | [12] |
| LDCM0473 | CL8 | HEK-293T | C18(1.02); C6(0.79) | LDD1676 | [12] |
| LDCM0474 | CL80 | HEK-293T | C18(1.36); C6(1.07) | LDD1677 | [12] |
| LDCM0475 | CL81 | HEK-293T | C18(1.35); C6(1.12) | LDD1678 | [12] |
| LDCM0476 | CL82 | HEK-293T | C18(1.33); C6(1.06) | LDD1679 | [12] |
| LDCM0477 | CL83 | HEK-293T | C18(1.11); C6(1.16) | LDD1680 | [12] |
| LDCM0478 | CL84 | HEK-293T | C18(1.22); C6(1.14) | LDD1681 | [12] |
| LDCM0479 | CL85 | HEK-293T | C18(0.93); C6(0.99) | LDD1682 | [12] |
| LDCM0480 | CL86 | HEK-293T | C18(0.89); C6(1.06) | LDD1683 | [12] |
| LDCM0481 | CL87 | HEK-293T | C18(1.07); C6(1.02) | LDD1684 | [12] |
| LDCM0482 | CL88 | HEK-293T | C18(1.01); C6(0.83) | LDD1685 | [12] |
| LDCM0483 | CL89 | HEK-293T | C18(0.96); C6(1.01) | LDD1686 | [12] |
| LDCM0484 | CL9 | HEK-293T | C18(1.22); C6(1.10) | LDD1687 | [12] |
| LDCM0485 | CL90 | HEK-293T | C18(0.89); C6(0.83) | LDD1688 | [12] |
| LDCM0486 | CL91 | HEK-293T | C18(1.03); C6(1.02) | LDD1689 | [12] |
| LDCM0487 | CL92 | HEK-293T | C18(1.20); C6(0.94) | LDD1690 | [12] |
| LDCM0488 | CL93 | HEK-293T | C18(1.14); C6(1.06) | LDD1691 | [12] |
| LDCM0489 | CL94 | HEK-293T | C18(1.18); C6(1.11) | LDD1692 | [12] |
| LDCM0490 | CL95 | HEK-293T | C18(0.98); C6(0.83) | LDD1693 | [12] |
| LDCM0491 | CL96 | HEK-293T | C18(1.01); C6(1.12) | LDD1694 | [12] |
| LDCM0492 | CL97 | HEK-293T | C18(1.06); C6(0.97) | LDD1695 | [12] |
| LDCM0493 | CL98 | HEK-293T | C18(1.13); C6(1.02) | LDD1696 | [12] |
| LDCM0494 | CL99 | HEK-293T | C18(1.19); C6(0.90) | LDD1697 | [12] |
| LDCM0495 | E2913 | HEK-293T | C18(0.96); C6(0.99) | LDD1698 | [12] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C18(1.67) | LDD1702 | [3] |
| LDCM0625 | F8 | Ramos | C6(0.96); C18(0.93) | LDD2187 | [13] |
| LDCM0572 | Fragment10 | Ramos | C6(0.52); C18(0.59) | LDD2189 | [13] |
| LDCM0573 | Fragment11 | Ramos | C6(1.18); C18(0.52) | LDD2190 | [13] |
| LDCM0574 | Fragment12 | Ramos | C6(0.72); C18(0.61) | LDD2191 | [13] |
| LDCM0575 | Fragment13 | Ramos | C6(0.88); C18(0.73) | LDD2192 | [13] |
| LDCM0576 | Fragment14 | Ramos | C6(1.17); C18(1.32) | LDD2193 | [13] |
| LDCM0579 | Fragment20 | Ramos | C6(0.55); C18(0.57) | LDD2194 | [13] |
| LDCM0580 | Fragment21 | Ramos | C6(0.90); C18(0.67) | LDD2195 | [13] |
| LDCM0582 | Fragment23 | Ramos | C6(0.27); C18(0.21) | LDD2196 | [13] |
| LDCM0578 | Fragment27 | Ramos | C6(0.87); C18(0.66) | LDD2197 | [13] |
| LDCM0586 | Fragment28 | Ramos | C6(0.80); C18(0.71) | LDD2198 | [13] |
| LDCM0588 | Fragment30 | Ramos | C6(0.90); C18(0.85) | LDD2199 | [13] |
| LDCM0589 | Fragment31 | Ramos | C6(0.92); C18(0.77) | LDD2200 | [13] |
| LDCM0590 | Fragment32 | Ramos | C6(0.55); C18(0.53) | LDD2201 | [13] |
| LDCM0468 | Fragment33 | HEK-293T | C18(0.99); C6(0.97) | LDD1671 | [12] |
| LDCM0596 | Fragment38 | Ramos | C6(0.91); C18(0.62) | LDD2203 | [13] |
| LDCM0566 | Fragment4 | Ramos | C6(0.82); C18(0.81) | LDD2184 | [13] |
| LDCM0427 | Fragment51 | HCT 116 | C18(0.29) | LDD0744 | [4] |
| LDCM0610 | Fragment52 | Ramos | C6(1.00); C18(0.72) | LDD2204 | [13] |
| LDCM0614 | Fragment56 | Ramos | C6(0.96); C18(0.91) | LDD2205 | [13] |
| LDCM0569 | Fragment7 | Ramos | C6(0.73); C18(0.75) | LDD2186 | [13] |
| LDCM0571 | Fragment9 | Ramos | C6(0.55); C18(0.46) | LDD2188 | [13] |
| LDCM0022 | KB02 | HEK-293T | C18(0.76); C6(0.97) | LDD1492 | [12] |
| LDCM0023 | KB03 | HEK-293T | C18(1.06); C6(0.98) | LDD1497 | [12] |
| LDCM0024 | KB05 | MEL167 | C6(2.37) | LDD3316 | [14] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C18(1.04) | LDD2102 | [3] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C18(0.58) | LDD2121 | [3] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C18(0.99) | LDD2089 | [3] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C18(1.20) | LDD2092 | [3] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C18(0.74) | LDD2093 | [3] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C18(3.24) | LDD2094 | [3] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C18(0.32) | LDD2096 | [3] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C18(0.77) | LDD2097 | [3] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C18(1.08) | LDD2098 | [3] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C18(0.99) | LDD2099 | [3] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C18(0.90) | LDD2101 | [3] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C18(0.60) | LDD2104 | [3] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C18(2.77) | LDD2105 | [3] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C18(0.94) | LDD2107 | [3] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C18(0.38) | LDD2108 | [3] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C18(0.75) | LDD2109 | [3] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C18(0.80) | LDD2111 | [3] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C18(0.64) | LDD2114 | [3] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C18(0.51) | LDD2115 | [3] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C18(0.70) | LDD2116 | [3] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C18(0.74) | LDD2118 | [3] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C18(2.49) | LDD2119 | [3] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C18(0.90) | LDD2120 | [3] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C18(0.60) | LDD2122 | [3] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C18(0.40) | LDD2124 | [3] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C18(0.86) | LDD2125 | [3] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C18(0.42) | LDD2126 | [3] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C18(1.08) | LDD2127 | [3] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C18(0.69) | LDD2128 | [3] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C18(0.97) | LDD2129 | [3] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C18(0.69); C6(1.15) | LDD2133 | [3] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C18(0.55) | LDD2134 | [3] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C18(0.88) | LDD2136 | [3] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C18(0.96) | LDD2137 | [3] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C18(1.36) | LDD1700 | [3] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C18(0.72) | LDD2140 | [3] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C18(0.44) | LDD2141 | [3] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C18(0.95) | LDD2143 | [3] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C18(3.10) | LDD2144 | [3] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C18(2.19) | LDD2145 | [3] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C18(0.87) | LDD2146 | [3] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C18(3.38) | LDD2147 | [3] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C18(0.75) | LDD2148 | [3] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C18(0.82) | LDD2149 | [3] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C18(0.29) | LDD2150 | [3] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C18(0.54) | LDD2151 | [3] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C18(0.80) | LDD2206 | [15] |
| LDCM0131 | RA190 | MM1.R | C6(1.13) | LDD0304 | [16] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transcription factor
Other
References
