Details of the Target
General Information of Target
| Target ID | LDTP10889 | |||||
|---|---|---|---|---|---|---|
| Target Name | Perilipin-2 (PLIN2) | |||||
| Gene Name | PLIN2 | |||||
| Gene ID | 123 | |||||
| Synonyms |
ADFP; Perilipin-2; Adipophilin; Adipose differentiation-related protein; ADRP |
|||||
| 3D Structure | ||||||
| Sequence |
MVWKVAVFLSVALGIGAVPIDDPEDGGKHWVVIVAGSNGWYNYRHQADACHAYQIIHRNG
IPDEQIVVMMYDDIAYSEDNPTPGIVINRPNGTDVYQGVPKDYTGEDVTPQNFLAVLRGD AEAVKGIGSGKVLKSGPQDHVFIYFTDHGSTGILVFPNEDLHVKDLNETIHYMYKHKMYR KMVFYIEACESGSMMNHLPDNINVYATTAANPRESSYACYYDEKRSTYLGDWYSVNWMED SDVEDLTKETLHKQYHLVKSHTNTSHVMQYGNKTISTMKVMQFQGMKRKASSPVPLPPVT HLDLTPSPDVPLTIMKRKLMNTNDLEESRQLTEEIQRHLDARHLIEKSVRKIVSLLAASE AEVEQLLSERAPLTGHSCYPEALLHFRTHCFNWHSPTYEYALRHLYVLVNLCEKPYPLHR IKLSMDHVCLGHY |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Perilipin family
|
|||||
| Subcellular location |
Membrane
|
|||||
| Function | Structural component of lipid droplets, which is required for the formation and maintenance of lipid storage droplets. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [2] | |
|
TG42 Probe Info |
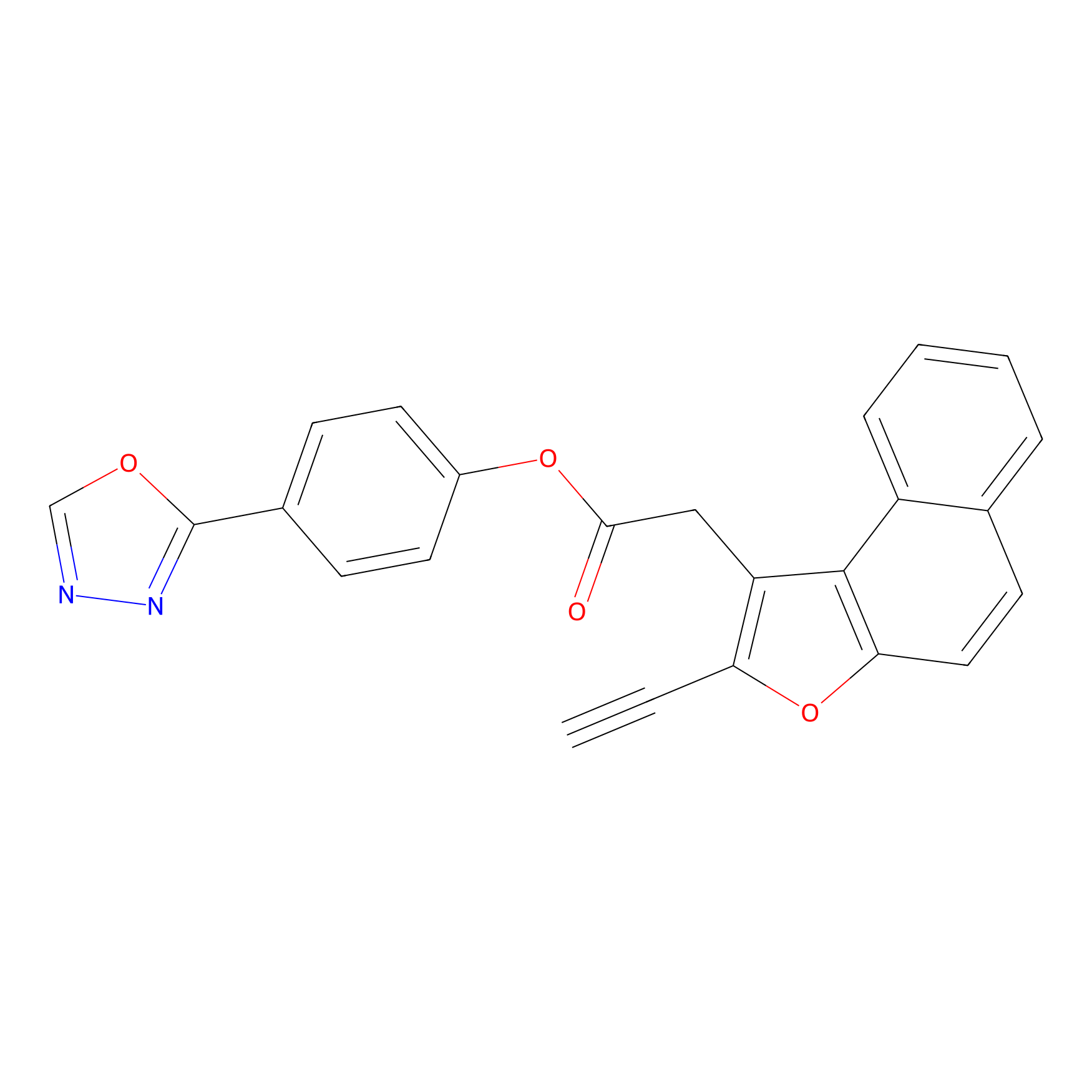 |
5.38 | LDD0326 | [3] | |
|
W1 Probe Info |
 |
28.71 | LDD0235 | [4] | |
|
TH211 Probe Info |
 |
Y82(10.02); Y216(7.99) | LDD0257 | [5] | |
|
TH216 Probe Info |
 |
Y82(12.98); Y217(5.11) | LDD0259 | [5] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [6] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [6] | |
|
STPyne Probe Info |
 |
K243(1.69); K265(6.25) | LDD0277 | [7] | |
|
BTD Probe Info |
 |
C47(0.65) | LDD2095 | [8] | |
|
FBPP2 Probe Info |
 |
2.46 | LDD0054 | [9] | |
|
AHL-Pu-1 Probe Info |
 |
C302(5.20) | LDD0171 | [10] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [11] | |
|
IA-alkyne Probe Info |
 |
C302(20.00) | LDD1706 | [12] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0225 | [13] | |
|
DBIA Probe Info |
 |
C84(1.68) | LDD0080 | [14] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [15] | |
|
NAIA_4 Probe Info |
 |
C302(0.00); C325(0.00) | LDD2226 | [16] | |
|
TFBX Probe Info |
 |
C47(0.00); C302(0.00) | LDD0027 | [15] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [17] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [17] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [17] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [18] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DR-1 Probe Info |
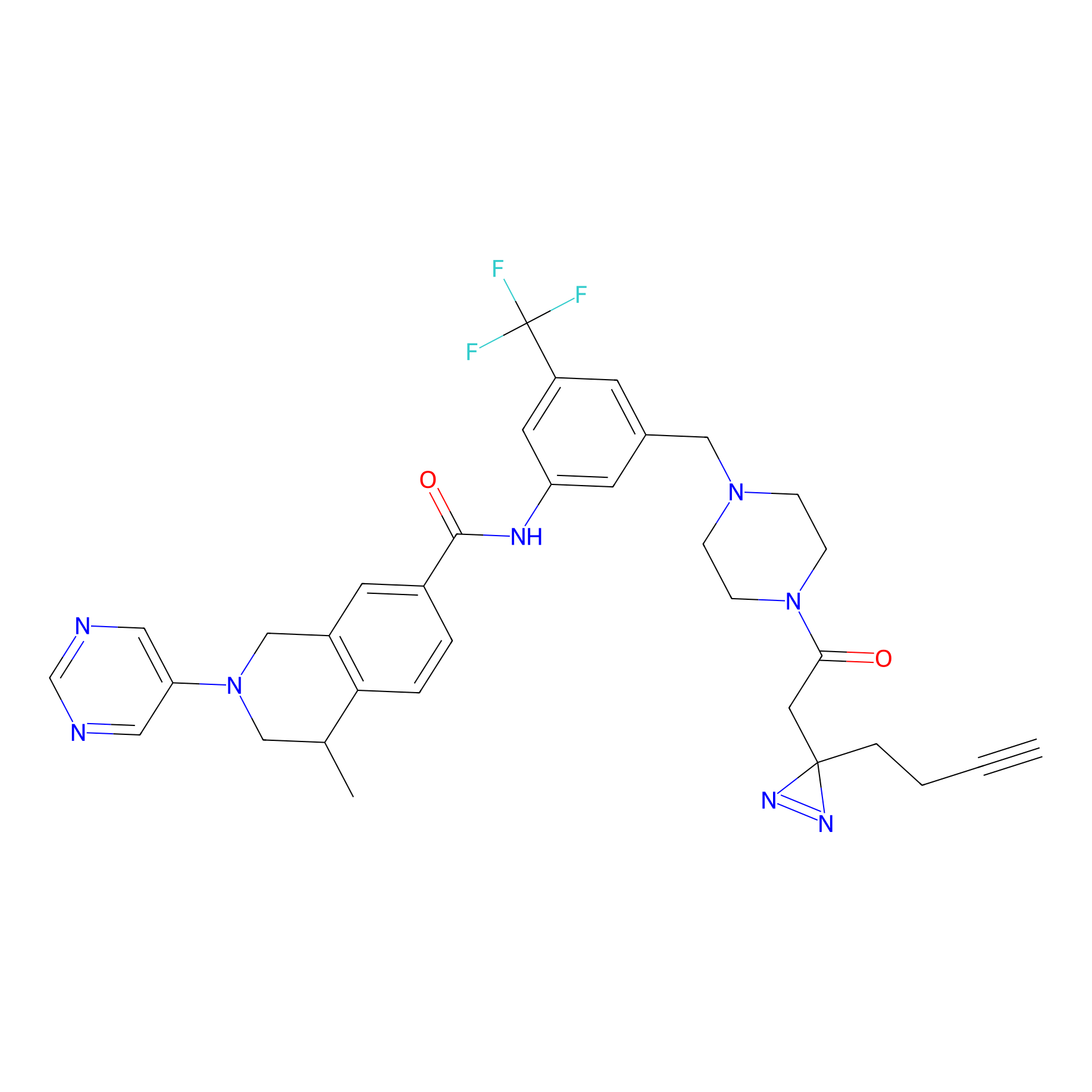 |
5.46 | LDD0398 | [19] | |
|
C362 Probe Info |
 |
43.71 | LDD2023 | [20] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [21] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [21] | |
|
FFF probe14 Probe Info |
 |
20.00 | LDD0477 | [21] | |
|
FFF probe2 Probe Info |
 |
20.00 | LDD0463 | [21] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0465 | [21] | |
|
A-DA Probe Info |
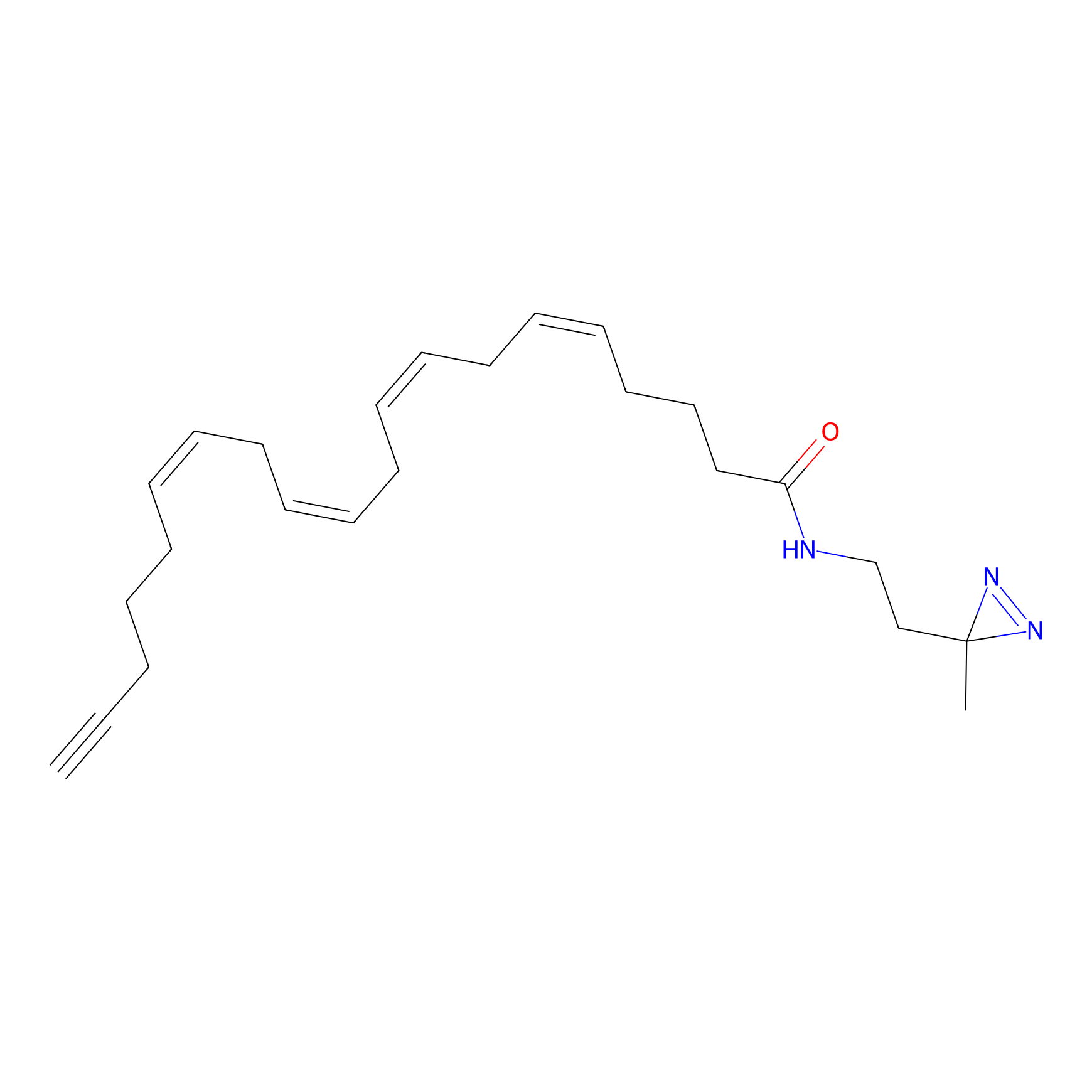 |
9.04 | LDD0143 | [22] | |
|
OEA-DA Probe Info |
 |
20.00 | LDD0046 | [23] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C47(0.65) | LDD2095 | [8] |
| LDCM0026 | 4SU-RNA+native RNA | DM93 | C302(5.20) | LDD0171 | [10] |
| LDCM0259 | AC14 | HEK-293T | C84(1.03) | LDD1512 | [24] |
| LDCM0282 | AC22 | HEK-293T | C84(1.01) | LDD1521 | [24] |
| LDCM0291 | AC30 | HEK-293T | C84(1.03) | LDD1530 | [24] |
| LDCM0299 | AC38 | HEK-293T | C84(1.21) | LDD1538 | [24] |
| LDCM0308 | AC46 | HEK-293T | C84(1.13) | LDD1547 | [24] |
| LDCM0317 | AC54 | HEK-293T | C84(1.06) | LDD1556 | [24] |
| LDCM0323 | AC6 | HEK-293T | C84(1.03) | LDD1562 | [24] |
| LDCM0326 | AC62 | HEK-293T | C84(0.91) | LDD1565 | [24] |
| LDCM0156 | Aniline | NCI-H1299 | 12.99 | LDD0403 | [1] |
| LDCM0083 | Avasimibe | A-549 | 9.04 | LDD0143 | [22] |
| LDCM0368 | CL10 | HEK-293T | C84(1.04) | LDD1572 | [24] |
| LDCM0410 | CL22 | HEK-293T | C84(1.04) | LDD1614 | [24] |
| LDCM0423 | CL34 | HEK-293T | C84(1.04) | LDD1627 | [24] |
| LDCM0436 | CL46 | HEK-293T | C84(1.07) | LDD1640 | [24] |
| LDCM0449 | CL58 | HEK-293T | C84(0.88) | LDD1652 | [24] |
| LDCM0463 | CL70 | HEK-293T | C84(1.19) | LDD1666 | [24] |
| LDCM0476 | CL82 | HEK-293T | C84(1.04) | LDD1679 | [24] |
| LDCM0489 | CL94 | HEK-293T | C84(1.10) | LDD1692 | [24] |
| LDCM0175 | Ethacrynic acid | HeLa | N.A. | LDD0440 | [11] |
| LDCM0022 | KB02 | HCT 116 | C84(1.68) | LDD0080 | [14] |
| LDCM0023 | KB03 | HCT 116 | C84(1.24) | LDD0081 | [14] |
| LDCM0024 | KB05 | HCT 116 | C84(1.51) | LDD0082 | [14] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0225 | [13] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C47(1.46) | LDD2137 | [8] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C302(1.35) | LDD2150 | [8] |
| LDCM0084 | Ro 48-8071 | A-549 | 9.15 | LDD0145 | [22] |
| LDCM0008 | Tranylcypromine | SH-SY5Y | 2.46 | LDD0054 | [9] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Deubiquitinase DESI2 (DESI2) | DeSI family | Q9BSY9 | |||
| 1-acylglycerol-3-phosphate O-acyltransferase ABHD5 (ABHD5) | Peptidase S33 family | Q8WTS1 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Wolframin (WFS1) | . | O76024 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Pogo transposable element with ZNF domain (POGZ) | . | Q7Z3K3 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Vesicle transport protein SFT2B (SFT2D2) | SFT2 family | O95562 | |||
References
