Details of the Target
General Information of Target
| Target ID | LDTP05023 | |||||
|---|---|---|---|---|---|---|
| Target Name | Large ribosomal subunit protein eL31 (RPL31) | |||||
| Gene Name | RPL31 | |||||
| Gene ID | 6160 | |||||
| Synonyms |
Large ribosomal subunit protein eL31; 60S ribosomal protein L31 |
|||||
| 3D Structure | ||||||
| Sequence |
MAPAKKGGEKKKGRSAINEVVTREYTINIHKRIHGVGFKKRAPRALKEIRKFAMKEMGTP
DVRIDTRLNKAVWAKGIRNVPYRIRVRLSRKRNEDEDSPNKLYTLVTYVPVTTFKNLQTV NVDEN |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Eukaryotic ribosomal protein eL31 family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function | Component of the large ribosomal subunit. The ribosome is a large ribonucleoprotein complex responsible for the synthesis of proteins in the cell. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P1 Probe Info |
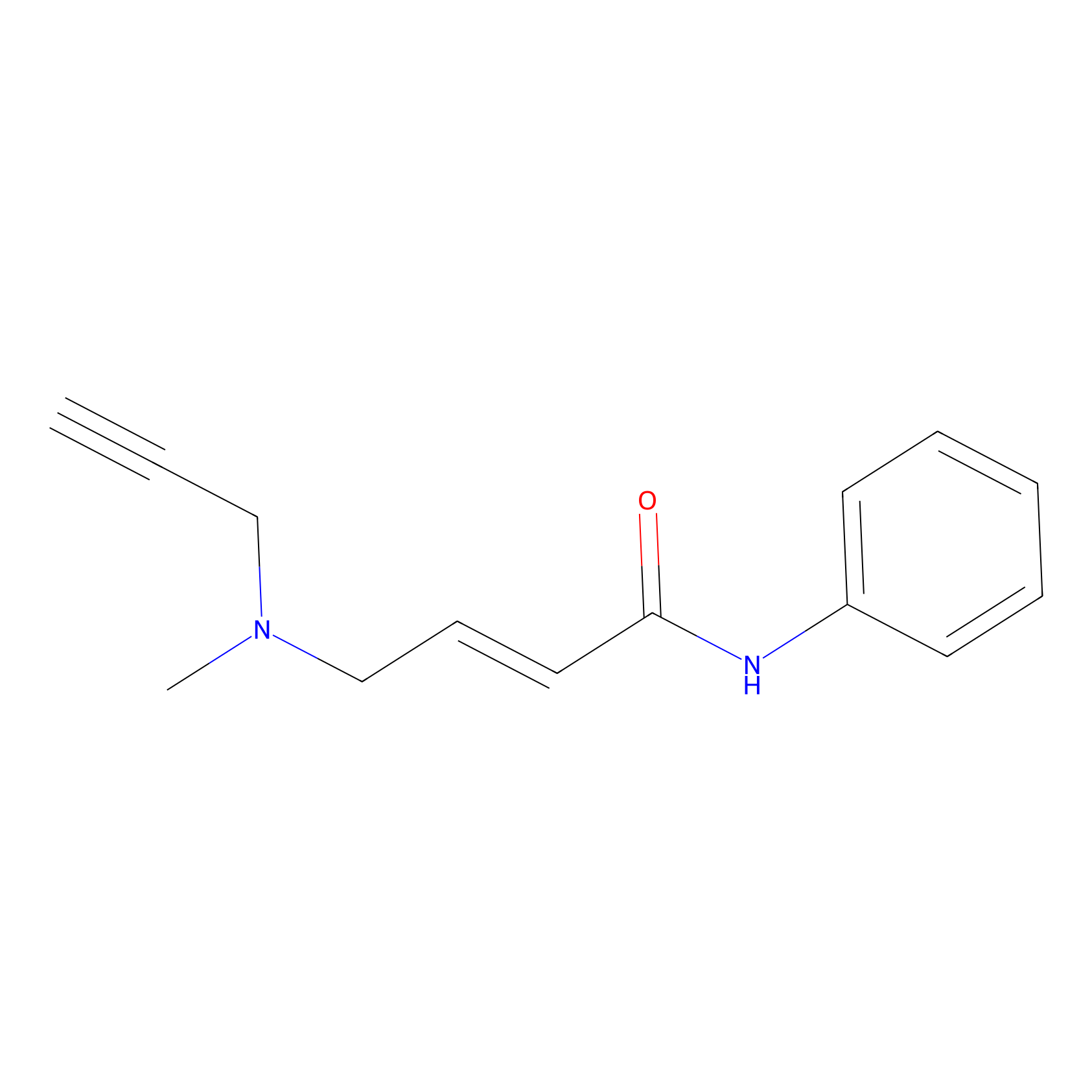 |
2.48 | LDD0452 | [1] | |
|
P2 Probe Info |
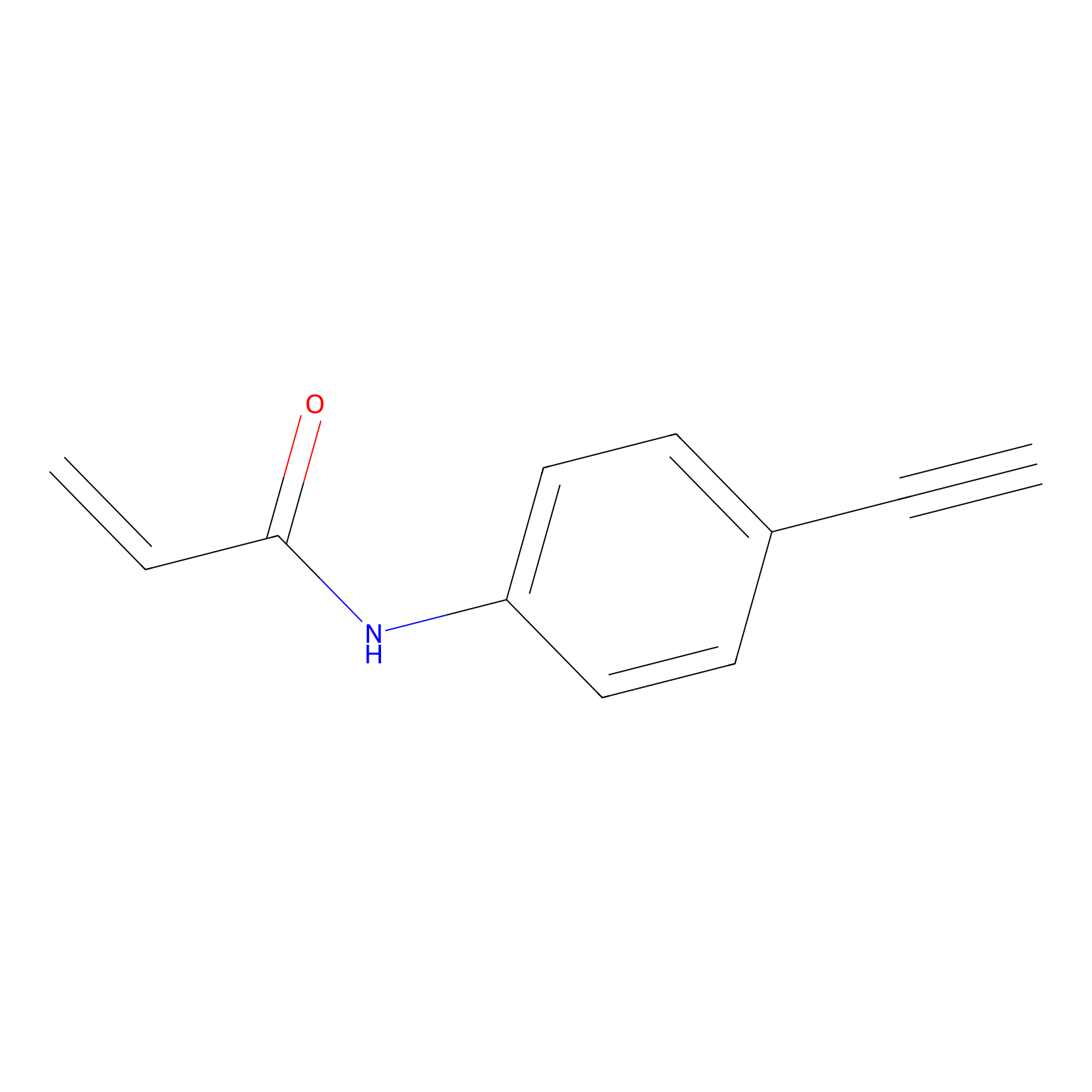 |
4.28 | LDD0453 | [1] | |
|
P3 Probe Info |
 |
1.79 | LDD0454 | [1] | |
|
A-EBA Probe Info |
 |
3.69 | LDD0215 | [2] | |
|
C-Sul Probe Info |
 |
7.12 | LDD0066 | [3] | |
|
TH211 Probe Info |
 |
Y25(20.00) | LDD0257 | [4] | |
|
TH214 Probe Info |
 |
Y25(20.00) | LDD0258 | [4] | |
|
TH216 Probe Info |
 |
Y103(20.00); Y108(20.00); Y25(6.36) | LDD0259 | [4] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [5] | |
|
ONAyne Probe Info |
 |
K31(10.00); K70(5.00); K75(0.72) | LDD0274 | [6] | |
|
STPyne Probe Info |
 |
K101(0.84); K31(6.67); K39(10.00); K70(10.00) | LDD0277 | [6] | |
|
AZ-9 Probe Info |
 |
E19(0.95) | LDD2208 | [7] | |
|
OPA-S-S-alkyne Probe Info |
 |
K31(1.97); K75(2.92); K70(12.86) | LDD3494 | [8] | |
|
HHS-482 Probe Info |
 |
Y103(1.79); Y108(0.81) | LDD0285 | [9] | |
|
HHS-475 Probe Info |
 |
Y103(0.62) | LDD0264 | [10] | |
|
HHS-465 Probe Info |
 |
Y108(5.15) | LDD2237 | [11] | |
|
ATP probe Probe Info |
 |
K75(0.00); K31(0.00); K55(0.00); K70(0.00) | LDD0199 | [12] | |
|
ATP probe Probe Info |
 |
K75(0.00); K31(0.00) | LDD0035 | [13] | |
|
1d-yne Probe Info |
 |
K115(0.00); K101(0.00) | LDD0356 | [14] | |
|
SF Probe Info |
 |
Y25(0.00); K91(0.00); Y108(0.00) | LDD0028 | [15] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [16] | |
|
1c-yne Probe Info |
 |
K55(0.00); K101(0.00); K115(0.00) | LDD0228 | [14] | |
|
Acrolein Probe Info |
 |
H34(0.00); H30(0.00) | LDD0217 | [17] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [17] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [17] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [18] | |
|
MPP-AC Probe Info |
 |
N.A. | LDD0428 | [19] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [19] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-1 Probe Info |
 |
N.A. | LDD0137 | [20] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | H30(0.00); H34(0.00) | LDD0222 | [17] |
| LDCM0116 | HHS-0101 | DM93 | Y103(0.62) | LDD0264 | [10] |
| LDCM0117 | HHS-0201 | DM93 | Y103(0.80) | LDD0265 | [10] |
| LDCM0118 | HHS-0301 | DM93 | Y103(0.82) | LDD0266 | [10] |
| LDCM0119 | HHS-0401 | DM93 | Y103(0.71) | LDD0267 | [10] |
| LDCM0120 | HHS-0701 | DM93 | Y103(0.56) | LDD0268 | [10] |
| LDCM0107 | IAA | HeLa | H30(0.00); H34(0.00) | LDD0221 | [17] |
| LDCM0123 | JWB131 | DM93 | Y103(1.79); Y108(0.81) | LDD0285 | [9] |
| LDCM0124 | JWB142 | DM93 | Y103(1.45) | LDD0286 | [9] |
| LDCM0125 | JWB146 | DM93 | Y103(1.86) | LDD0287 | [9] |
| LDCM0126 | JWB150 | DM93 | Y103(6.13) | LDD0288 | [9] |
| LDCM0127 | JWB152 | DM93 | Y103(5.25); Y108(2.78) | LDD0289 | [9] |
| LDCM0128 | JWB198 | DM93 | Y103(2.50) | LDD0290 | [9] |
| LDCM0129 | JWB202 | DM93 | Y103(1.63) | LDD0291 | [9] |
| LDCM0130 | JWB211 | DM93 | Y103(2.19) | LDD0292 | [9] |
| LDCM0109 | NEM | HeLa | H30(0.00); H34(0.00) | LDD0223 | [17] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Fibroblast growth factor receptor 3 (FGFR3) | Tyr protein kinase family | P22607 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Huntingtin (HTT) | Huntingtin family | P42858 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Gelsolin (GSN) | Villin/gelsolin family | P06396 | |||
| Ubiquilin-1 (UBQLN1) | . | Q9UMX0 | |||
References
