Details of the Target
General Information of Target
| Target ID | LDTP04558 | |||||
|---|---|---|---|---|---|---|
| Target Name | Arfaptin-1 (ARFIP1) | |||||
| Gene Name | ARFIP1 | |||||
| Gene ID | 27236 | |||||
| Synonyms |
Arfaptin-1; ADP-ribosylation factor-interacting protein 1 |
|||||
| 3D Structure | ||||||
| Sequence |
MAQESPKNSAAEIPVTSNGEVDDSREHSFNRDLKHSLPSGLGLSETQITSHGFDNTKEGV
IEAGAFQGSPAPPLPSVMSPSRVAASRLAQQGSDLIVPAGGQRTQTKSGPVILADEIKNP AMEKLELVRKWSLNTYKCTRQIISEKLGRGSRTVDLELEAQIDILRDNKKKYENILKLAQ TLSTQLFQMVHTQRQLGDAFADLSLKSLELHEEFGYNADTQKLLAKNGETLLGAINFFIA SVNTLVNKTIEDTLMTVKQYESARIEYDAYRTDLEELNLGPRDANTLPKIEQSQHLFQAH KEKYDKMRNDVSVKLKFLEENKVKVLHNQLVLFHNAIAAYFAGNQKQLEQTLKQFHIKLK TPGVDAPSWLEEQ |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Golgi apparatus
|
|||||
| Function |
Plays a role in controlling biogenesis of secretory granules at the trans-Golgi network. Mechanistically, binds ARF-GTP at the neck of a growing secretory granule precursor and forms a protective scaffold. Once the granule precursor has been completely loaded, active PRKD1 phosphorylates ARFIP1 and releases it from ARFs. In turn, ARFs induce fission. Through this mechanism, ensures proper secretory granule formation at the Golgi of pancreatic beta cells.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
ONAyne Probe Info |
 |
K146(1.43) | LDD0274 | [2] | |
|
Jackson_1 Probe Info |
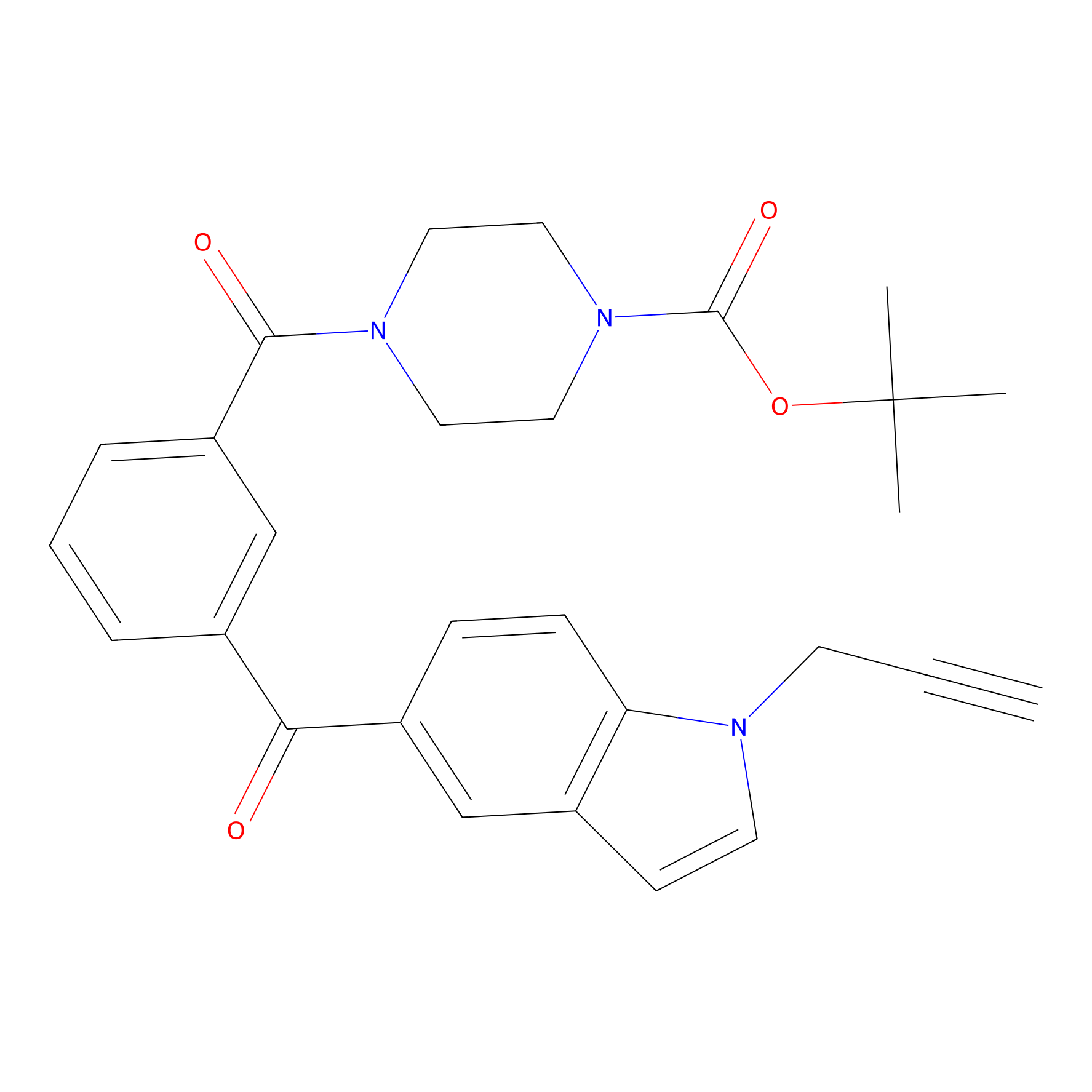 |
4.31 | LDD0120 | [3] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0221 | [4] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [5] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [6] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [6] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [7] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [8] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C027 Probe Info |
 |
5.31 | LDD1733 | [9] | |
|
C094 Probe Info |
 |
46.53 | LDD1785 | [9] | |
|
C235 Probe Info |
 |
26.54 | LDD1908 | [9] | |
|
C258 Probe Info |
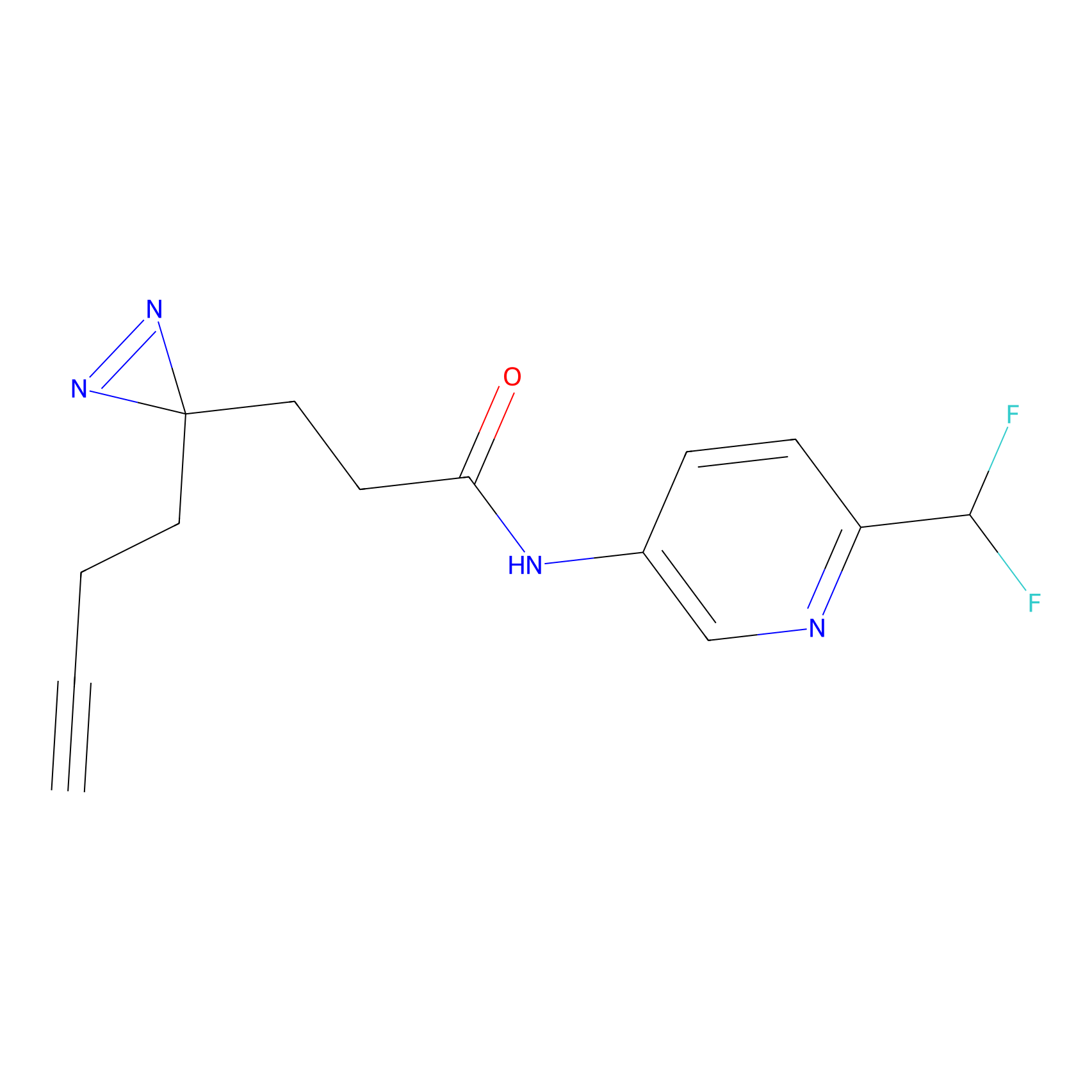 |
5.50 | LDD1931 | [9] | |
|
C299 Probe Info |
 |
13.83 | LDD1968 | [9] | |
|
C320 Probe Info |
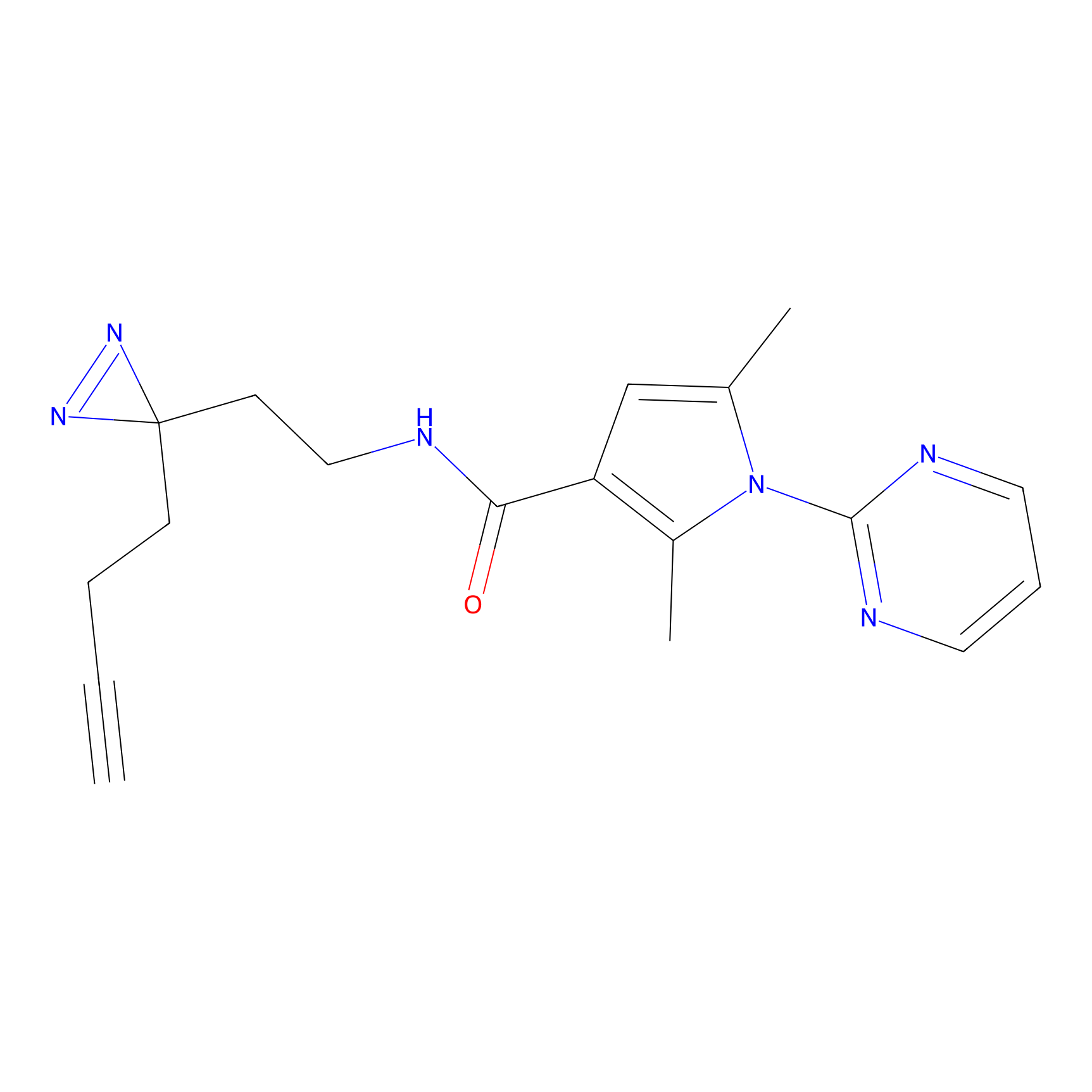 |
6.54 | LDD1986 | [9] | |
|
C361 Probe Info |
 |
23.43 | LDD2022 | [9] | |
|
C383 Probe Info |
 |
18.13 | LDD2042 | [9] | |
|
C391 Probe Info |
 |
14.93 | LDD2050 | [9] | |
|
FFF probe2 Probe Info |
 |
6.22 | LDD0463 | [10] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0464 | [10] | |
|
Alk-rapa Probe Info |
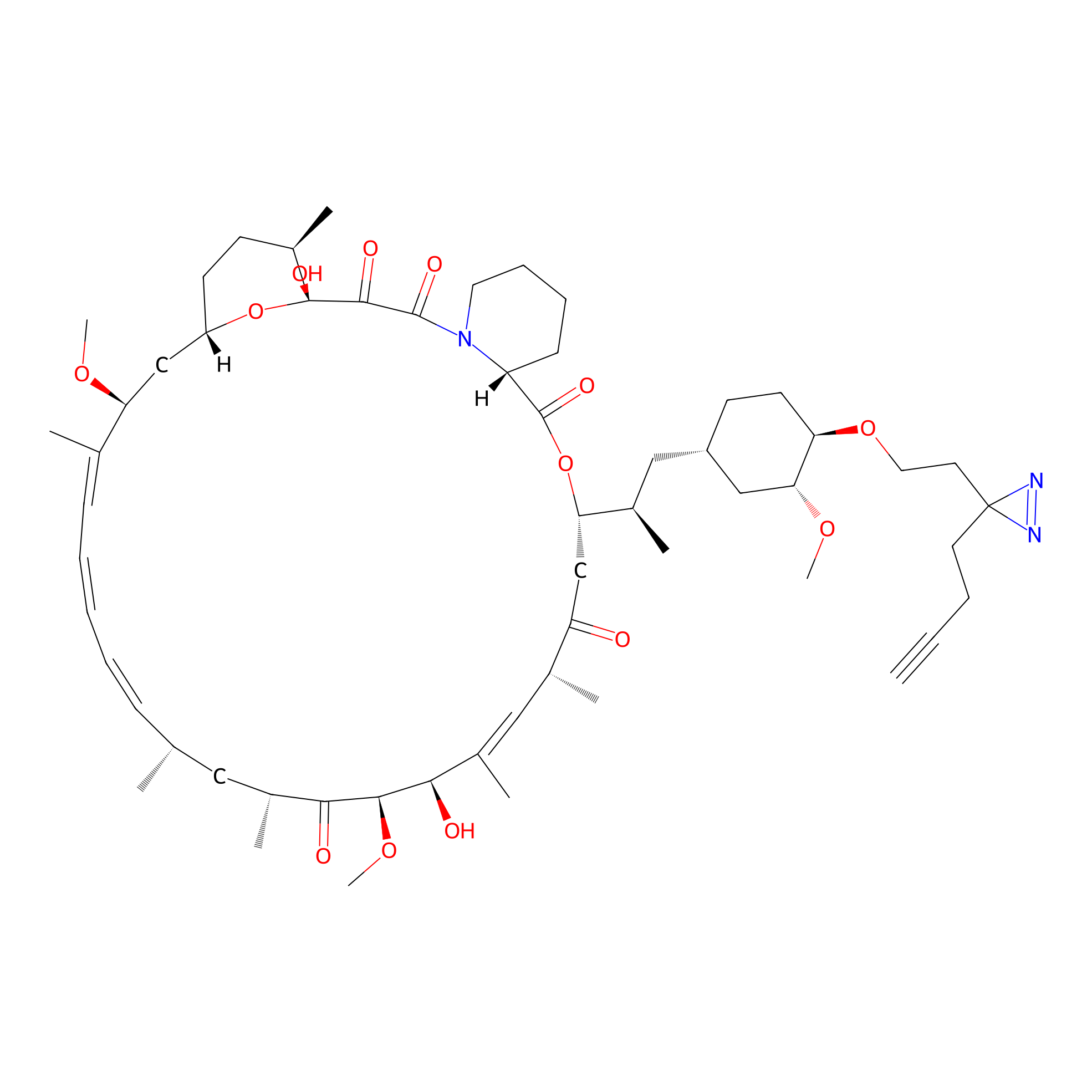 |
11.51 | LDD0213 | [11] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0156 | Aniline | NCI-H1299 | 11.40 | LDD0403 | [1] |
| LDCM0070 | Cisar_cp37 | MDA-MB-231 | 4.31 | LDD0120 | [3] |
| LDCM0191 | Compound 21 | HEK-293T | 12.27 | LDD0508 | [10] |
| LDCM0192 | Compound 35 | HEK-293T | 6.17 | LDD0509 | [10] |
| LDCM0193 | Compound 36 | HEK-293T | 9.43 | LDD0511 | [10] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [4] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [4] |
| LDCM0090 | Rapamycin | JHH-7 | 11.51 | LDD0213 | [11] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Secretory carrier-associated membrane protein 1 (SCAMP1) | SCAMP family | O15126 | |||
| Barttin (BSND) | . | Q8WZ55 | |||
Cytokine and receptor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Tumor necrosis factor receptor superfamily member 10D (TNFRSF10D) | . | Q9UBN6 | |||
Other
References
