Details of the Target
General Information of Target
| Target ID | LDTP00372 | |||||
|---|---|---|---|---|---|---|
| Target Name | Acyl carrier protein, mitochondrial (NDUFAB1) | |||||
| Gene Name | NDUFAB1 | |||||
| Gene ID | 4706 | |||||
| Synonyms |
Acyl carrier protein, mitochondrial; ACP; CI-SDAP; NADH-ubiquinone oxidoreductase 9.6 kDa subunit |
|||||
| 3D Structure | ||||||
| Sequence |
MASRVLSAYVSRLPAAFAPLPRVRMLAVARPLSTALCSAGTQTRLGTLQPALVLAQVPGR
VTQLCRQYSDMPPLTLEGIQDRVLYVLKLYDKIDPEKLSVNSHFMKDLGLDSLDQVEIIM AMEDEFGFEIPDIDAEKLMCPQEIVDYIADKKDVYE |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Acyl carrier protein (ACP) family
|
|||||
| Subcellular location |
Mitochondrion
|
|||||
| Function |
Carrier of the growing fatty acid chain in fatty acid biosynthesis. Accessory and non-catalytic subunit of the mitochondrial membrane respiratory chain NADH dehydrogenase (Complex I), which functions in the transfer of electrons from NADH to the respiratory chain. Accessory protein, of the core iron-sulfur cluster (ISC) assembly complex, that regulates, in association with LYRM4, the stability and the cysteine desulfurase activity of NFS1 and participates in the [2Fe-2S] clusters assembly on the scaffolding protein ISCU. The core iron-sulfur cluster (ISC) assembly complex is involved in the de novo synthesis of a [2Fe-2S] cluster, the first step of the mitochondrial iron-sulfur protein biogenesis. This process is initiated by the cysteine desulfurase complex (NFS1:LYRM4:NDUFAB1) that produces persulfide which is delivered on the scaffold protein ISCU in a FXN-dependent manner. Then this complex is stabilized by FDX2 which provides reducing equivalents to accomplish the [2Fe-2S] cluster assembly. Finally, the [2Fe-2S] cluster is transferred from ISCU to chaperone proteins, including HSCB, HSPA9 and GLRX5.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
10.25 | LDD0402 | [1] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [2] | |
|
TH211 Probe Info |
 |
Y85(20.00); Y147(20.00) | LDD0260 | [3] | |
|
STPyne Probe Info |
 |
K88(20.00) | LDD2217 | [4] | |
|
Probe 1 Probe Info |
 |
Y85(63.98) | LDD3495 | [5] | |
|
BTD Probe Info |
 |
C140(1.15) | LDD2098 | [6] | |
|
DBIA Probe Info |
 |
C140(1.31) | LDD0531 | [7] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [8] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [8] | |
|
IPIAA_H Probe Info |
 |
N.A. | LDD0030 | [9] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [9] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [8] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [10] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [10] | |
|
aHNE Probe Info |
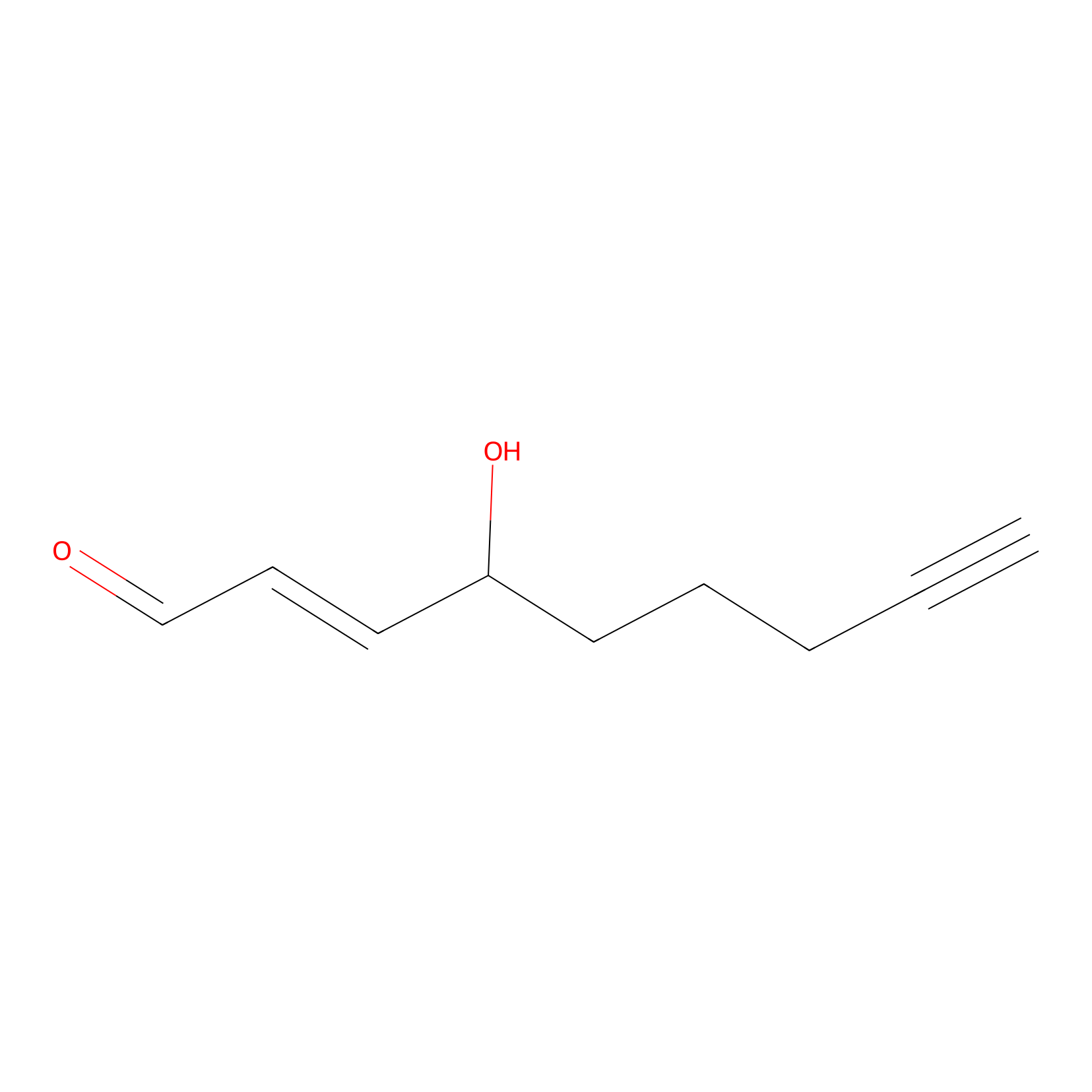 |
N.A. | LDD0001 | [10] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [10] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [11] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [12] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [10] | |
|
YN-1 Probe Info |
 |
N.A. | LDD0446 | [13] | |
|
1c-yne Probe Info |
 |
K97(0.00); K88(0.00) | LDD0228 | [14] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [15] | |
|
Crotonaldehyde Probe Info |
 |
C140(0.00); H103(0.00) | LDD0219 | [15] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [15] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [16] | |
|
AOyne Probe Info |
 |
11.70 | LDD0443 | [17] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [18] | |
|
HHS-465 Probe Info |
 |
N.A. | LDD2240 | [19] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe3 Probe Info |
 |
20.00 | LDD0465 | [20] | |
|
FFF probe6 Probe Info |
 |
20.00 | LDD0468 | [20] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C140(0.50) | LDD2142 | [6] |
| LDCM0214 | AC1 | HCT 116 | C140(1.31) | LDD0531 | [7] |
| LDCM0215 | AC10 | PaTu 8988t | C140(1.17) | LDD1094 | [7] |
| LDCM0216 | AC100 | HEK-293T | C140(1.09) | LDD0814 | [7] |
| LDCM0217 | AC101 | HEK-293T | C140(1.19) | LDD0815 | [7] |
| LDCM0218 | AC102 | HEK-293T | C140(0.92) | LDD0816 | [7] |
| LDCM0219 | AC103 | HEK-293T | C140(0.95) | LDD0817 | [7] |
| LDCM0220 | AC104 | HEK-293T | C140(0.88) | LDD0818 | [7] |
| LDCM0221 | AC105 | HEK-293T | C140(0.75) | LDD0819 | [7] |
| LDCM0222 | AC106 | HEK-293T | C140(1.23) | LDD0820 | [7] |
| LDCM0223 | AC107 | HEK-293T | C140(0.66) | LDD0821 | [7] |
| LDCM0224 | AC108 | HEK-293T | C140(0.64) | LDD0822 | [7] |
| LDCM0225 | AC109 | HEK-293T | C140(0.77) | LDD0823 | [7] |
| LDCM0226 | AC11 | PaTu 8988t | C140(1.06) | LDD1105 | [7] |
| LDCM0227 | AC110 | HEK-293T | C140(0.96) | LDD0825 | [7] |
| LDCM0228 | AC111 | HEK-293T | C140(0.78) | LDD0826 | [7] |
| LDCM0229 | AC112 | HEK-293T | C140(1.16) | LDD0827 | [7] |
| LDCM0230 | AC113 | HCT 116 | C140(1.13) | LDD0547 | [7] |
| LDCM0231 | AC114 | HCT 116 | C140(1.26) | LDD0548 | [7] |
| LDCM0232 | AC115 | HCT 116 | C140(1.46) | LDD0549 | [7] |
| LDCM0233 | AC116 | HCT 116 | C140(1.40) | LDD0550 | [7] |
| LDCM0234 | AC117 | HCT 116 | C140(1.24) | LDD0551 | [7] |
| LDCM0235 | AC118 | HCT 116 | C140(1.32) | LDD0552 | [7] |
| LDCM0236 | AC119 | HCT 116 | C140(1.36) | LDD0553 | [7] |
| LDCM0237 | AC12 | PaTu 8988t | C140(1.00) | LDD1116 | [7] |
| LDCM0238 | AC120 | HCT 116 | C140(1.27) | LDD0555 | [7] |
| LDCM0239 | AC121 | HCT 116 | C140(1.23) | LDD0556 | [7] |
| LDCM0240 | AC122 | HCT 116 | C140(1.26) | LDD0557 | [7] |
| LDCM0241 | AC123 | HCT 116 | C140(1.13) | LDD0558 | [7] |
| LDCM0242 | AC124 | HCT 116 | C140(1.08) | LDD0559 | [7] |
| LDCM0243 | AC125 | HCT 116 | C140(1.30) | LDD0560 | [7] |
| LDCM0244 | AC126 | HCT 116 | C140(1.25) | LDD0561 | [7] |
| LDCM0245 | AC127 | HCT 116 | C140(1.40) | LDD0562 | [7] |
| LDCM0246 | AC128 | HCT 116 | C140(1.26) | LDD0563 | [7] |
| LDCM0247 | AC129 | HCT 116 | C140(0.85) | LDD0564 | [7] |
| LDCM0249 | AC130 | HCT 116 | C140(1.10) | LDD0566 | [7] |
| LDCM0250 | AC131 | HCT 116 | C140(0.84) | LDD0567 | [7] |
| LDCM0251 | AC132 | HCT 116 | C140(0.99) | LDD0568 | [7] |
| LDCM0252 | AC133 | HCT 116 | C140(1.13) | LDD0569 | [7] |
| LDCM0253 | AC134 | HCT 116 | C140(1.14) | LDD0570 | [7] |
| LDCM0254 | AC135 | HCT 116 | C140(1.11) | LDD0571 | [7] |
| LDCM0255 | AC136 | HCT 116 | C140(1.11) | LDD0572 | [7] |
| LDCM0256 | AC137 | HCT 116 | C140(1.07) | LDD0573 | [7] |
| LDCM0257 | AC138 | HCT 116 | C140(1.16) | LDD0574 | [7] |
| LDCM0258 | AC139 | HCT 116 | C140(1.04) | LDD0575 | [7] |
| LDCM0259 | AC14 | PaTu 8988t | C140(1.03) | LDD1138 | [7] |
| LDCM0260 | AC140 | HCT 116 | C140(1.15) | LDD0577 | [7] |
| LDCM0261 | AC141 | HCT 116 | C140(1.09) | LDD0578 | [7] |
| LDCM0262 | AC142 | HCT 116 | C140(0.94) | LDD0579 | [7] |
| LDCM0263 | AC143 | HCT 116 | C140(1.19) | LDD0580 | [7] |
| LDCM0264 | AC144 | HCT 116 | C140(1.28) | LDD0581 | [7] |
| LDCM0265 | AC145 | HCT 116 | C140(1.20) | LDD0582 | [7] |
| LDCM0266 | AC146 | HCT 116 | C140(1.26) | LDD0583 | [7] |
| LDCM0267 | AC147 | HCT 116 | C140(1.39) | LDD0584 | [7] |
| LDCM0268 | AC148 | HCT 116 | C140(1.42) | LDD0585 | [7] |
| LDCM0269 | AC149 | HCT 116 | C140(1.48) | LDD0586 | [7] |
| LDCM0270 | AC15 | PaTu 8988t | C140(1.02) | LDD1149 | [7] |
| LDCM0271 | AC150 | HCT 116 | C140(1.08) | LDD0588 | [7] |
| LDCM0272 | AC151 | HCT 116 | C140(1.19) | LDD0589 | [7] |
| LDCM0273 | AC152 | HCT 116 | C140(1.41) | LDD0590 | [7] |
| LDCM0274 | AC153 | HCT 116 | C140(1.47) | LDD0591 | [7] |
| LDCM0621 | AC154 | HCT 116 | C140(1.34) | LDD2158 | [7] |
| LDCM0622 | AC155 | HCT 116 | C140(1.30) | LDD2159 | [7] |
| LDCM0623 | AC156 | HCT 116 | C140(0.97) | LDD2160 | [7] |
| LDCM0624 | AC157 | HCT 116 | C140(0.91) | LDD2161 | [7] |
| LDCM0276 | AC17 | PaTu 8988t | C140(0.96) | LDD1155 | [7] |
| LDCM0277 | AC18 | PaTu 8988t | C140(1.36) | LDD1156 | [7] |
| LDCM0278 | AC19 | PaTu 8988t | C140(1.15) | LDD1157 | [7] |
| LDCM0279 | AC2 | HCT 116 | C140(1.31) | LDD0596 | [7] |
| LDCM0280 | AC20 | PaTu 8988t | C140(1.19) | LDD1159 | [7] |
| LDCM0281 | AC21 | PaTu 8988t | C140(1.12) | LDD1160 | [7] |
| LDCM0282 | AC22 | PaTu 8988t | C140(1.22) | LDD1161 | [7] |
| LDCM0283 | AC23 | PaTu 8988t | C140(1.28) | LDD1162 | [7] |
| LDCM0284 | AC24 | PaTu 8988t | C140(1.22) | LDD1163 | [7] |
| LDCM0285 | AC25 | HEK-293T | C140(0.95) | LDD0883 | [7] |
| LDCM0286 | AC26 | HEK-293T | C140(1.04) | LDD0884 | [7] |
| LDCM0287 | AC27 | HEK-293T | C140(1.34) | LDD0885 | [7] |
| LDCM0288 | AC28 | HEK-293T | C140(0.83) | LDD0886 | [7] |
| LDCM0289 | AC29 | HEK-293T | C140(1.01) | LDD0887 | [7] |
| LDCM0290 | AC3 | HCT 116 | C140(1.25) | LDD0607 | [7] |
| LDCM0291 | AC30 | HEK-293T | C140(0.77) | LDD0889 | [7] |
| LDCM0292 | AC31 | HEK-293T | C140(0.97) | LDD0890 | [7] |
| LDCM0293 | AC32 | HEK-293T | C140(1.03) | LDD0891 | [7] |
| LDCM0294 | AC33 | HEK-293T | C140(1.02) | LDD0892 | [7] |
| LDCM0295 | AC34 | HEK-293T | C140(0.88) | LDD0893 | [7] |
| LDCM0296 | AC35 | PaTu 8988t | C140(1.20) | LDD1175 | [7] |
| LDCM0297 | AC36 | PaTu 8988t | C140(1.36) | LDD1176 | [7] |
| LDCM0298 | AC37 | PaTu 8988t | C140(1.39) | LDD1177 | [7] |
| LDCM0299 | AC38 | PaTu 8988t | C140(1.35) | LDD1178 | [7] |
| LDCM0300 | AC39 | PaTu 8988t | C140(1.44) | LDD1179 | [7] |
| LDCM0301 | AC4 | HCT 116 | C140(1.20) | LDD0618 | [7] |
| LDCM0302 | AC40 | PaTu 8988t | C140(1.32) | LDD1181 | [7] |
| LDCM0303 | AC41 | PaTu 8988t | C140(1.20) | LDD1182 | [7] |
| LDCM0304 | AC42 | PaTu 8988t | C140(1.23) | LDD1183 | [7] |
| LDCM0305 | AC43 | PaTu 8988t | C140(1.22) | LDD1184 | [7] |
| LDCM0306 | AC44 | PaTu 8988t | C140(1.42) | LDD1185 | [7] |
| LDCM0307 | AC45 | PaTu 8988t | C140(1.34) | LDD1186 | [7] |
| LDCM0308 | AC46 | PaTu 8988t | C140(1.11) | LDD1187 | [7] |
| LDCM0309 | AC47 | PaTu 8988t | C140(1.15) | LDD1188 | [7] |
| LDCM0310 | AC48 | PaTu 8988t | C140(1.25) | LDD1189 | [7] |
| LDCM0311 | AC49 | PaTu 8988t | C140(1.35) | LDD1190 | [7] |
| LDCM0312 | AC5 | HCT 116 | C140(1.19) | LDD0629 | [7] |
| LDCM0313 | AC50 | PaTu 8988t | C140(1.25) | LDD1192 | [7] |
| LDCM0314 | AC51 | PaTu 8988t | C140(0.86) | LDD1193 | [7] |
| LDCM0315 | AC52 | PaTu 8988t | C140(1.20) | LDD1194 | [7] |
| LDCM0316 | AC53 | PaTu 8988t | C140(1.34) | LDD1195 | [7] |
| LDCM0317 | AC54 | PaTu 8988t | C140(1.49) | LDD1196 | [7] |
| LDCM0318 | AC55 | PaTu 8988t | C140(1.25) | LDD1197 | [7] |
| LDCM0319 | AC56 | PaTu 8988t | C140(1.43) | LDD1198 | [7] |
| LDCM0320 | AC57 | PaTu 8988t | C140(1.16) | LDD1199 | [7] |
| LDCM0321 | AC58 | PaTu 8988t | C140(1.15) | LDD1200 | [7] |
| LDCM0322 | AC59 | PaTu 8988t | C140(1.03) | LDD1201 | [7] |
| LDCM0323 | AC6 | PaTu 8988t | C140(1.05) | LDD1202 | [7] |
| LDCM0324 | AC60 | PaTu 8988t | C140(1.18) | LDD1203 | [7] |
| LDCM0325 | AC61 | PaTu 8988t | C140(1.05) | LDD1204 | [7] |
| LDCM0326 | AC62 | PaTu 8988t | C140(1.09) | LDD1205 | [7] |
| LDCM0327 | AC63 | PaTu 8988t | C140(1.03) | LDD1206 | [7] |
| LDCM0328 | AC64 | PaTu 8988t | C140(1.02) | LDD1207 | [7] |
| LDCM0329 | AC65 | PaTu 8988t | C140(1.07) | LDD1208 | [7] |
| LDCM0330 | AC66 | PaTu 8988t | C140(1.07) | LDD1209 | [7] |
| LDCM0331 | AC67 | PaTu 8988t | C140(1.12) | LDD1210 | [7] |
| LDCM0332 | AC68 | HCT 116 | C140(1.09) | LDD0649 | [7] |
| LDCM0333 | AC69 | HCT 116 | C140(1.00) | LDD0650 | [7] |
| LDCM0334 | AC7 | PaTu 8988t | C140(0.95) | LDD1213 | [7] |
| LDCM0335 | AC70 | HCT 116 | C140(0.89) | LDD0652 | [7] |
| LDCM0336 | AC71 | HCT 116 | C140(1.04) | LDD0653 | [7] |
| LDCM0337 | AC72 | HCT 116 | C140(1.05) | LDD0654 | [7] |
| LDCM0338 | AC73 | HCT 116 | C140(1.00) | LDD0655 | [7] |
| LDCM0339 | AC74 | HCT 116 | C140(0.96) | LDD0656 | [7] |
| LDCM0340 | AC75 | HCT 116 | C140(0.96) | LDD0657 | [7] |
| LDCM0341 | AC76 | HCT 116 | C140(1.06) | LDD0658 | [7] |
| LDCM0342 | AC77 | HCT 116 | C140(1.06) | LDD0659 | [7] |
| LDCM0343 | AC78 | HCT 116 | C140(0.92) | LDD0660 | [7] |
| LDCM0344 | AC79 | HCT 116 | C140(0.95) | LDD0661 | [7] |
| LDCM0345 | AC8 | PaTu 8988t | C140(0.92) | LDD1224 | [7] |
| LDCM0346 | AC80 | HCT 116 | C140(1.04) | LDD0663 | [7] |
| LDCM0347 | AC81 | HCT 116 | C140(0.71) | LDD0664 | [7] |
| LDCM0348 | AC82 | HCT 116 | C140(0.82) | LDD0665 | [7] |
| LDCM0349 | AC83 | PaTu 8988t | C140(0.81) | LDD1228 | [7] |
| LDCM0350 | AC84 | PaTu 8988t | C140(0.90) | LDD1229 | [7] |
| LDCM0351 | AC85 | PaTu 8988t | C140(0.96) | LDD1230 | [7] |
| LDCM0352 | AC86 | PaTu 8988t | C140(0.93) | LDD1231 | [7] |
| LDCM0353 | AC87 | PaTu 8988t | C140(0.86) | LDD1232 | [7] |
| LDCM0354 | AC88 | PaTu 8988t | C140(0.81) | LDD1233 | [7] |
| LDCM0355 | AC89 | PaTu 8988t | C140(0.86) | LDD1234 | [7] |
| LDCM0357 | AC90 | PaTu 8988t | C140(0.80) | LDD1236 | [7] |
| LDCM0358 | AC91 | PaTu 8988t | C140(0.84) | LDD1237 | [7] |
| LDCM0359 | AC92 | PaTu 8988t | C140(0.93) | LDD1238 | [7] |
| LDCM0360 | AC93 | PaTu 8988t | C140(0.77) | LDD1239 | [7] |
| LDCM0361 | AC94 | PaTu 8988t | C140(0.80) | LDD1240 | [7] |
| LDCM0362 | AC95 | PaTu 8988t | C140(0.87) | LDD1241 | [7] |
| LDCM0363 | AC96 | PaTu 8988t | C140(0.81) | LDD1242 | [7] |
| LDCM0364 | AC97 | PaTu 8988t | C140(0.74) | LDD1243 | [7] |
| LDCM0365 | AC98 | HEK-293T | C140(1.20) | LDD0963 | [7] |
| LDCM0366 | AC99 | HEK-293T | C140(0.90) | LDD0964 | [7] |
| LDCM0248 | AKOS034007472 | PaTu 8988t | C140(1.14) | LDD1127 | [7] |
| LDCM0356 | AKOS034007680 | PaTu 8988t | C140(1.22) | LDD1235 | [7] |
| LDCM0275 | AKOS034007705 | PaTu 8988t | C140(1.13) | LDD1154 | [7] |
| LDCM0156 | Aniline | NCI-H1299 | 11.00 | LDD0403 | [1] |
| LDCM0108 | Chloroacetamide | HeLa | C140(0.00); H103(0.00) | LDD0222 | [15] |
| LDCM0632 | CL-Sc | Hep-G2 | C140(1.26) | LDD2227 | [18] |
| LDCM0367 | CL1 | PaTu 8988t | C140(1.15) | LDD1246 | [7] |
| LDCM0368 | CL10 | PaTu 8988t | C140(1.22) | LDD1247 | [7] |
| LDCM0369 | CL100 | HCT 116 | C140(1.16) | LDD0686 | [7] |
| LDCM0370 | CL101 | PaTu 8988t | C140(0.99) | LDD1249 | [7] |
| LDCM0371 | CL102 | PaTu 8988t | C140(1.06) | LDD1250 | [7] |
| LDCM0372 | CL103 | PaTu 8988t | C140(1.21) | LDD1251 | [7] |
| LDCM0373 | CL104 | PaTu 8988t | C140(1.13) | LDD1252 | [7] |
| LDCM0374 | CL105 | PaTu 8988t | C140(1.07) | LDD1253 | [7] |
| LDCM0375 | CL106 | PaTu 8988t | C140(0.99) | LDD1254 | [7] |
| LDCM0376 | CL107 | PaTu 8988t | C140(1.05) | LDD1255 | [7] |
| LDCM0377 | CL108 | PaTu 8988t | C140(1.11) | LDD1256 | [7] |
| LDCM0378 | CL109 | PaTu 8988t | C140(1.19) | LDD1257 | [7] |
| LDCM0379 | CL11 | PaTu 8988t | C140(1.05) | LDD1258 | [7] |
| LDCM0380 | CL110 | PaTu 8988t | C140(1.15) | LDD1259 | [7] |
| LDCM0381 | CL111 | PaTu 8988t | C140(1.24) | LDD1260 | [7] |
| LDCM0382 | CL112 | HEK-293T | C140(1.19) | LDD0980 | [7] |
| LDCM0383 | CL113 | HEK-293T | C140(0.98) | LDD0981 | [7] |
| LDCM0384 | CL114 | HEK-293T | C140(1.08) | LDD0982 | [7] |
| LDCM0385 | CL115 | HEK-293T | C140(0.82) | LDD0983 | [7] |
| LDCM0386 | CL116 | HEK-293T | C140(1.04) | LDD0984 | [7] |
| LDCM0387 | CL117 | PaTu 8988t | C140(1.10) | LDD1266 | [7] |
| LDCM0388 | CL118 | PaTu 8988t | C140(1.14) | LDD1267 | [7] |
| LDCM0389 | CL119 | PaTu 8988t | C140(1.24) | LDD1268 | [7] |
| LDCM0390 | CL12 | PaTu 8988t | C140(1.07) | LDD1269 | [7] |
| LDCM0391 | CL120 | PaTu 8988t | C140(1.19) | LDD1270 | [7] |
| LDCM0392 | CL121 | PaTu 8988t | C140(1.14) | LDD1271 | [7] |
| LDCM0393 | CL122 | PaTu 8988t | C140(1.45) | LDD1272 | [7] |
| LDCM0394 | CL123 | PaTu 8988t | C140(1.37) | LDD1273 | [7] |
| LDCM0395 | CL124 | PaTu 8988t | C140(1.37) | LDD1274 | [7] |
| LDCM0396 | CL125 | PaTu 8988t | C140(1.03) | LDD1275 | [7] |
| LDCM0397 | CL126 | PaTu 8988t | C140(0.99) | LDD1276 | [7] |
| LDCM0398 | CL127 | PaTu 8988t | C140(0.94) | LDD1277 | [7] |
| LDCM0399 | CL128 | PaTu 8988t | C140(1.12) | LDD1278 | [7] |
| LDCM0400 | CL13 | PaTu 8988t | C140(1.24) | LDD1279 | [7] |
| LDCM0401 | CL14 | PaTu 8988t | C140(1.12) | LDD1280 | [7] |
| LDCM0402 | CL15 | PaTu 8988t | C140(1.02) | LDD1281 | [7] |
| LDCM0403 | CL16 | HCT 116 | C140(1.20) | LDD0720 | [7] |
| LDCM0404 | CL17 | HCT 116 | C140(1.19) | LDD0721 | [7] |
| LDCM0405 | CL18 | HCT 116 | C140(1.22) | LDD0722 | [7] |
| LDCM0406 | CL19 | HCT 116 | C140(1.46) | LDD0723 | [7] |
| LDCM0407 | CL2 | PaTu 8988t | C140(1.18) | LDD1286 | [7] |
| LDCM0408 | CL20 | HCT 116 | C140(1.43) | LDD0725 | [7] |
| LDCM0409 | CL21 | HCT 116 | C140(1.41) | LDD0726 | [7] |
| LDCM0410 | CL22 | HCT 116 | C140(1.79) | LDD0727 | [7] |
| LDCM0411 | CL23 | HCT 116 | C140(1.39) | LDD0728 | [7] |
| LDCM0412 | CL24 | HCT 116 | C140(1.55) | LDD0729 | [7] |
| LDCM0413 | CL25 | HCT 116 | C140(1.60) | LDD0730 | [7] |
| LDCM0414 | CL26 | HCT 116 | C140(1.24) | LDD0731 | [7] |
| LDCM0415 | CL27 | HCT 116 | C140(1.44) | LDD0732 | [7] |
| LDCM0416 | CL28 | HCT 116 | C140(1.48) | LDD0733 | [7] |
| LDCM0417 | CL29 | HCT 116 | C140(1.46) | LDD0734 | [7] |
| LDCM0418 | CL3 | PaTu 8988t | C140(1.51) | LDD1297 | [7] |
| LDCM0419 | CL30 | HCT 116 | C140(1.19) | LDD0736 | [7] |
| LDCM0420 | CL31 | HEK-293T | C140(1.60) | LDD1018 | [7] |
| LDCM0421 | CL32 | HEK-293T | C140(1.46) | LDD1019 | [7] |
| LDCM0422 | CL33 | HEK-293T | C140(1.57) | LDD1020 | [7] |
| LDCM0423 | CL34 | HEK-293T | C140(1.36) | LDD1021 | [7] |
| LDCM0424 | CL35 | HEK-293T | C140(1.50) | LDD1022 | [7] |
| LDCM0425 | CL36 | HEK-293T | C140(1.80) | LDD1023 | [7] |
| LDCM0426 | CL37 | HEK-293T | C140(1.41) | LDD1024 | [7] |
| LDCM0428 | CL39 | HEK-293T | C140(1.43) | LDD1026 | [7] |
| LDCM0429 | CL4 | PaTu 8988t | C140(1.27) | LDD1308 | [7] |
| LDCM0430 | CL40 | HEK-293T | C140(1.38) | LDD1028 | [7] |
| LDCM0431 | CL41 | HEK-293T | C140(1.32) | LDD1029 | [7] |
| LDCM0432 | CL42 | HEK-293T | C140(1.39) | LDD1030 | [7] |
| LDCM0433 | CL43 | HEK-293T | C140(1.61) | LDD1031 | [7] |
| LDCM0434 | CL44 | HEK-293T | C140(1.30) | LDD1032 | [7] |
| LDCM0435 | CL45 | HEK-293T | C140(1.54) | LDD1033 | [7] |
| LDCM0436 | CL46 | HCT 116 | C140(0.94) | LDD0753 | [7] |
| LDCM0437 | CL47 | HCT 116 | C140(1.03) | LDD0754 | [7] |
| LDCM0438 | CL48 | HCT 116 | C140(0.93) | LDD0755 | [7] |
| LDCM0439 | CL49 | HCT 116 | C140(0.93) | LDD0756 | [7] |
| LDCM0440 | CL5 | PaTu 8988t | C140(1.41) | LDD1319 | [7] |
| LDCM0441 | CL50 | HCT 116 | C140(0.85) | LDD0758 | [7] |
| LDCM0442 | CL51 | HCT 116 | C140(1.08) | LDD0759 | [7] |
| LDCM0443 | CL52 | HCT 116 | C140(1.21) | LDD0760 | [7] |
| LDCM0444 | CL53 | HCT 116 | C140(0.80) | LDD0761 | [7] |
| LDCM0445 | CL54 | HCT 116 | C140(0.93) | LDD0762 | [7] |
| LDCM0446 | CL55 | HCT 116 | C140(1.00) | LDD0763 | [7] |
| LDCM0447 | CL56 | HCT 116 | C140(1.04) | LDD0764 | [7] |
| LDCM0448 | CL57 | HCT 116 | C140(0.97) | LDD0765 | [7] |
| LDCM0449 | CL58 | HCT 116 | C140(0.93) | LDD0766 | [7] |
| LDCM0450 | CL59 | HCT 116 | C140(0.96) | LDD0767 | [7] |
| LDCM0451 | CL6 | PaTu 8988t | C140(1.26) | LDD1330 | [7] |
| LDCM0452 | CL60 | HCT 116 | C140(0.90) | LDD0769 | [7] |
| LDCM0453 | CL61 | HCT 116 | C140(1.09) | LDD0770 | [7] |
| LDCM0454 | CL62 | HCT 116 | C140(1.11) | LDD0771 | [7] |
| LDCM0455 | CL63 | HCT 116 | C140(1.15) | LDD0772 | [7] |
| LDCM0456 | CL64 | HCT 116 | C140(1.19) | LDD0773 | [7] |
| LDCM0457 | CL65 | HCT 116 | C140(1.12) | LDD0774 | [7] |
| LDCM0458 | CL66 | HCT 116 | C140(1.37) | LDD0775 | [7] |
| LDCM0459 | CL67 | HCT 116 | C140(1.18) | LDD0776 | [7] |
| LDCM0460 | CL68 | HCT 116 | C140(1.05) | LDD0777 | [7] |
| LDCM0461 | CL69 | HCT 116 | C140(0.85) | LDD0778 | [7] |
| LDCM0462 | CL7 | PaTu 8988t | C140(1.27) | LDD1341 | [7] |
| LDCM0463 | CL70 | HCT 116 | C140(1.06) | LDD0780 | [7] |
| LDCM0464 | CL71 | HCT 116 | C140(0.99) | LDD0781 | [7] |
| LDCM0465 | CL72 | HCT 116 | C140(0.94) | LDD0782 | [7] |
| LDCM0466 | CL73 | HCT 116 | C140(1.10) | LDD0783 | [7] |
| LDCM0467 | CL74 | HCT 116 | C140(1.07) | LDD0784 | [7] |
| LDCM0469 | CL76 | HCT 116 | C140(1.09) | LDD0786 | [7] |
| LDCM0470 | CL77 | HCT 116 | C140(0.92) | LDD0787 | [7] |
| LDCM0471 | CL78 | HCT 116 | C140(1.16) | LDD0788 | [7] |
| LDCM0472 | CL79 | HCT 116 | C140(1.35) | LDD0789 | [7] |
| LDCM0473 | CL8 | PaTu 8988t | C140(1.24) | LDD1352 | [7] |
| LDCM0474 | CL80 | HCT 116 | C140(1.07) | LDD0791 | [7] |
| LDCM0475 | CL81 | HCT 116 | C140(1.39) | LDD0792 | [7] |
| LDCM0476 | CL82 | HCT 116 | C140(1.47) | LDD0793 | [7] |
| LDCM0477 | CL83 | HCT 116 | C140(1.47) | LDD0794 | [7] |
| LDCM0478 | CL84 | HCT 116 | C140(1.37) | LDD0795 | [7] |
| LDCM0479 | CL85 | HCT 116 | C140(1.20) | LDD0796 | [7] |
| LDCM0480 | CL86 | HCT 116 | C140(1.06) | LDD0797 | [7] |
| LDCM0481 | CL87 | HCT 116 | C140(1.24) | LDD0798 | [7] |
| LDCM0482 | CL88 | HCT 116 | C140(1.62) | LDD0799 | [7] |
| LDCM0483 | CL89 | HCT 116 | C140(1.45) | LDD0800 | [7] |
| LDCM0484 | CL9 | PaTu 8988t | C140(1.22) | LDD1363 | [7] |
| LDCM0485 | CL90 | HCT 116 | C140(0.89) | LDD0802 | [7] |
| LDCM0486 | CL91 | HCT 116 | C140(1.36) | LDD0803 | [7] |
| LDCM0487 | CL92 | HCT 116 | C140(1.18) | LDD0804 | [7] |
| LDCM0488 | CL93 | HCT 116 | C140(1.05) | LDD0805 | [7] |
| LDCM0489 | CL94 | HCT 116 | C140(0.99) | LDD0806 | [7] |
| LDCM0490 | CL95 | HCT 116 | C140(1.06) | LDD0807 | [7] |
| LDCM0491 | CL96 | HCT 116 | C140(1.25) | LDD0808 | [7] |
| LDCM0492 | CL97 | HCT 116 | C140(1.18) | LDD0809 | [7] |
| LDCM0493 | CL98 | HCT 116 | C140(1.27) | LDD0810 | [7] |
| LDCM0494 | CL99 | HCT 116 | C140(1.60) | LDD0811 | [7] |
| LDCM0495 | E2913 | HEK-293T | C140(1.16) | LDD1698 | [21] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C140(0.89) | LDD1702 | [6] |
| LDCM0625 | F8 | Ramos | C140(0.74) | LDD2187 | [22] |
| LDCM0572 | Fragment10 | Ramos | C140(0.84) | LDD2189 | [22] |
| LDCM0573 | Fragment11 | Ramos | C140(0.97) | LDD2190 | [22] |
| LDCM0574 | Fragment12 | Ramos | C140(0.96) | LDD2191 | [22] |
| LDCM0575 | Fragment13 | Ramos | C140(1.06) | LDD2192 | [22] |
| LDCM0576 | Fragment14 | Ramos | C140(1.01) | LDD2193 | [22] |
| LDCM0579 | Fragment20 | Ramos | C140(0.84) | LDD2194 | [22] |
| LDCM0580 | Fragment21 | Ramos | C140(1.26) | LDD2195 | [22] |
| LDCM0582 | Fragment23 | Ramos | C140(1.02) | LDD2196 | [22] |
| LDCM0578 | Fragment27 | Ramos | C140(1.56) | LDD2197 | [22] |
| LDCM0586 | Fragment28 | Ramos | C140(0.48) | LDD2198 | [22] |
| LDCM0588 | Fragment30 | Ramos | C140(1.18) | LDD2199 | [22] |
| LDCM0589 | Fragment31 | Ramos | C140(1.38) | LDD2200 | [22] |
| LDCM0590 | Fragment32 | Ramos | C140(1.11) | LDD2201 | [22] |
| LDCM0468 | Fragment33 | HCT 116 | C140(1.13) | LDD0785 | [7] |
| LDCM0596 | Fragment38 | Ramos | C140(1.48) | LDD2203 | [22] |
| LDCM0566 | Fragment4 | Ramos | C140(0.63) | LDD2184 | [22] |
| LDCM0427 | Fragment51 | HEK-293T | C140(2.18) | LDD1025 | [7] |
| LDCM0610 | Fragment52 | Ramos | C140(1.08) | LDD2204 | [22] |
| LDCM0614 | Fragment56 | Ramos | C140(1.32) | LDD2205 | [22] |
| LDCM0569 | Fragment7 | Ramos | C140(0.52) | LDD2186 | [22] |
| LDCM0571 | Fragment9 | Ramos | C140(0.67) | LDD2188 | [22] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [15] |
| LDCM0022 | KB02 | HEK-293T | C140(0.99) | LDD1492 | [21] |
| LDCM0023 | KB03 | HEK-293T | C140(1.07) | LDD1497 | [21] |
| LDCM0024 | KB05 | HMCB | C140(0.82) | LDD3312 | [23] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0224 | [15] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C140(1.15) | LDD2098 | [6] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C140(0.90) | LDD2107 | [6] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C140(0.97) | LDD2109 | [6] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C140(0.29) | LDD2114 | [6] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C140(0.88) | LDD2137 | [6] |
| LDCM0131 | RA190 | MM1.R | C140(1.07) | LDD0304 | [24] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transcription factor
Cytokine and receptor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Ciliary neurotrophic factor (CNTF) | CNTF family | P26441 | |||
Other
References
