Details of the Target
General Information of Target
| Target ID | LDTP07283 | |||||
|---|---|---|---|---|---|---|
| Target Name | E3 ubiquitin-protein ligase RNF213 (RNF213) | |||||
| Gene Name | RNF213 | |||||
| Gene ID | 57674 | |||||
| Synonyms |
ALO17; C17orf27; KIAA1554; KIAA1618; MYSTR; E3 ubiquitin-protein ligase RNF213; EC 2.3.2.27; EC 3.6.4.-; ALK lymphoma oligomerization partner on chromosome 17; E3 ubiquitin-lipopolysaccharide ligase RNF213; EC 2.3.2.-; Mysterin; RING finger protein 213
|
|||||
| 3D Structure | ||||||
| Sequence |
MECPSCQHVSKEETPKFCSQCGERLPPAAPIADSENNNSTMASASEGEMECGQELKEEGG
PCLFPGSDSWQENPEEPCSKASWTVQESKKKKRKKKKKGNKSASSELASLPLSPASPCHL TLLSNPWPQDTALPHSQAQQSGPTGQPSQPPGTATTPLEGDGLSAPTEVGDSPLQAQALG EAGVATGSEAQSSPQFQDHTEGEDQDASIPSGGRGLSQEGTGPPTSAGEGHSRTEDAAQE LLLPESKGGSSEPGTELQTTEQQAGASASMAVDAVAEPANAVKGAGKEMKEKTQRMKQPP ATTPPFKTHCQEAETKTKDEMAAAEEKVGKNEQGEPEDLKKPEGKNRSAAAVKNEKEQKN QEADVQEVKASTLSPGGGVTVFFHAIISLHFPFNPDLHKVFIRGGEEFGESKWDSNICEL HYTRDLGHDRVLVEGIVCISKKHLDKYIPYKYVIYNGESFEYEFIYKHQQKKGEYVNRCL FIKSSLLGSGDWHQYYDIVYMKPHGRLQKVMNHITDGPRKDLVKGKQIAAALMLDSTFSI LQTWDTINLNSFFTQFEQFCFVLQQPMIYEGQAQLWTDLQYREKEVKRYLWQHLKKHVVP LPDGKSTDFLPVDCPVRSKLKTGLIVLFVVEKIELLLEGSLDWLCHLLTSDASSPDEFHR DLSHILGIPQSWRLYLVNLCQRCMDTRTYTWLGALPVLHCCMELAPRHKDAWRQPEDTWA ALEGLSFSPFREQMLDTSSLLQFMREKQHLLSIDEPLFRSWFSLLPLSHLVMYMENFIEH LGRFPAHILDCLSGIYYRLPGLEQVLNTQDVQDVQNVQNILEMLLRLLDTYRDKIPEEAL SPSYLTVCLKLHEAICSSTKLLKFYELPALSAEIVCRMIRLLSLVDSAGQRDETGNNSVQ TVFQGTLAATKRWLREVFTKNMLTSSGASFTYVKEIEVWRRLVEIQFPAEHGWKESLLGD MEWRLTKEEPLSQITAYCNSCWDTKGLEDSVAKTFEKCIIEAVSSACQSQTSILQGFSYS DLRKFGIVLSAVITKSWPRTADNFNDILKHLLTLADVKHVFRLCGTDEKILANVTEDAKR LIAVADSVLTKVVGDLLSGTILVGQLELIIKHKNQFLDIWQLREKSLSPQDEQCAVEEAL DWRREELLLLKKEKRCVDSLLKMCGNVKHLIQVDFGVLAVRHSQDLSSKRLNDTVTVRLS TSSNSQRATHYHLSSQVQEMAGKIDLLRDSHIFQLFWREAAEPLSEPKEDQEAAELLSEP EEESERHILELEEVYDYLYQPSYRKFIKLHQDLKSGEVTLAEIDVIFKDFVNKYTDLDSE LKIMCTVDHQDQRDWIKDRVEQIKEYHHLHQAVHAAKVILQVKESLGLNGDFSVLNTLLN FTDNFDDFRRETLDQINQELIQAKKLLQDISEARCKGLQALSLRKEFICWVREALGGINE LKVFVDLASISAGENDIDVDRVACFHDAVQGYASLLFKLDPSVDFSAFMKHLKKLWKALD KDQYLPRKLCDSARNLEWLKTVNESHGSVERSSLTLATAINQRGIYVIQAPKGGQKISPD TVLHLILPESPGSHEESREYSLEEVKELLNKLMLMSGKKDRNNTEVERFSEVFCSVQRLS QAFIDLHSAGNMLFRTWIAMAYCSPKQGVSLQMDFGLDLVTELKEGGDVTELLAALCRQM EHFLDSWKRFVTQKRMEHFYLNFYTAEQLVYLSTELRKQPPSDAALTMLSFIKSNCTLRD VLRASVGCGSEAARYRMRRVMEELPLMLLSEFSLVDKLRIIMEQSMRCLPAFLPDCLDLE TLGHCLAHLAGMGGSPVERCLPRGLQVGQPNLVVCGHSEVLPAALAVYMQTPSQPLPTYD EVLLCTPATTFEEVALLLRRCLTLGSLGHKVYSLLFADQLSYEVARQAEELFHNLCTQQH REDYQLVMVCDGDWEHCYLPSAFSQHKVFVTPQAPLEAIQAYLAGHYRVPKQTLSAAAVF NDRLCVGIVASERAGVGKSLYVKRLHDKMKMQLNVKNVPLKTIRLIDPQVDESRVLGALL PFLDAQYQKVPVLFHLDVTSSVQTGIWVFLFKLLILQYLMDINGKMWLRNPCHLYIVEIL ERRTSVPSRSSSALRTRVPQFSFLDIFPKVTCRPPKEVIDMELSALRSDTEPGMDLWEFC SETFQRPYQYLRRFNQNQDLDTFQYQEGSVEGTPEECLQHFLFHCGVINPSWSELRNFAR FLNYQLRDCEASLFCNPSFIGDTLRGFKKFVVTFMIFMARDFATPSLHTSDQSPGKHMVT MDGVREEDLAPFSLRKRWESEPHPYVFFNDDHTTMTFIGFHLQPNINGSVDAISHLTGKV IKRDVMTRDLYQGLLLQRVPFNVDFDKLPRHKKLERLCLTLGIPQATDPDKTYELTTDNM LKILAIEMRFRCGIPVIIMGETGCGKTRLIKFLSDLRRGGTNADTIKLVKVHGGTTADMI YSRVREAENVAFANKDQHQLDTILFFDEANTTEAISCIKEVLCDHMVDGQPLAEDSGLHI IAACNPYRKHSEEMICRLESAGLGYRVSMEETADRLGSIPLRQLVYRVHALPPSLIPLVW DFGQLSDVAEKLYIQQIVQRLVESISLDENGTRVITEVLCASQGFMRKTEDECSFVSLRD VERCVKVFRWFHEHSAMLLAQLNAFLSKSSVSKNHTERDPVLWSLMLAIGVCYHASLEKK DSYRKAIARFFPKPYDDSRLLLDEITRAQDLFLDGVPLRKTIAKNLALKENVFMMVVCIE LKIPLFLVGKPGSSKSLAKTIVADAMQGPAAYSDLFRSLKQVHLVSFQCSPHSTPQGIIS TFRQCARFQQGKDLQQYVSVVVLDEVGLAEDSPKMPLKTLHPLLEDGCIEDDPAPHKKVG FVGISNWALDPAKMNRGIFVSRGSPNETELIESAKGICSSDILVQDRVQGYFASFAKAYE TVCKRQDKEFFGLRDYYSLIKMVFAAAKASNRKPSPQDIAQAVLRNFSGKDDIQALDIFL ANLPEAKCSEEVSPMQLIKQNIFGPSQKVPGGEQEDAESRYLLVLTKNYVALQILQQTFF EGDQQPEIIFGSGFPKDQEYTQLCRNINRVKICMETGKMVLLLNLQNLYESLYDALNQYY VHLGGQKYVDLGLGTHRVKCRVHPNFRLIVIEEKDVVYKHFPIPLINRLEKHYLDINTVL EKWQKSIVEELCAWVEKFINVKAHHFQKRHKYSPSDVFIGYHSDACASVVLQVIERQGPR ALTEELHQKVSEEAKSILLNCATPDAVVRLSAYSLGGFAAEWLSQEYFHRQRHNSFADFL QAHLHTADLERHAIFTEITTFSRLLTSHDCEILESEVTGRAPKPTLLWLQQFDTEYSFLK EVRNCLTNTAKCKILIFQTDFEDGIRSAQLIASAKYSVINEINKIRENEDRIFVYFITKL SRVGRGTAYVGFHGGLWQSVHIDDLRRSTLMVSDVTRLQHVTISQLFAPGDLPELGLEHR AEDGHEEAMETEASTSGEVAEVAEEAMETESSEKVGKETSELGGSDVSILDTTRLLRSCV QSAVGMLRDQNESCTRNMRRVVLLLGLLNEDDACHASFLRVSKMRLSVFLKKQEESQFHP LEWLAREACNQDALQEAGTFRHTLWKRVQGAVTPLLASMISFIDRDGNLELLTRPDTPPW ARDLWMFIFSDTMLLNIPLVMNNERHKGEMAYIVVQNHMNLSENASNNVPFSWKIKDYLE ELWVQAQYITDAEGLPKKFVDIFQQTPLGRFLAQLHGEPQQELLQCYLKDFILLTMRVST EEELKFLQMALWSCTRKLKAASEAPEEEVSLPWVHLAYQRFRSRLQNFSRILTIYPQVLH SLMEARWNHELAGCEMTLDAFAAMACTEMLTRNTLKPSPQAWLQLVKNLSMPLELICSDE HMQGSGSLAQAVIREVRAQWSRIFSTALFVEHVLLGTESRVPELQGLVTEHVFLLDKCLR ENSDVKTHGPFEAVMRTLCECKETASKTLSRFGIQPCSICLGDAKDPVCLPCDHVHCLRC LRAWFASEQMICPYCLTALPDEFSPAVSQAHREAIEKHARFRQMCNSFFVDLVSTICFKD NAPPEKEVIESLLSLLFVQKGRLRDAAQRHCEHTKSLSPFNDVVDKTPVIRSVILKLLLK YSFHDVKDYIQEYLTLLKKKAFITEDKTELYMLFINCLEDSILEKTSAYSRNDELNHLEE EGRFLKAYSPASRGREPANEASVEYLQEVARIRLCLDRAADFLSEPEGGPEMAKEKQCYL QQVKQFCIRVENDWHRVYLVRKLSSQRGMEFVQGLSKPGRPHQWVFPKDVVKQQGLRQDH PGQMDRYLVYGDEYKALRDAVAKAVLECKPLGIKTALKACKTPQSQQSAYFLLTLFREVA ILYRSHNASLHPTPEQCEAVSKFIGECKILSPPDISRFATSLVDNSVPLLRAGPSDSNLD GTVTEMAIHAAAVLLCGQNELLEPLKNLAFSPATMAHAFLPTMPEDLLAQARRWKGLERV HWYTCPNGHPCSVGECGRPMEQSICIDCHAPIGGIDHKPRDGFHLVKDKADRTQTGHVLG NPQRRDVVTCDRGLPPVVFLLIRLLTHLALLLGASQSSQALINIIKPPVRDPKGFLQQHI LKDLEQLAKMLGHSADETIGVVHLVLRRLLQEQHQLSSRRLLNFDTELSTKEMRNNWEKE IAAVISPELEHLDKTLPTMNNLISQDKRISSNPVAKIIYGDPVTFLPHLPRKSVVHCSKI WSCRKRITVEYLQHIVEQKNGKERVPILWHFLQKEAELRLVKFLPEILALQRDLVKQFQN VQQVEYSSIRGFLSKHSSDGLRQLLHNRITVFLSTWNKLRRSLETNGEINLPKDYCSTDL DLDTEFEILLPRRRGLGLCATALVSYLIRLHNEIVYAVEKLSKENNSYSVDAAEVTELHV ISYEVERDLTPLILSNCQYQVEEGRETVQEFDLEKIQRQIVSRFLQGKPRLSLKGIPTLV YRHDWNYEHLFMDIKNKMAQDSLPSSVISAISGQLQSYSDACEVLSVVEVTLGFLSTAGG DPNMQLNVYTQDILQMGDQTIHVLKALNRCQLKHTIALWQFLSAHKSEQLLRLHKEPFGE ISSRYKADLSPENAKLLSTFLNQTGLDAFLLELHEMIILKLKNPQTQTEERFRPQWSLRD TLVSYMQTKESEILPEMASQFPEEILLASCVSVWKTAAVLKWNREMR |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
AAA ATPase family
|
|||||
| Subcellular location |
Cytoplasm, cytosol
|
|||||
| Function |
Atypical E3 ubiquitin ligase that can catalyze ubiquitination of both proteins and lipids, and which is involved in various processes, such as lipid metabolism, angiogenesis and cell-autonomous immunity. Acts as a key immune sensor by catalyzing ubiquitination of the lipid A moiety of bacterial lipopolysaccharide (LPS) via its RZ-type zinc-finger: restricts the proliferation of cytosolic bacteria, such as Salmonella, by generating the bacterial ubiquitin coat through the ubiquitination of LPS. Also acts indirectly by mediating the recruitment of the LUBAC complex, which conjugates linear polyubiquitin chains. Ubiquitination of LPS triggers cell-autonomous immunity, such as antibacterial autophagy, leading to degradation of the microbial invader. Involved in lipid metabolism by regulating fat storage and lipid droplet formation; act by inhibiting the lipolytic process. Also regulates lipotoxicity by inhibiting desaturation of fatty acids. Also acts as an E3 ubiquitin-protein ligase via its RING-type zinc finger: mediates 'Lys-63'-linked ubiquitination of target proteins. Involved in the non-canonical Wnt signaling pathway in vascular development: acts by mediating ubiquitination and degradation of FLNA and NFATC2 downstream of RSPO3, leading to inhibit the non-canonical Wnt signaling pathway and promoting vessel regression. Also has ATPase activity; ATPase activity is required for ubiquitination of LPS.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
13.90 | LDD0402 | [2] | |
|
C-Sul Probe Info |
 |
14.42 | LDD0066 | [3] | |
|
STPyne Probe Info |
 |
K1151(10.00); K1404(2.12); K1508(8.33); K2367(0.74) | LDD0277 | [4] | |
|
Probe 1 Probe Info |
 |
Y475(40.47); Y4225(16.23); Y4327(17.23); Y4806(14.48) | LDD3495 | [5] | |
|
BTD Probe Info |
 |
C2132(2.64) | LDD1700 | [6] | |
|
AHL-Pu-1 Probe Info |
 |
C614(2.05) | LDD0168 | [7] | |
|
DBIA Probe Info |
 |
CAmbiguous(26.23); C3084(14.51); C2536(5.58); C3330(4.31) | LDD0209 | [8] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [9] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C1748(0.00); C4348(0.00); C2943(0.00); C1881(0.00) | LDD0038 | [10] | |
|
IA-alkyne Probe Info |
 |
C1748(0.00); C4348(0.00); C3008(0.00); C1156(0.00) | LDD0036 | [10] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [11] | |
|
Lodoacetamide azide Probe Info |
 |
C4348(0.00); C2943(0.00); C1881(0.00); C3008(0.00) | LDD0037 | [10] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [12] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [13] | |
|
TFBX Probe Info |
 |
C614(0.00); C3261(0.00) | LDD0027 | [12] | |
|
WYneN Probe Info |
 |
C1748(0.00); C3084(0.00); C1156(0.00); C3609(0.00) | LDD0021 | [14] | |
|
WYneO Probe Info |
 |
C4111(0.00); C3330(0.00); C2943(0.00); C614(0.00) | LDD0022 | [14] | |
|
Compound 10 Probe Info |
 |
C1510(0.00); C1748(0.00); C3008(0.00); C614(0.00) | LDD2216 | [15] | |
|
Compound 11 Probe Info |
 |
C1510(0.00); C1748(0.00); C3008(0.00); C614(0.00) | LDD2213 | [15] | |
|
ENE Probe Info |
 |
C3554(0.00); C614(0.00) | LDD0006 | [14] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [14] | |
|
PF-06672131 Probe Info |
 |
C614(0.00); C4348(0.00) | LDD0017 | [16] | |
|
Acrolein Probe Info |
 |
C3609(0.00); C3330(0.00); C856(0.00); C3261(0.00) | LDD0217 | [17] | |
|
Crotonaldehyde Probe Info |
 |
C1748(0.00); C1614(0.00); C3261(0.00) | LDD0219 | [17] | |
|
Methacrolein Probe Info |
 |
C310(0.00); C3261(0.00); C856(0.00); C1748(0.00) | LDD0218 | [17] | |
|
W1 Probe Info |
 |
C614(0.00); C680(0.00); C3554(0.00); C310(0.00) | LDD0236 | [18] | |
|
AOyne Probe Info |
 |
12.30 | LDD0443 | [19] | |
|
NAIA_5 Probe Info |
 |
C4348(0.00); C5190(0.00); C310(0.00); C1643(0.00) | LDD2223 | [13] | |
|
HHS-482 Probe Info |
 |
Y2792(1.47); Y3273(0.96); Y4806(1.05) | LDD2239 | [20] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C174 Probe Info |
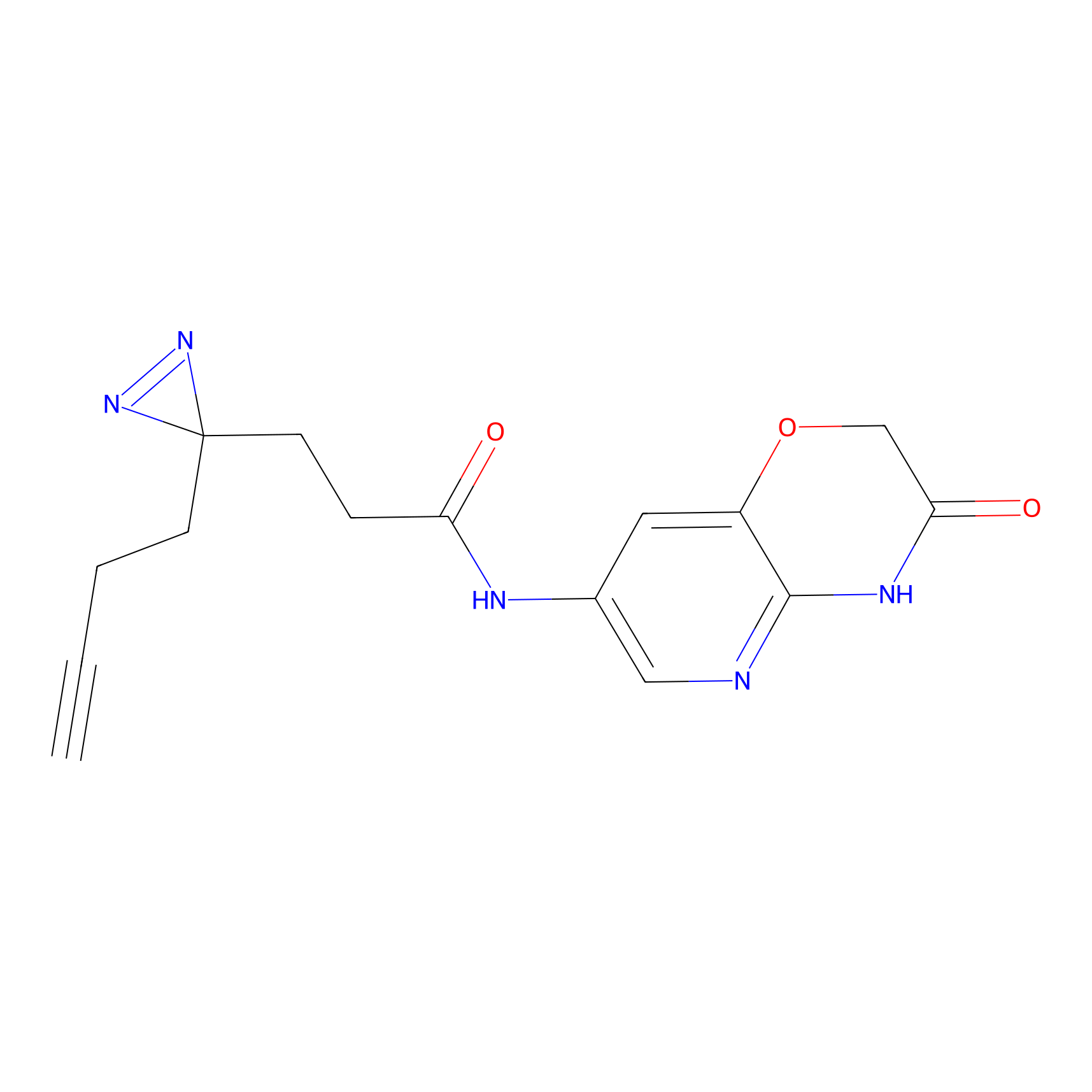 |
4.99 | LDD1854 | [21] | |
|
Alk-rapa Probe Info |
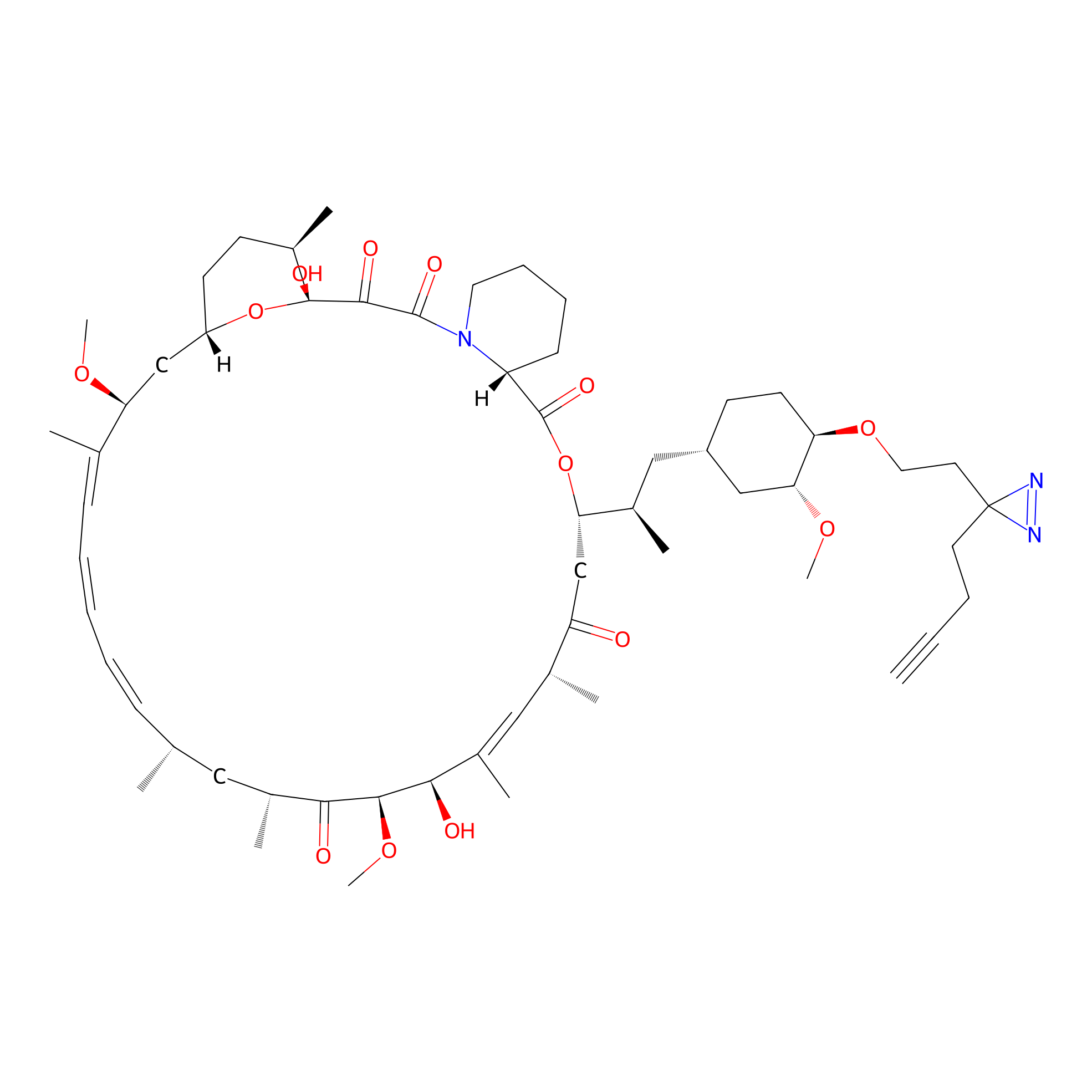 |
9.93 | LDD0213 | [22] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C1510(0.91) | LDD2112 | [6] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C1748(0.99) | LDD2130 | [6] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C856(0.89); C1748(0.97); C1510(0.82); C2132(0.83) | LDD2117 | [6] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C856(1.03); C1510(1.20); C2132(1.05) | LDD2152 | [6] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C1510(0.86) | LDD2103 | [6] |
| LDCM0025 | 4SU-RNA | HEK-293T | C614(2.05) | LDD0168 | [7] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C614(4.49); C310(2.27) | LDD0169 | [7] |
| LDCM0214 | AC1 | HEK-293T | C2536(1.04); C3554(0.91); C310(1.05); C1748(1.01) | LDD1507 | [23] |
| LDCM0215 | AC10 | HEK-293T | C2536(1.01); C3554(0.97); C310(1.02); C1748(1.15) | LDD1508 | [23] |
| LDCM0226 | AC11 | HEK-293T | C2536(1.03); C3554(1.03); C310(1.00); C1748(1.08) | LDD1509 | [23] |
| LDCM0237 | AC12 | HEK-293T | C2536(1.06); C3554(0.93); C310(1.01); C1748(1.05) | LDD1510 | [23] |
| LDCM0259 | AC14 | HEK-293T | C2536(0.94); C3554(1.26); C310(0.90); C1748(0.94) | LDD1512 | [23] |
| LDCM0270 | AC15 | HEK-293T | C2536(0.89); C3554(0.89); C310(1.07); C1748(1.03) | LDD1513 | [23] |
| LDCM0276 | AC17 | HEK-293T | C2536(1.04); C3554(0.85); C310(1.10); C1748(1.25) | LDD1515 | [23] |
| LDCM0277 | AC18 | HEK-293T | C2536(0.88); C3554(1.04); C310(1.07); C1748(0.94) | LDD1516 | [23] |
| LDCM0278 | AC19 | HEK-293T | C2536(1.13); C3554(1.01); C310(1.02); C1748(1.03) | LDD1517 | [23] |
| LDCM0279 | AC2 | HEK-293T | C2536(0.99); C3554(0.97); C310(1.06); C1748(0.97) | LDD1518 | [23] |
| LDCM0280 | AC20 | HEK-293T | C2536(1.01); C3554(1.00); C310(1.03); C1748(0.96) | LDD1519 | [23] |
| LDCM0281 | AC21 | HEK-293T | C2536(1.00); C3554(0.89); C310(0.98); C1748(1.03) | LDD1520 | [23] |
| LDCM0282 | AC22 | HEK-293T | C2536(0.93); C3554(1.18); C310(0.94); C1748(0.92) | LDD1521 | [23] |
| LDCM0283 | AC23 | HEK-293T | C2536(0.97); C3554(1.05); C310(1.04); C1748(1.02) | LDD1522 | [23] |
| LDCM0284 | AC24 | HEK-293T | C2536(0.86); C3554(0.93); C310(0.95); C1748(1.02) | LDD1523 | [23] |
| LDCM0285 | AC25 | HEK-293T | C2536(1.02); C3554(0.89); C310(1.05); C1748(1.09) | LDD1524 | [23] |
| LDCM0286 | AC26 | HEK-293T | C2536(0.95); C3554(1.09); C310(1.01); C1748(1.03) | LDD1525 | [23] |
| LDCM0287 | AC27 | HEK-293T | C2536(1.01); C3554(0.90); C310(1.00); C1748(0.94) | LDD1526 | [23] |
| LDCM0288 | AC28 | HEK-293T | C2536(0.98); C3554(1.08); C310(0.99); C1748(0.94) | LDD1527 | [23] |
| LDCM0289 | AC29 | HEK-293T | C2536(1.12); C3554(1.07); C310(1.00); C1748(1.02) | LDD1528 | [23] |
| LDCM0290 | AC3 | HEK-293T | C2536(0.88); C3554(1.20); C310(0.98); C1748(1.02) | LDD1529 | [23] |
| LDCM0291 | AC30 | HEK-293T | C2536(0.96); C3554(1.12); C310(0.94); C1748(0.93) | LDD1530 | [23] |
| LDCM0292 | AC31 | HEK-293T | C2536(1.02); C3554(1.09); C310(1.00); C1748(1.10) | LDD1531 | [23] |
| LDCM0293 | AC32 | HEK-293T | C2536(0.88); C3554(0.88); C310(1.02); C1748(1.03) | LDD1532 | [23] |
| LDCM0294 | AC33 | HEK-293T | C2536(1.03); C3554(0.85); C310(1.14); C1748(0.94) | LDD1533 | [23] |
| LDCM0295 | AC34 | HEK-293T | C2536(0.97); C3554(0.97); C310(1.05); C1748(0.94) | LDD1534 | [23] |
| LDCM0296 | AC35 | HEK-293T | C2536(1.01); C3554(0.99); C310(0.99); C1748(1.12) | LDD1535 | [23] |
| LDCM0297 | AC36 | HEK-293T | C2536(1.04); C3554(0.95); C310(0.95); C1748(1.01) | LDD1536 | [23] |
| LDCM0298 | AC37 | HEK-293T | C2536(0.90); C3554(1.08); C310(0.92); C1748(1.12) | LDD1537 | [23] |
| LDCM0299 | AC38 | HEK-293T | C2536(0.90); C3554(1.01); C310(0.91); C1748(0.93) | LDD1538 | [23] |
| LDCM0300 | AC39 | HEK-293T | C2536(1.02); C3554(0.99); C310(0.97); C1748(1.03) | LDD1539 | [23] |
| LDCM0301 | AC4 | HEK-293T | C2536(1.07); C3554(0.96); C310(1.01); C1748(0.98) | LDD1540 | [23] |
| LDCM0302 | AC40 | HEK-293T | C2536(0.91); C3554(0.93); C310(0.97); C1748(0.93) | LDD1541 | [23] |
| LDCM0303 | AC41 | HEK-293T | C2536(1.00); C3554(0.89); C310(1.14); C1748(1.09) | LDD1542 | [23] |
| LDCM0304 | AC42 | HEK-293T | C2536(0.96); C3554(1.19); C310(1.06); C1748(1.04) | LDD1543 | [23] |
| LDCM0305 | AC43 | HEK-293T | C2536(1.11); C3554(0.98); C310(1.02); C1748(1.07) | LDD1544 | [23] |
| LDCM0306 | AC44 | HEK-293T | C2536(0.97); C3554(0.95); C310(0.97); C1748(1.03) | LDD1545 | [23] |
| LDCM0307 | AC45 | HEK-293T | C2536(1.08); C3554(1.04); C310(0.99); C1748(1.13) | LDD1546 | [23] |
| LDCM0308 | AC46 | HEK-293T | C2536(1.10); C3554(1.15); C310(0.89); C1748(1.01) | LDD1547 | [23] |
| LDCM0309 | AC47 | HEK-293T | C2536(1.12); C3554(1.10); C310(1.05); C1748(1.00) | LDD1548 | [23] |
| LDCM0310 | AC48 | HEK-293T | C2536(0.95); C3554(1.02); C310(0.95); C1748(0.93) | LDD1549 | [23] |
| LDCM0311 | AC49 | HEK-293T | C2536(1.13); C3554(0.85); C310(1.21); C1748(0.95) | LDD1550 | [23] |
| LDCM0312 | AC5 | HEK-293T | C2536(1.01); C3554(0.98); C310(0.94); C1748(1.05) | LDD1551 | [23] |
| LDCM0313 | AC50 | HEK-293T | C2536(1.05); C3554(1.02); C310(1.00); C1748(1.04) | LDD1552 | [23] |
| LDCM0314 | AC51 | HEK-293T | C2536(0.98); C3554(1.08); C310(1.02); C1748(1.06) | LDD1553 | [23] |
| LDCM0315 | AC52 | HEK-293T | C2536(1.04); C3554(1.00); C310(0.96); C1748(1.00) | LDD1554 | [23] |
| LDCM0316 | AC53 | HEK-293T | C2536(1.29); C3554(1.02); C310(1.00); C1748(1.10) | LDD1555 | [23] |
| LDCM0317 | AC54 | HEK-293T | C2536(0.98); C3554(0.94); C310(0.97); C1748(1.00) | LDD1556 | [23] |
| LDCM0318 | AC55 | HEK-293T | C2536(0.90); C3554(1.02); C310(0.98); C1748(0.96) | LDD1557 | [23] |
| LDCM0319 | AC56 | HEK-293T | C2536(0.85); C3554(1.02); C310(0.96); C1748(0.94) | LDD1558 | [23] |
| LDCM0320 | AC57 | HEK-293T | C2536(1.00); C3554(1.02); C310(1.10); C1748(1.08) | LDD1559 | [23] |
| LDCM0321 | AC58 | HEK-293T | C2536(0.96); C3554(0.98); C310(0.99); C1748(1.02) | LDD1560 | [23] |
| LDCM0322 | AC59 | HEK-293T | C2536(0.96); C3554(1.13); C310(1.00); C1748(1.08) | LDD1561 | [23] |
| LDCM0323 | AC6 | HEK-293T | C2536(0.97); C3554(1.11); C310(0.97); C1748(0.91) | LDD1562 | [23] |
| LDCM0324 | AC60 | HEK-293T | C2536(1.05); C3554(1.07); C310(0.95); C1748(0.97) | LDD1563 | [23] |
| LDCM0325 | AC61 | HEK-293T | C2536(1.11); C3554(1.06); C310(0.97); C1748(1.09) | LDD1564 | [23] |
| LDCM0326 | AC62 | HEK-293T | C2536(1.05); C3554(1.25); C310(0.92); C1748(1.03) | LDD1565 | [23] |
| LDCM0327 | AC63 | HEK-293T | C2536(1.02); C3554(0.94); C310(1.02); C1748(1.03) | LDD1566 | [23] |
| LDCM0328 | AC64 | HEK-293T | C2536(0.89); C3554(0.95); C310(1.00); C1748(1.04) | LDD1567 | [23] |
| LDCM0334 | AC7 | HEK-293T | C2536(0.94); C3554(1.00); C310(0.96); C1748(0.94) | LDD1568 | [23] |
| LDCM0345 | AC8 | HEK-293T | C2536(0.87); C3554(1.06); C310(0.97); C1748(1.02) | LDD1569 | [23] |
| LDCM0248 | AKOS034007472 | HEK-293T | C2536(1.05); C3554(0.90); C310(0.94); C1748(1.09) | LDD1511 | [23] |
| LDCM0356 | AKOS034007680 | HEK-293T | C2536(1.07); C3554(0.89); C310(1.21); C1748(1.09) | LDD1570 | [23] |
| LDCM0275 | AKOS034007705 | HEK-293T | C2536(0.91); C3554(1.04); C310(0.96); C1748(0.94) | LDD1514 | [23] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C856(0.89) | LDD2091 | [6] |
| LDCM0108 | Chloroacetamide | HeLa | C3330(0.00); C1748(0.00); C1614(0.00); C2633(0.00) | LDD0222 | [17] |
| LDCM0632 | CL-Sc | Hep-G2 | C1748(20.00); C2620(20.00); C310(0.94) | LDD2227 | [13] |
| LDCM0367 | CL1 | HEK-293T | C2536(1.00); C3554(1.02); C310(0.97); C1748(1.03) | LDD1571 | [23] |
| LDCM0368 | CL10 | HEK-293T | C2536(1.20); C3554(1.49); C310(0.87); C1748(1.01) | LDD1572 | [23] |
| LDCM0369 | CL100 | HEK-293T | C2536(0.82); C3554(0.72); C1748(1.06); C2132(1.02) | LDD1573 | [23] |
| LDCM0370 | CL101 | HEK-293T | C2536(0.97); C3554(1.03); C310(1.10); C1748(1.03) | LDD1574 | [23] |
| LDCM0371 | CL102 | HEK-293T | C2536(1.12); C3554(0.96); C310(0.97); C1748(1.08) | LDD1575 | [23] |
| LDCM0372 | CL103 | HEK-293T | C2536(0.96); C3554(1.02); C310(1.04); C1748(1.10) | LDD1576 | [23] |
| LDCM0373 | CL104 | HEK-293T | C2536(0.94); C3554(0.88); C1748(1.08); C2132(1.00) | LDD1577 | [23] |
| LDCM0374 | CL105 | HEK-293T | C2536(0.93); C3554(1.02); C310(1.07); C1748(0.99) | LDD1578 | [23] |
| LDCM0375 | CL106 | HEK-293T | C2536(0.92); C3554(1.22); C310(1.06); C1748(1.04) | LDD1579 | [23] |
| LDCM0376 | CL107 | HEK-293T | C2536(0.83); C3554(1.00); C310(1.01); C1748(0.96) | LDD1580 | [23] |
| LDCM0377 | CL108 | HEK-293T | C2536(0.94); C3554(0.88); C1748(1.00); C2132(0.85) | LDD1581 | [23] |
| LDCM0378 | CL109 | HEK-293T | C2536(1.01); C3554(0.87); C310(1.03); C1748(1.03) | LDD1582 | [23] |
| LDCM0379 | CL11 | HEK-293T | C2536(1.10); C3554(1.25); C310(0.94); C1748(1.05) | LDD1583 | [23] |
| LDCM0380 | CL110 | HEK-293T | C2536(1.19); C3554(1.10); C310(0.99); C1748(1.06) | LDD1584 | [23] |
| LDCM0381 | CL111 | HEK-293T | C2536(0.99); C3554(1.26); C310(1.00); C1748(1.05) | LDD1585 | [23] |
| LDCM0382 | CL112 | HEK-293T | C2536(0.94); C3554(0.92); C1748(1.08); C2132(0.97) | LDD1586 | [23] |
| LDCM0383 | CL113 | HEK-293T | C2536(1.09); C3554(0.95); C310(1.05); C1748(1.03) | LDD1587 | [23] |
| LDCM0384 | CL114 | HEK-293T | C2536(1.09); C3554(1.08); C310(0.90); C1748(1.11) | LDD1588 | [23] |
| LDCM0385 | CL115 | HEK-293T | C2536(0.97); C3554(1.02); C310(1.05); C1748(1.08) | LDD1589 | [23] |
| LDCM0386 | CL116 | HEK-293T | C2536(0.80); C3554(0.77); C1748(1.07); C2132(1.00) | LDD1590 | [23] |
| LDCM0387 | CL117 | HEK-293T | C2536(0.99); C3554(0.89); C310(1.05); C1748(1.04) | LDD1591 | [23] |
| LDCM0388 | CL118 | HEK-293T | C2536(1.10); C3554(0.86); C310(0.99); C1748(1.00) | LDD1592 | [23] |
| LDCM0389 | CL119 | HEK-293T | C2536(0.91); C3554(1.03); C310(1.06); C1748(1.05) | LDD1593 | [23] |
| LDCM0390 | CL12 | HEK-293T | C2536(0.94); C3554(1.14); C310(0.91); C1748(0.98) | LDD1594 | [23] |
| LDCM0391 | CL120 | HEK-293T | C2536(1.08); C3554(0.76); C1748(1.06); C2132(1.10) | LDD1595 | [23] |
| LDCM0392 | CL121 | HEK-293T | C2536(1.01); C3554(1.12); C310(0.97); C1748(1.00) | LDD1596 | [23] |
| LDCM0393 | CL122 | HEK-293T | C2536(1.24); C3554(0.96); C310(1.00); C1748(1.08) | LDD1597 | [23] |
| LDCM0394 | CL123 | HEK-293T | C2536(0.93); C3554(1.63); C310(0.97); C1748(1.05) | LDD1598 | [23] |
| LDCM0395 | CL124 | HEK-293T | C2536(1.61); C3554(1.31); C1748(0.98); C2132(0.95) | LDD1599 | [23] |
| LDCM0396 | CL125 | HEK-293T | C2536(0.95); C3554(0.93); C310(1.00); C1748(0.97) | LDD1600 | [23] |
| LDCM0397 | CL126 | HEK-293T | C2536(0.93); C3554(1.04); C310(0.95); C1748(1.02) | LDD1601 | [23] |
| LDCM0398 | CL127 | HEK-293T | C2536(0.97); C3554(1.07); C310(1.17); C1748(1.06) | LDD1602 | [23] |
| LDCM0399 | CL128 | HEK-293T | C2536(0.81); C3554(0.93); C1748(0.94); C2132(0.97) | LDD1603 | [23] |
| LDCM0400 | CL13 | HEK-293T | C2536(1.00); C3554(0.97); C310(1.05); C1748(0.94) | LDD1604 | [23] |
| LDCM0401 | CL14 | HEK-293T | C2536(0.87); C3554(0.93); C310(0.92); C1748(0.93) | LDD1605 | [23] |
| LDCM0402 | CL15 | HEK-293T | C2536(0.97); C3554(1.44); C310(0.94); C1748(1.11) | LDD1606 | [23] |
| LDCM0403 | CL16 | HEK-293T | C2536(0.76); C3554(0.83); C1748(0.99); C2132(1.07) | LDD1607 | [23] |
| LDCM0404 | CL17 | HEK-293T | C2536(0.99); C3554(1.26); C310(1.02); C1748(0.94) | LDD1608 | [23] |
| LDCM0405 | CL18 | HEK-293T | C2536(0.92); C3554(0.95); C310(1.02); C1748(1.04) | LDD1609 | [23] |
| LDCM0406 | CL19 | HEK-293T | C2536(1.05); C3554(1.07); C310(1.02); C1748(1.18) | LDD1610 | [23] |
| LDCM0407 | CL2 | HEK-293T | C2536(0.95); C3554(1.06); C310(0.97); C1748(0.98) | LDD1611 | [23] |
| LDCM0408 | CL20 | HEK-293T | C2536(1.17); C3554(1.09); C310(0.98); C1748(1.04) | LDD1612 | [23] |
| LDCM0409 | CL21 | HEK-293T | C2536(1.02); C3554(1.50); C310(0.89); C1748(1.24) | LDD1613 | [23] |
| LDCM0410 | CL22 | HEK-293T | C2536(1.08); C3554(1.39); C310(1.14); C1748(0.87) | LDD1614 | [23] |
| LDCM0411 | CL23 | HEK-293T | C2536(1.11); C3554(1.16); C310(1.02); C1748(1.09) | LDD1615 | [23] |
| LDCM0412 | CL24 | HEK-293T | C2536(0.92); C3554(1.08); C310(0.96); C1748(1.03) | LDD1616 | [23] |
| LDCM0413 | CL25 | HEK-293T | C2536(0.84); C3554(1.30); C310(0.95); C1748(1.09) | LDD1617 | [23] |
| LDCM0414 | CL26 | HEK-293T | C2536(0.88); C3554(0.99); C310(0.99); C1748(1.01) | LDD1618 | [23] |
| LDCM0415 | CL27 | HEK-293T | C2536(0.94); C3554(1.12); C310(0.93); C1748(1.07) | LDD1619 | [23] |
| LDCM0416 | CL28 | HEK-293T | C2536(0.77); C3554(0.74); C1748(1.02); C2132(0.98) | LDD1620 | [23] |
| LDCM0417 | CL29 | HEK-293T | C2536(1.13); C3554(0.80); C310(1.01); C1748(1.01) | LDD1621 | [23] |
| LDCM0418 | CL3 | HEK-293T | C2536(1.00); C3554(1.07); C310(1.09); C1748(1.03) | LDD1622 | [23] |
| LDCM0419 | CL30 | HEK-293T | C2536(0.92); C3554(1.01); C310(1.00); C1748(1.08) | LDD1623 | [23] |
| LDCM0420 | CL31 | HEK-293T | C2536(0.96); C3554(1.08); C310(1.05); C1748(1.08) | LDD1624 | [23] |
| LDCM0421 | CL32 | HEK-293T | C2536(1.06); C3554(1.22); C310(1.02); C1748(1.04) | LDD1625 | [23] |
| LDCM0422 | CL33 | HEK-293T | C2536(1.36); C3554(1.15); C310(0.80); C1748(1.19) | LDD1626 | [23] |
| LDCM0423 | CL34 | HEK-293T | C2536(1.02); C3554(1.22); C310(0.97); C1748(0.98) | LDD1627 | [23] |
| LDCM0424 | CL35 | HEK-293T | C2536(1.18); C3554(0.95); C310(1.10); C1748(1.11) | LDD1628 | [23] |
| LDCM0425 | CL36 | HEK-293T | C2536(0.97); C3554(1.18); C310(0.95); C1748(1.18) | LDD1629 | [23] |
| LDCM0426 | CL37 | HEK-293T | C2536(0.94); C3554(0.95); C310(0.96); C1748(0.94) | LDD1630 | [23] |
| LDCM0428 | CL39 | HEK-293T | C2536(0.96); C3554(0.95); C310(0.97); C1748(1.05) | LDD1632 | [23] |
| LDCM0429 | CL4 | HEK-293T | C2536(0.90); C3554(1.29); C1748(1.02); C2132(1.05) | LDD1633 | [23] |
| LDCM0430 | CL40 | HEK-293T | C2536(0.89); C3554(0.76); C1748(1.08); C2132(0.95) | LDD1634 | [23] |
| LDCM0431 | CL41 | HEK-293T | C2536(1.03); C3554(1.03); C310(1.14); C1748(0.93) | LDD1635 | [23] |
| LDCM0432 | CL42 | HEK-293T | C2536(0.93); C3554(1.16); C310(1.00); C1748(0.96) | LDD1636 | [23] |
| LDCM0433 | CL43 | HEK-293T | C2536(0.98); C3554(1.09); C310(1.01); C1748(1.10) | LDD1637 | [23] |
| LDCM0434 | CL44 | HEK-293T | C2536(1.02); C3554(1.09); C310(1.02); C1748(1.03) | LDD1638 | [23] |
| LDCM0435 | CL45 | HEK-293T | C2536(1.17); C3554(0.88); C310(1.04); C1748(1.11) | LDD1639 | [23] |
| LDCM0436 | CL46 | HEK-293T | C2536(1.11); C3554(1.16); C310(0.92); C1748(1.17) | LDD1640 | [23] |
| LDCM0437 | CL47 | HEK-293T | C2536(1.11); C3554(0.99); C310(1.18); C1748(1.10) | LDD1641 | [23] |
| LDCM0438 | CL48 | HEK-293T | C2536(1.02); C3554(1.17); C310(0.95); C1748(1.29) | LDD1642 | [23] |
| LDCM0439 | CL49 | HEK-293T | C2536(0.91); C3554(1.03); C310(0.96); C1748(1.13) | LDD1643 | [23] |
| LDCM0440 | CL5 | HEK-293T | C2536(1.07); C3554(0.93); C310(1.23); C1748(0.86) | LDD1644 | [23] |
| LDCM0441 | CL50 | HEK-293T | C2536(1.05); C3554(1.21); C310(0.96); C1748(1.01) | LDD1645 | [23] |
| LDCM0443 | CL52 | HEK-293T | C2536(0.77); C3554(0.96); C1748(1.06); C2132(1.06) | LDD1646 | [23] |
| LDCM0444 | CL53 | HEK-293T | C2536(1.01); C3554(0.95); C310(1.09); C1748(1.06) | LDD1647 | [23] |
| LDCM0445 | CL54 | HEK-293T | C2536(1.08); C3554(1.16); C310(0.96); C1748(1.00) | LDD1648 | [23] |
| LDCM0446 | CL55 | HEK-293T | C2536(1.01); C3554(1.02); C310(1.00); C1748(1.12) | LDD1649 | [23] |
| LDCM0447 | CL56 | HEK-293T | C2536(1.08); C3554(1.02); C310(0.99); C1748(1.12) | LDD1650 | [23] |
| LDCM0448 | CL57 | HEK-293T | C2536(1.24); C3554(0.74); C310(0.89); C1748(1.31) | LDD1651 | [23] |
| LDCM0449 | CL58 | HEK-293T | C2536(1.11); C3554(1.36); C310(0.89); C1748(1.03) | LDD1652 | [23] |
| LDCM0450 | CL59 | HEK-293T | C2536(1.04); C3554(1.06); C310(0.98); C1748(1.09) | LDD1653 | [23] |
| LDCM0451 | CL6 | HEK-293T | C2536(1.02); C3554(1.24); C310(1.01); C1748(1.02) | LDD1654 | [23] |
| LDCM0452 | CL60 | HEK-293T | C2536(0.87); C3554(1.27); C310(0.92); C1748(0.96) | LDD1655 | [23] |
| LDCM0453 | CL61 | HEK-293T | C2536(1.01); C3554(1.05); C310(0.99); C1748(1.00) | LDD1656 | [23] |
| LDCM0454 | CL62 | HEK-293T | C2536(1.68); C3554(0.81); C310(0.99); C1748(1.04) | LDD1657 | [23] |
| LDCM0455 | CL63 | HEK-293T | C2536(1.02); C3554(1.01); C310(1.06); C1748(1.10) | LDD1658 | [23] |
| LDCM0456 | CL64 | HEK-293T | C2536(0.92); C3554(1.12); C1748(1.10); C2132(1.13) | LDD1659 | [23] |
| LDCM0457 | CL65 | HEK-293T | C2536(1.05); C3554(0.91); C310(1.20); C1748(1.08) | LDD1660 | [23] |
| LDCM0458 | CL66 | HEK-293T | C2536(1.08); C3554(1.03); C310(0.99); C1748(1.02) | LDD1661 | [23] |
| LDCM0459 | CL67 | HEK-293T | C2536(1.14); C3554(1.08); C310(0.97); C1748(1.13) | LDD1662 | [23] |
| LDCM0460 | CL68 | HEK-293T | C2536(1.10); C3554(1.14); C310(0.97); C1748(1.04) | LDD1663 | [23] |
| LDCM0461 | CL69 | HEK-293T | C2536(1.24); C3554(0.98); C310(0.97); C1748(1.16) | LDD1664 | [23] |
| LDCM0462 | CL7 | HEK-293T | C2536(1.04); C3554(0.97); C310(0.98); C1748(1.15) | LDD1665 | [23] |
| LDCM0463 | CL70 | HEK-293T | C2536(1.02); C3554(1.31); C310(0.91); C1748(0.94) | LDD1666 | [23] |
| LDCM0464 | CL71 | HEK-293T | C2536(1.07); C3554(1.10); C310(0.99); C1748(1.12) | LDD1667 | [23] |
| LDCM0465 | CL72 | HEK-293T | C2536(0.95); C3554(1.05); C310(0.95); C1748(1.17) | LDD1668 | [23] |
| LDCM0466 | CL73 | HEK-293T | C2536(0.89); C3554(1.12); C310(0.98); C1748(1.01) | LDD1669 | [23] |
| LDCM0467 | CL74 | HEK-293T | C2536(0.90); C3554(1.09); C310(0.97); C1748(1.01) | LDD1670 | [23] |
| LDCM0469 | CL76 | HEK-293T | C2536(0.80); C3554(1.07); C1748(0.98); C2132(1.09) | LDD1672 | [23] |
| LDCM0470 | CL77 | HEK-293T | C2536(1.04); C3554(0.95); C310(1.08); C1748(0.93) | LDD1673 | [23] |
| LDCM0471 | CL78 | HEK-293T | C2536(0.87); C3554(1.17); C310(1.02); C1748(1.24) | LDD1674 | [23] |
| LDCM0472 | CL79 | HEK-293T | C2536(1.04); C3554(0.92); C310(0.98); C1748(1.14) | LDD1675 | [23] |
| LDCM0473 | CL8 | HEK-293T | C2536(1.33); C3554(1.65); C310(0.69); C1748(1.16) | LDD1676 | [23] |
| LDCM0474 | CL80 | HEK-293T | C2536(0.94); C3554(0.97); C310(0.99); C1748(1.01) | LDD1677 | [23] |
| LDCM0475 | CL81 | HEK-293T | C2536(1.32); C3554(1.01); C310(1.02); C1748(1.27) | LDD1678 | [23] |
| LDCM0476 | CL82 | HEK-293T | C2536(1.02); C3554(0.99); C310(0.89); C1748(0.92) | LDD1679 | [23] |
| LDCM0477 | CL83 | HEK-293T | C2536(0.97); C3554(1.29); C310(1.01); C1748(1.08) | LDD1680 | [23] |
| LDCM0478 | CL84 | HEK-293T | C2536(0.93); C3554(1.14); C310(0.91); C1748(1.02) | LDD1681 | [23] |
| LDCM0479 | CL85 | HEK-293T | C2536(0.92); C3554(0.90); C310(1.01); C1748(0.95) | LDD1682 | [23] |
| LDCM0480 | CL86 | HEK-293T | C2536(0.82); C3554(0.93); C310(0.92); C1748(1.00) | LDD1683 | [23] |
| LDCM0481 | CL87 | HEK-293T | C2536(0.95); C3554(1.10); C310(1.04); C1748(0.99) | LDD1684 | [23] |
| LDCM0482 | CL88 | HEK-293T | C2536(0.90); C3554(1.34); C1748(1.03); C2132(0.96) | LDD1685 | [23] |
| LDCM0483 | CL89 | HEK-293T | C2536(1.00); C3554(1.03); C310(1.09); C1748(1.03) | LDD1686 | [23] |
| LDCM0484 | CL9 | HEK-293T | C2536(1.33); C3554(1.01); C310(0.90); C1748(1.12) | LDD1687 | [23] |
| LDCM0485 | CL90 | HEK-293T | C2536(1.16); C3554(1.33); C310(0.81); C1748(1.21) | LDD1688 | [23] |
| LDCM0486 | CL91 | HEK-293T | C2536(1.08); C3554(0.93); C310(0.94); C1748(1.10) | LDD1689 | [23] |
| LDCM0487 | CL92 | HEK-293T | C2536(1.07); C3554(1.48); C310(0.95); C1748(1.07) | LDD1690 | [23] |
| LDCM0488 | CL93 | HEK-293T | C2536(1.07); C3554(1.05); C310(0.89); C1748(1.27) | LDD1691 | [23] |
| LDCM0489 | CL94 | HEK-293T | C2536(1.03); C3554(1.37); C310(0.91); C1748(1.02) | LDD1692 | [23] |
| LDCM0490 | CL95 | HEK-293T | C2536(1.19); C3554(1.48); C310(0.95); C1748(1.53) | LDD1693 | [23] |
| LDCM0491 | CL96 | HEK-293T | C2536(0.93); C3554(1.33); C310(0.93); C1748(1.18) | LDD1694 | [23] |
| LDCM0492 | CL97 | HEK-293T | C2536(1.01); C3554(0.97); C310(1.09); C1748(1.02) | LDD1695 | [23] |
| LDCM0493 | CL98 | HEK-293T | C2536(1.05); C3554(1.12); C310(0.99); C1748(0.97) | LDD1696 | [23] |
| LDCM0494 | CL99 | HEK-293T | C2536(0.91); C3554(1.06); C310(1.05); C1748(1.04) | LDD1697 | [23] |
| LDCM0198 | Dimethyl Fumarate(DMF) | T cell | C4407(4.11) | LDD0513 | [24] |
| LDCM0495 | E2913 | HEK-293T | C2536(0.95); C3554(1.06); C310(1.09); C1748(1.10) | LDD1698 | [23] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C4348(4.45); C1881(2.44) | LDD1702 | [6] |
| LDCM0203 | EV-96 | T cell | C614(5.01) | LDD0523 | [24] |
| LDCM0205 | EV-98 | T cell | C4407(4.78) | LDD0519 | [24] |
| LDCM0625 | F8 | Ramos | C1748(1.23); C3261(0.50) | LDD2187 | [25] |
| LDCM0572 | Fragment10 | Ramos | C1748(0.84); C3261(0.66); C3609(0.96) | LDD2189 | [25] |
| LDCM0573 | Fragment11 | Ramos | C3261(0.78) | LDD2190 | [25] |
| LDCM0574 | Fragment12 | Ramos | C1748(0.85); C3261(0.87) | LDD2191 | [25] |
| LDCM0575 | Fragment13 | Ramos | C1748(0.80); C3261(0.92); C3609(0.72) | LDD2192 | [25] |
| LDCM0576 | Fragment14 | Ramos | C1748(1.51) | LDD2193 | [25] |
| LDCM0579 | Fragment20 | Ramos | C1748(0.78) | LDD2194 | [25] |
| LDCM0580 | Fragment21 | Ramos | C1748(0.88); C3261(0.85); C3609(1.05) | LDD2195 | [25] |
| LDCM0582 | Fragment23 | Ramos | C1748(0.68) | LDD2196 | [25] |
| LDCM0578 | Fragment27 | Ramos | C1748(0.93); C3261(0.87); C3609(0.98) | LDD2197 | [25] |
| LDCM0586 | Fragment28 | Ramos | C1748(0.52); C3261(0.42); C3609(1.06) | LDD2198 | [25] |
| LDCM0588 | Fragment30 | Ramos | C1748(1.01); C3261(0.65); C3609(1.04) | LDD2199 | [25] |
| LDCM0589 | Fragment31 | Ramos | C1748(0.93); C3261(0.88); C3609(0.66) | LDD2200 | [25] |
| LDCM0590 | Fragment32 | Ramos | C1748(0.87); C3609(0.76) | LDD2201 | [25] |
| LDCM0468 | Fragment33 | HEK-293T | C2536(0.93); C3554(1.01); C310(1.07); C1748(1.07) | LDD1671 | [23] |
| LDCM0596 | Fragment38 | Ramos | C1748(0.81); C3261(0.78); C3609(1.18) | LDD2203 | [25] |
| LDCM0566 | Fragment4 | Ramos | C1748(0.97); C3609(0.93) | LDD2184 | [25] |
| LDCM0427 | Fragment51 | HEK-293T | C2536(0.90); C3554(0.86); C310(0.95); C1748(1.02) | LDD1631 | [23] |
| LDCM0610 | Fragment52 | Ramos | C1748(1.52); C3261(0.88) | LDD2204 | [25] |
| LDCM0614 | Fragment56 | Ramos | C1748(1.52) | LDD2205 | [25] |
| LDCM0569 | Fragment7 | Ramos | C1748(0.93); C3261(0.76); C3609(1.01) | LDD2186 | [25] |
| LDCM0571 | Fragment9 | Ramos | C1748(1.10) | LDD2188 | [25] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [17] |
| LDCM0022 | KB02 | HEK-293T | C3554(0.84); C310(1.01); C2536(0.90); C1748(0.90) | LDD1492 | [23] |
| LDCM0023 | KB03 | Jurkat | CAmbiguous(26.23); C3084(14.51); C2536(5.58); C3330(4.31) | LDD0209 | [8] |
| LDCM0024 | KB05 | G361 | C1797(5.94); C2992(3.68); C3603(3.28); C1478(3.62) | LDD3311 | [26] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C1510(1.22) | LDD2102 | [6] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0224 | [17] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C1510(0.83) | LDD2090 | [6] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C1510(0.88) | LDD2092 | [6] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C856(1.16); C1510(1.10) | LDD2093 | [6] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C1510(0.87) | LDD2094 | [6] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C1510(1.20) | LDD2096 | [6] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C1510(1.08) | LDD2098 | [6] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C856(1.00); C1748(0.89); C1510(0.60); C2132(1.01) | LDD2099 | [6] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C1510(0.93) | LDD2104 | [6] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C1510(1.44) | LDD2105 | [6] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C856(0.91); C1748(0.86); C1510(0.92); C2132(1.06) | LDD2107 | [6] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C856(0.81); C1748(0.74); C2132(0.89) | LDD2109 | [6] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C856(0.84); C1748(0.93) | LDD2111 | [6] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C1510(0.63) | LDD2114 | [6] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C856(0.90); C2132(0.84); C614(1.27) | LDD2123 | [6] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C856(0.91); C1748(0.68); C1510(0.90); C2132(0.83) | LDD2125 | [6] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C856(0.86); C1748(0.77); C2132(1.04) | LDD2127 | [6] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C856(1.36) | LDD2129 | [6] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C856(1.22); C1748(1.29); C2132(0.88); C1736(0.94) | LDD2136 | [6] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C856(0.66); C1748(1.02) | LDD2137 | [6] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C2132(2.64) | LDD1700 | [6] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C1510(0.71); C2132(0.75) | LDD2140 | [6] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C1510(0.85); C2132(0.89) | LDD2146 | [6] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C856(1.28); C1510(0.59) | LDD2150 | [6] |
| LDCM0131 | RA190 | MM1.R | C78(1.46) | LDD0304 | [27] |
| LDCM0090 | Rapamycin | JHH-7 | 9.93 | LDD0213 | [22] |
| LDCM0112 | W16 | Hep-G2 | C614(0.64) | LDD0239 | [18] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| E3 ubiquitin-protein ligase TRIM23 (TRIM23) | Arf family | P36406 | |||
| Zinc finger protein RFP (TRIM27) | TRIM/RBCC family | P14373 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Proto-oncogene c-Rel (REL) | . | Q04864 | |||
Other
References
