Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P2 Probe Info |
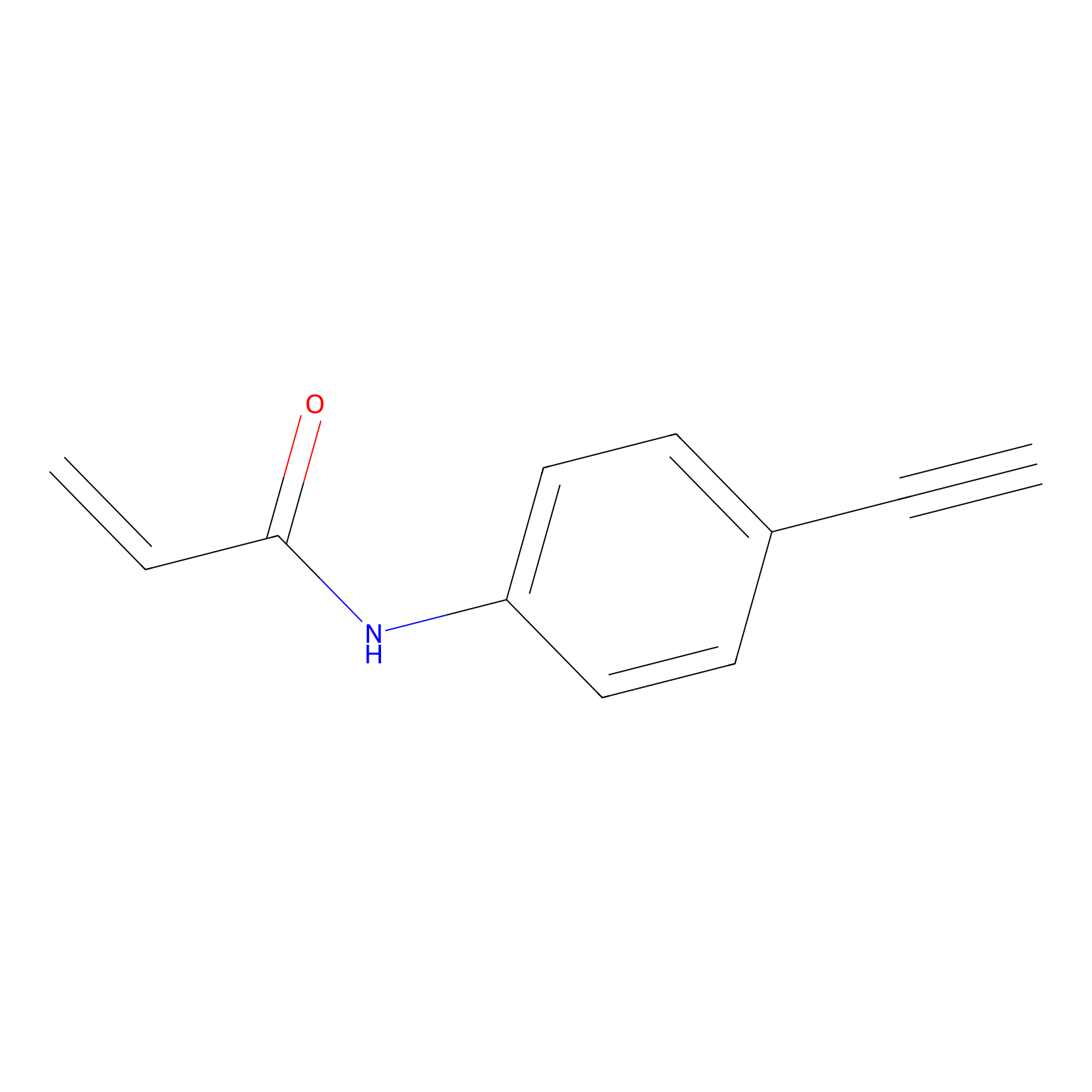 |
10.00 | LDD0449 | [1] | |
|
P8 Probe Info |
 |
10.00 | LDD0451 | [1] | |
|
FBPP2 Probe Info |
 |
5.87 | LDD0318 | [2] | |
|
C-Sul Probe Info |
 |
11.23 | LDD0066 | [3] | |
|
TH211 Probe Info |
 |
Y76(13.51) | LDD0257 | [4] | |
|
TH216 Probe Info |
 |
Y88(10.49) | LDD0259 | [4] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [5] | |
|
AZ-9 Probe Info |
 |
E75(0.99) | LDD2208 | [6] | |
|
ONAyne Probe Info |
 |
K36(0.00); K28(0.00); K56(0.00); K70(0.00) | LDD0273 | [7] | |
|
OPA-S-S-alkyne Probe Info |
 |
K86(1.85); K8(4.22); K99(4.89) | LDD3494 | [8] | |
|
Probe 1 Probe Info |
 |
Y76(8.69); Y88(85.18) | LDD3495 | [9] | |
|
m-APA Probe Info |
 |
6.16 | LDD0403 | [10] | |
|
HHS-482 Probe Info |
 |
Y76(2.92) | LDD0285 | [11] | |
|
HHS-475 Probe Info |
 |
Y76(1.28) | LDD0264 | [12] | |
|
W1 Probe Info |
 |
K8(1.78) | LDD0237 | [13] | |
|
AMP probe Probe Info |
 |
K56(0.00); K40(0.00); K86(0.00) | LDD0200 | [14] | |
|
ATP probe Probe Info |
 |
K56(0.00); K40(0.00); K86(0.00); K28(0.00) | LDD0199 | [14] | |
|
Alkyne tyramide Probe Info |
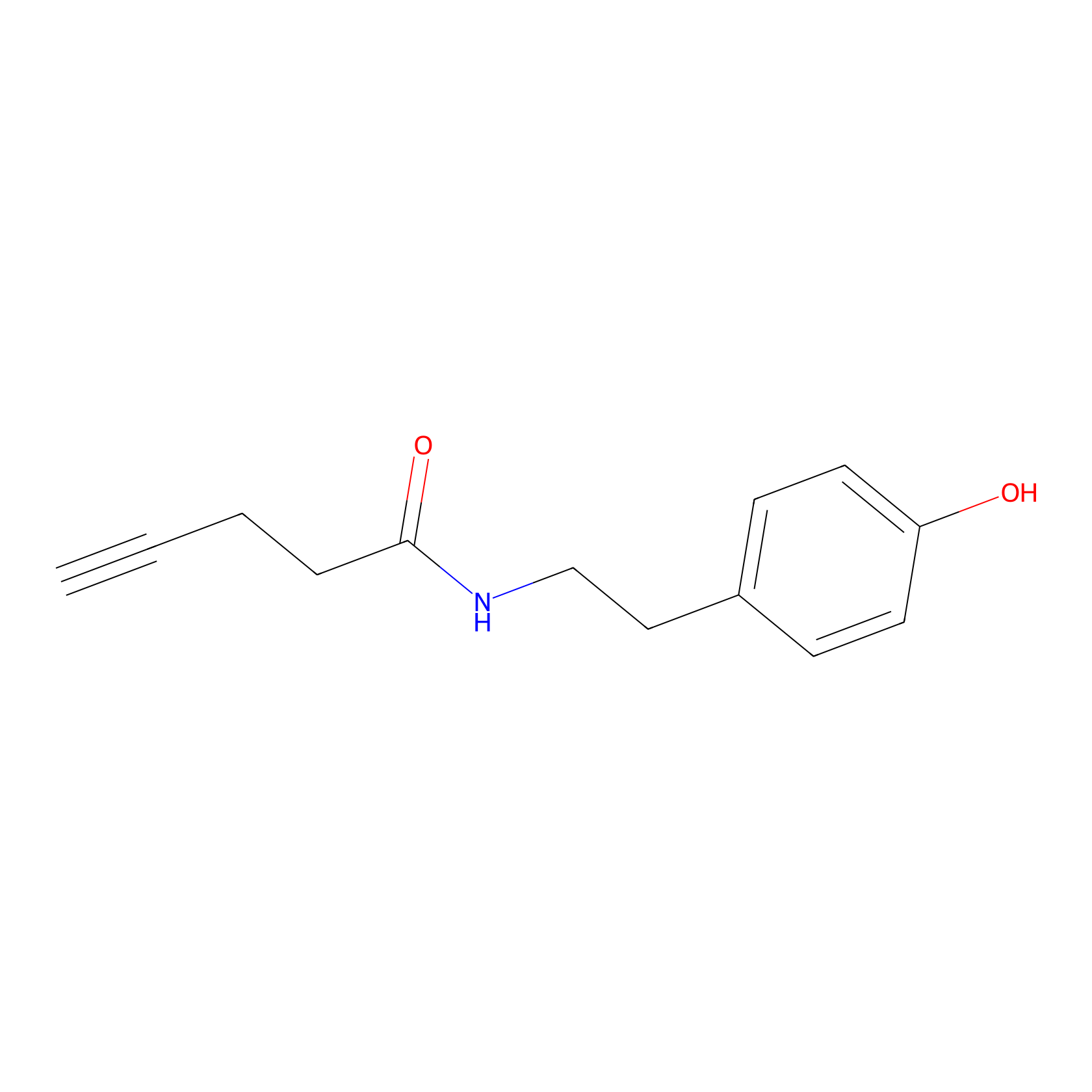 |
Y76(0.00); Y88(0.00) | LDD0003 | [15] | |
|
2PCA Probe Info |
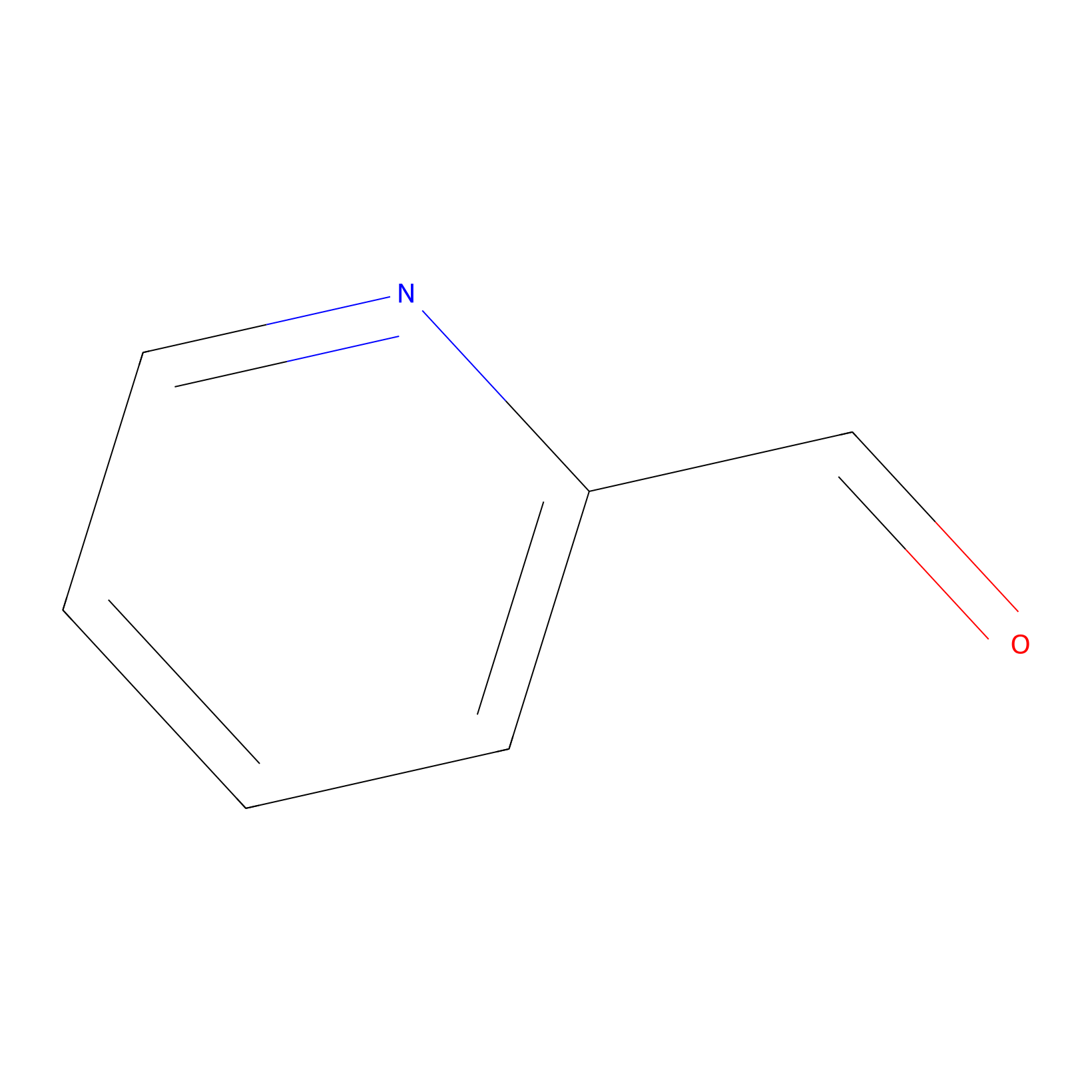 |
N.A. | LDD0034 | [16] | |
|
ATP probe Probe Info |
 |
K36(0.00); K8(0.00); K70(0.00); K66(0.00) | LDD0035 | [17] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0356 | [18] | |
|
NHS Probe Info |
 |
K36(0.00); K56(0.00); K70(0.00); K80(0.00) | LDD0010 | [15] | |
|
OSF Probe Info |
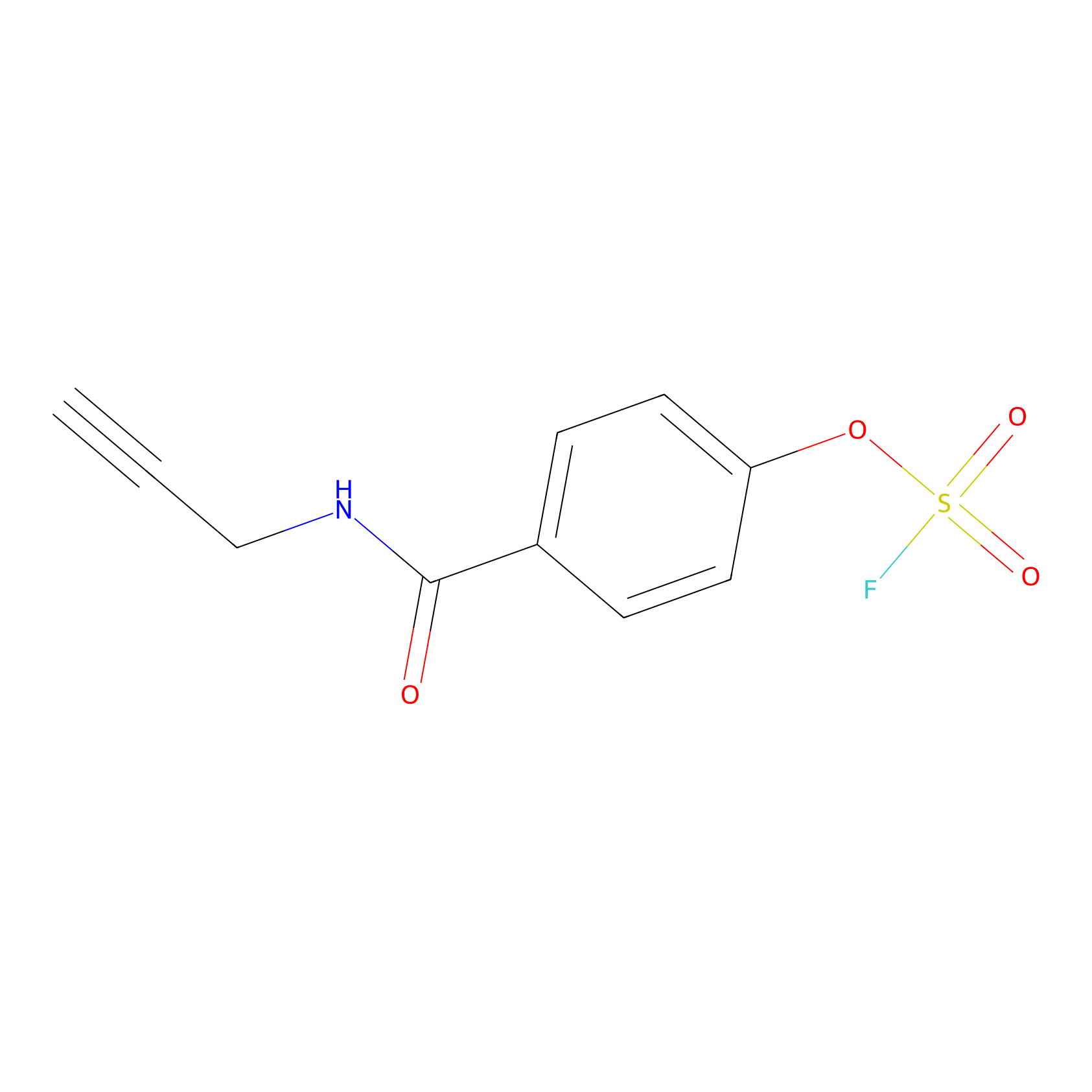 |
N.A. | LDD0029 | [19] | |
|
SF Probe Info |
 |
K56(0.00); K36(0.00); K40(0.00); Y76(0.00) | LDD0028 | [19] | |
|
STPyne Probe Info |
 |
K70(0.00); K54(0.00); K28(0.00); K56(0.00) | LDD0009 | [15] | |
|
1c-yne Probe Info |
 |
K40(0.00); K70(0.00); K66(0.00); K80(0.00) | LDD0228 | [18] | |
|
HHS-465 Probe Info |
 |
K36(0.00); K40(0.00); K54(0.00); K56(0.00) | LDD2240 | [20] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe13 Probe Info |
 |
9.94 | LDD0475 | [21] | |
|
FFF probe2 Probe Info |
 |
7.28 | LDD0463 | [21] | |
|
CR-1 Probe Info |
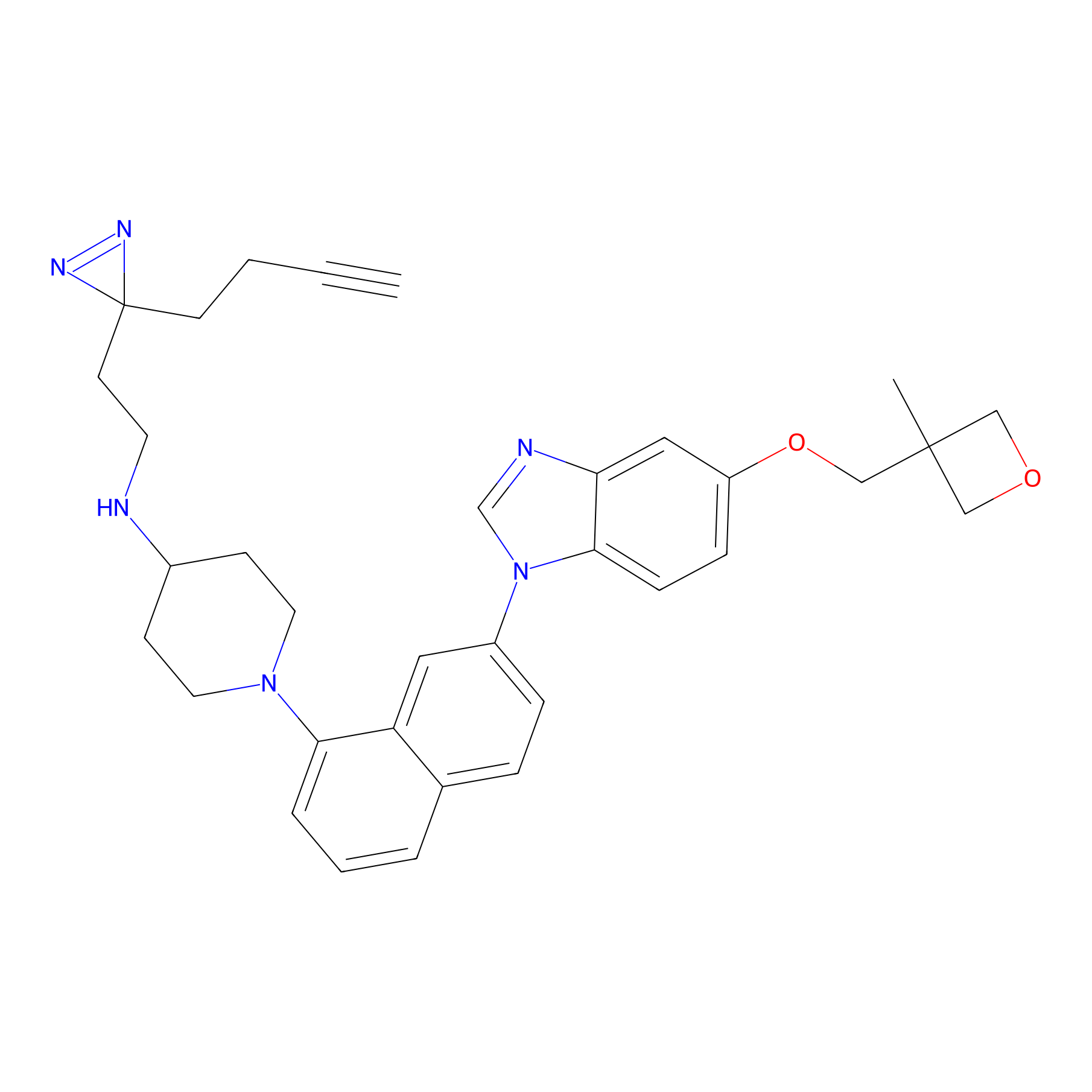 |
2.88 | LDD0430 | [22] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0156 | Aniline | NCI-H1299 | 6.16 | LDD0403 | [10] |
| LDCM0168 | Crenolanib | MV4-11 | 2.88 | LDD0430 | [22] |
| LDCM0116 | HHS-0101 | DM93 | Y76(1.28) | LDD0264 | [12] |
| LDCM0117 | HHS-0201 | DM93 | Y76(0.92) | LDD0265 | [12] |
| LDCM0118 | HHS-0301 | DM93 | Y76(1.13) | LDD0266 | [12] |
| LDCM0119 | HHS-0401 | DM93 | Y76(0.82) | LDD0267 | [12] |
| LDCM0120 | HHS-0701 | DM93 | Y76(1.23) | LDD0268 | [12] |
| LDCM0123 | JWB131 | DM93 | Y76(2.92) | LDD0285 | [11] |
| LDCM0124 | JWB142 | DM93 | Y76(3.27) | LDD0286 | [11] |
| LDCM0125 | JWB146 | DM93 | Y76(2.34) | LDD0287 | [11] |
| LDCM0126 | JWB150 | DM93 | Y76(8.33) | LDD0288 | [11] |
| LDCM0127 | JWB152 | DM93 | Y76(7.13) | LDD0289 | [11] |
| LDCM0128 | JWB198 | DM93 | Y76(2.42) | LDD0290 | [11] |
| LDCM0129 | JWB202 | DM93 | Y76(3.32) | LDD0291 | [11] |
| LDCM0130 | JWB211 | DM93 | Y76(2.52) | LDD0292 | [11] |
| LDCM0110 | W12 | Hep-G2 | K8(1.78) | LDD0237 | [13] |
The Interaction Atlas With This Target
References
