Details of the Target
General Information of Target
| Target ID | LDTP04925 | |||||
|---|---|---|---|---|---|---|
| Target Name | Pterin-4-alpha-carbinolamine dehydratase (PCBD1) | |||||
| Gene Name | PCBD1 | |||||
| Gene ID | 5092 | |||||
| Synonyms |
DCOH; PCBD; Pterin-4-alpha-carbinolamine dehydratase; PHS; EC 4.2.1.96; 4-alpha-hydroxy-tetrahydropterin dehydratase; Dimerization cofactor of hepatocyte nuclear factor 1-alpha; DCoH; Dimerization cofactor of HNF1; Phenylalanine hydroxylase-stimulating protein; Pterin carbinolamine dehydratase; PCD
|
|||||
| 3D Structure | ||||||
| Sequence |
MAGKAHRLSAEERDQLLPNLRAVGWNELEGRDAIFKQFHFKDFNRAFGFMTRVALQAEKL
DHHPEWFNVYNKVHITLSTHECAGLSERDINLASFIEQVAVSMT |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Pterin-4-alpha-carbinolamine dehydratase family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Involved in tetrahydrobiopterin biosynthesis. Seems to both prevent the formation of 7-pterins and accelerate the formation of quinonoid-BH2. Coactivator for HNF1A-dependent transcription. Regulates the dimerization of homeodomain protein HNF1A and enhances its transcriptional activity. Also acts as a coactivator for HNF1B-dependent transcription.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C-Sul Probe Info |
 |
18.62 | LDD0066 | [1] | |
|
BTD Probe Info |
 |
C82(1.05) | LDD2092 | [2] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [3] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [4] | |
|
m-APA Probe Info |
 |
N.A. | LDD2231 | [3] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C269 Probe Info |
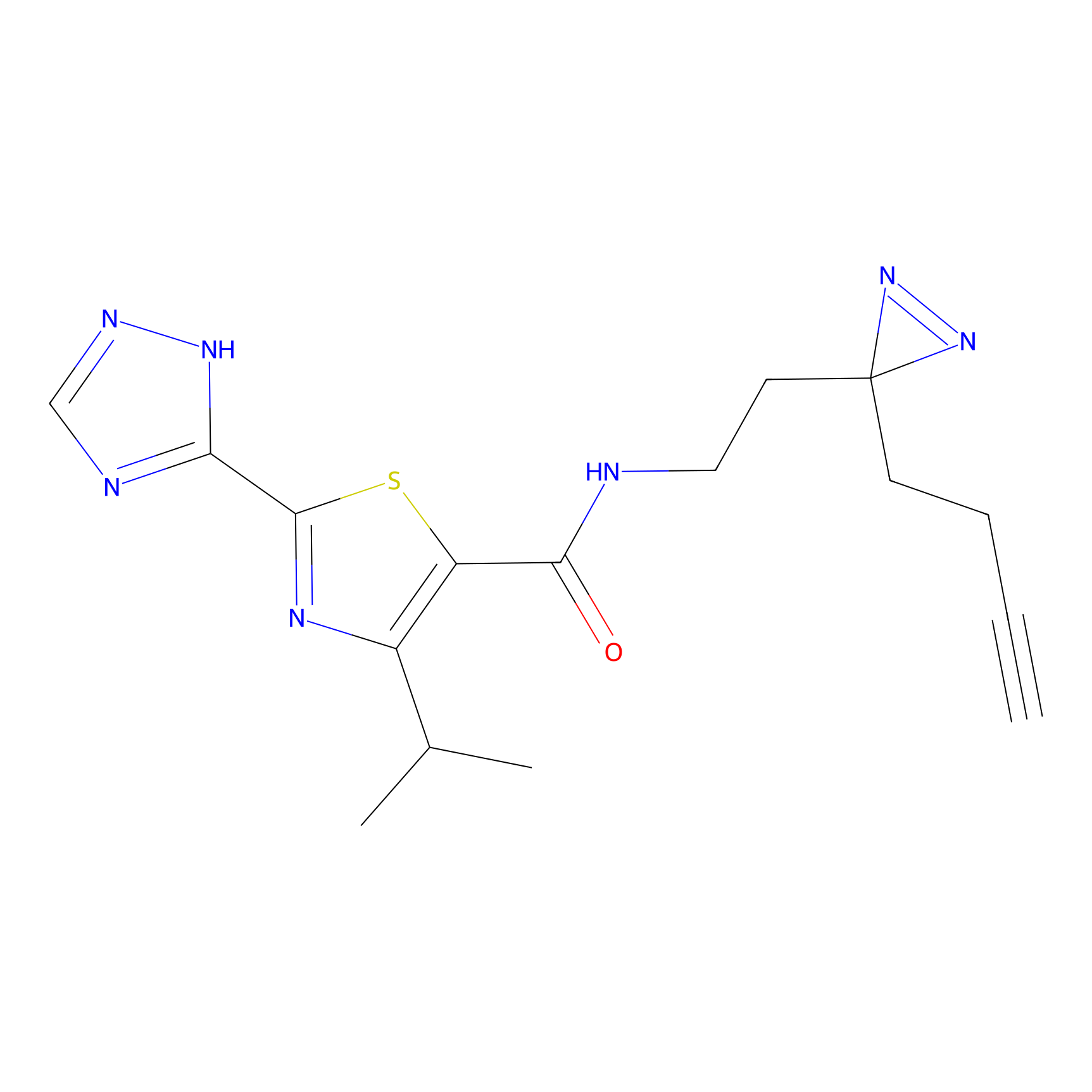 |
5.70 | LDD1939 | [5] | |
|
C278 Probe Info |
 |
61.39 | LDD1948 | [5] | |
|
C280 Probe Info |
 |
10.56 | LDD1950 | [5] | |
|
C282 Probe Info |
 |
16.22 | LDD1952 | [5] | |
|
C284 Probe Info |
 |
29.04 | LDD1954 | [5] | |
|
C348 Probe Info |
 |
13.74 | LDD2009 | [5] | |
|
C349 Probe Info |
 |
9.51 | LDD2010 | [5] | |
|
C350 Probe Info |
 |
26.72 | LDD2011 | [5] | |
|
C353 Probe Info |
 |
7.46 | LDD2014 | [5] | |
|
C354 Probe Info |
 |
11.63 | LDD2015 | [5] | |
|
C420 Probe Info |
 |
10.85 | LDD2075 | [5] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C82(0.26) | LDD2142 | [2] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C82(0.42) | LDD2112 | [2] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C82(0.59) | LDD2103 | [2] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C82(0.81) | LDD2113 | [2] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C82(1.11) | LDD2102 | [2] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C82(1.05) | LDD2092 | [2] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C82(2.25) | LDD2094 | [2] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C82(0.32) | LDD2096 | [2] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C82(0.94) | LDD2097 | [2] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C82(0.60) | LDD2100 | [2] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C82(0.96) | LDD2101 | [2] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C82(0.47) | LDD2104 | [2] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C82(0.81) | LDD2105 | [2] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C82(0.46) | LDD2108 | [2] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C82(0.80) | LDD2114 | [2] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C82(0.53) | LDD2115 | [2] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C82(0.52) | LDD2116 | [2] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C82(0.71) | LDD2120 | [2] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C82(0.71) | LDD2128 | [2] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C82(0.53) | LDD2134 | [2] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C82(0.67) | LDD2143 | [2] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C82(0.36) | LDD2148 | [2] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C82(0.34) | LDD2151 | [2] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Amyloid-beta precursor protein (APP) | APP family | P05067 | |||
| Apoptotic chromatin condensation inducer in the nucleus (ACIN1) | . | Q9UKV3 | |||
| Syntenin-1 (SDCBP) | . | O00560 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Hepatocyte nuclear factor 1-alpha (HNF1A) | HNF1 homeobox family | P20823 | |||
| Hepatocyte nuclear factor 1-beta (HNF1B) | HNF1 homeobox family | P35680 | |||
Other
References
