Details of the Target
General Information of Target
| Target ID | LDTP19892 | |||||
|---|---|---|---|---|---|---|
| Target Name | DGAT1/2-independent enzyme synthesizing storage lipids (TMEM68) | |||||
| Gene Name | TMEM68 | |||||
| Gene ID | 137695 | |||||
| Synonyms |
DGAT1/2-independent enzyme synthesizing storage lipids; DIESL; EC 2.3.1.-; 2-acylglycerol/1,2-diacylglycerol O-acyltransferase; Monoacylglycerol/Diacylglycerol O-acyltransferase; MGAT/DGAT; EC 2.3.1.20, EC 2.3.1.22; Transmembrane protein 68
|
|||||
| 3D Structure | ||||||
| Sequence |
MSSQKGNVARSRPQKHQNTFSFKNDKFDKSVQTKKINAKLHDGVCQRCKEVLEWRVKYSK
YKPLSKPKKCVKCLQKTVKDSYHIMCRPCACELEVCAKCGKKEDIVIPWSLPLLPRLECS GRILAHHNLRLPCSSDSPASASRVAGTTGAHHHAQLIFVFLVEMGFHYVGQAGLELLTS |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Subcellular location |
Membrane
|
|||||
| Function |
Catalytic subunit of the alternative triglyceride biosynthesis pathway, which mediates formation of triacylglycerol from diacylglycerol and membrane phospholipids. Synthesizes triacylglycerol at the expense of membrane phospholipids, such as phosphatidylcholine (PC) and its ether-linked form (ePC), thereby altering the composition of membranes. The alternative triglyceride biosynthesis pathway is probably required to provide the energy required for rapid growth when fuel sources are limiting. It maintains mitochondrial function during periods of extracellular lipid starvation. Can also use acyl-CoA as donor: acts as a acyl-CoA:monoacylglycerol acyltransferase (MGAT), but also shows acyl-CoA:diacylglycerol acyltransferase (DGAT) activity.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| KASUMI1 | SNV: p.V213I | . | |||
| NCIH1155 | Deletion: p.S317TfsTer19 | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
9.28 | LDD0402 | [1] | |
|
STPyne Probe Info |
 |
K307(8.91) | LDD0277 | [2] | |
|
DBIA Probe Info |
 |
C183(1.57) | LDD3311 | [3] | |
|
Jackson_1 Probe Info |
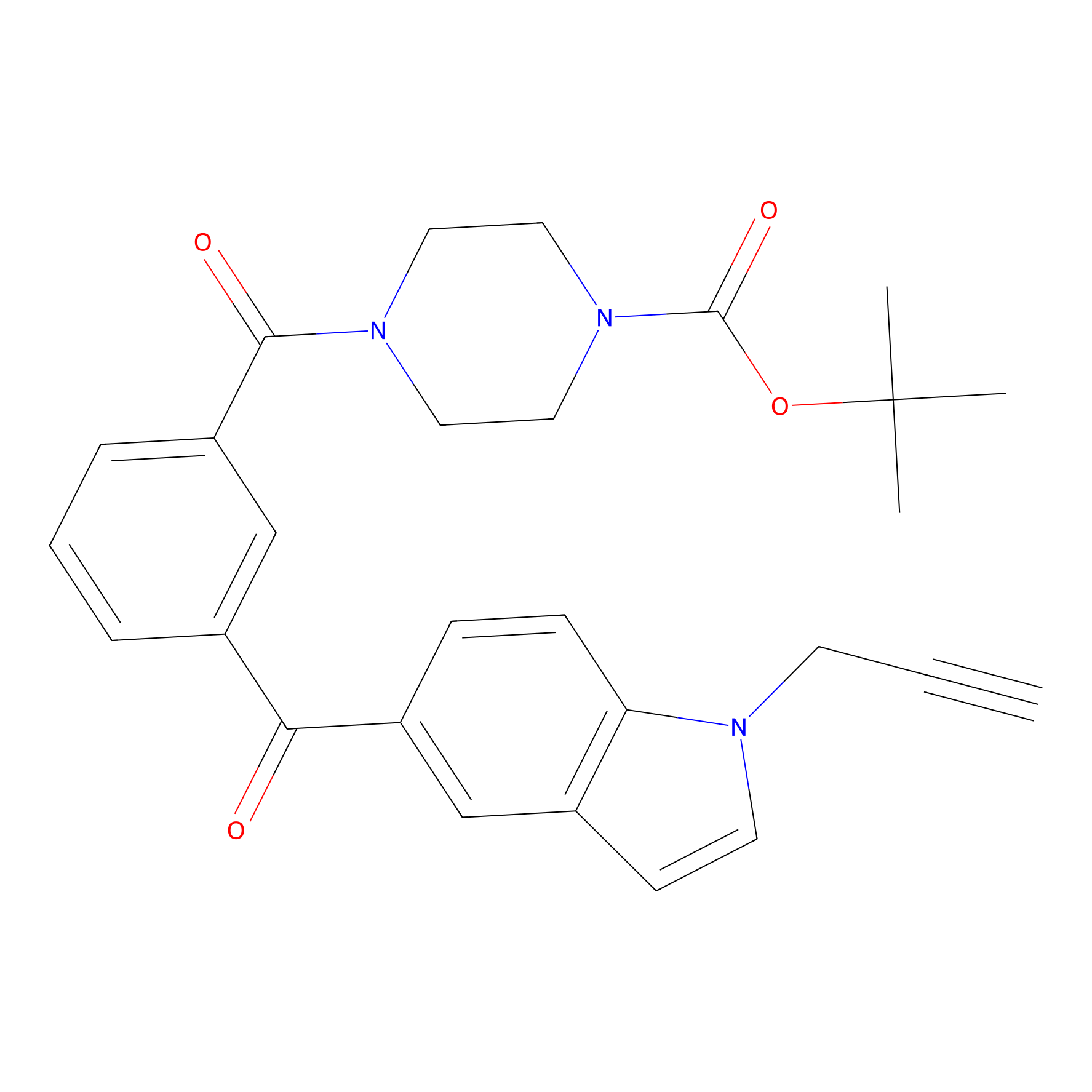 |
2.56 | LDD0120 | [4] | |
|
Jackson_14 Probe Info |
 |
3.33 | LDD0123 | [4] | |
|
Johansson_61 Probe Info |
 |
_(5.34) | LDD1485 | [5] | |
|
HPAP Probe Info |
 |
3.26 | LDD0063 | [6] | |
|
Alkyne-RA190 Probe Info |
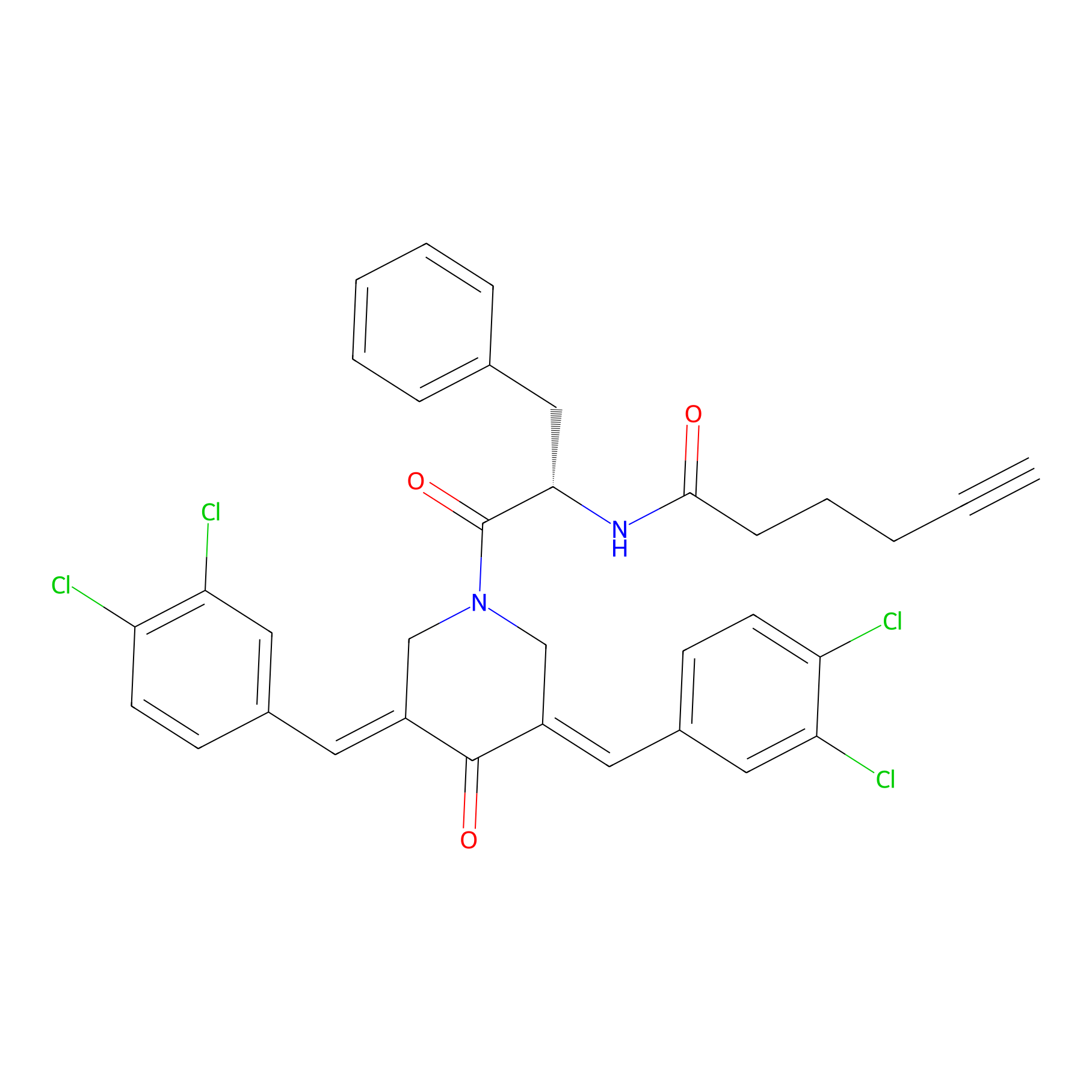 |
4.71 | LDD0299 | [7] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0165 | [8] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [9] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C053 Probe Info |
 |
14.72 | LDD1751 | [10] | |
|
C112 Probe Info |
 |
62.68 | LDD1799 | [10] | |
|
C407 Probe Info |
 |
23.10 | LDD2064 | [10] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0472 | [11] | |
|
FFF probe14 Probe Info |
 |
20.00 | LDD0477 | [11] | |
|
OEA-DA Probe Info |
 |
3.92 | LDD0046 | [12] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0259 | AC14 | HEK-293T | C174(0.85) | LDD1512 | [13] |
| LDCM0282 | AC22 | HEK-293T | C174(0.79) | LDD1521 | [13] |
| LDCM0291 | AC30 | HEK-293T | C174(0.83) | LDD1530 | [13] |
| LDCM0299 | AC38 | HEK-293T | C174(1.04) | LDD1538 | [13] |
| LDCM0308 | AC46 | HEK-293T | C174(0.92) | LDD1547 | [13] |
| LDCM0317 | AC54 | HEK-293T | C174(0.96) | LDD1556 | [13] |
| LDCM0323 | AC6 | HEK-293T | C174(0.85) | LDD1562 | [13] |
| LDCM0326 | AC62 | HEK-293T | C174(0.89) | LDD1565 | [13] |
| LDCM0156 | Aniline | NCI-H1299 | 12.31 | LDD0403 | [1] |
| LDCM0070 | Cisar_cp37 | MDA-MB-231 | 2.56 | LDD0120 | [4] |
| LDCM0367 | CL1 | HEK-293T | C174(1.06) | LDD1571 | [13] |
| LDCM0368 | CL10 | HEK-293T | C174(1.00) | LDD1572 | [13] |
| LDCM0369 | CL100 | HEK-293T | C174(0.98) | LDD1573 | [13] |
| LDCM0370 | CL101 | HEK-293T | C174(1.01) | LDD1574 | [13] |
| LDCM0373 | CL104 | HEK-293T | C174(1.31) | LDD1577 | [13] |
| LDCM0374 | CL105 | HEK-293T | C174(0.98) | LDD1578 | [13] |
| LDCM0377 | CL108 | HEK-293T | C174(1.09) | LDD1581 | [13] |
| LDCM0378 | CL109 | HEK-293T | C174(1.00) | LDD1582 | [13] |
| LDCM0382 | CL112 | HEK-293T | C174(0.99) | LDD1586 | [13] |
| LDCM0383 | CL113 | HEK-293T | C174(0.94) | LDD1587 | [13] |
| LDCM0386 | CL116 | HEK-293T | C174(1.14) | LDD1590 | [13] |
| LDCM0387 | CL117 | HEK-293T | C174(0.94) | LDD1591 | [13] |
| LDCM0391 | CL120 | HEK-293T | C174(1.16) | LDD1595 | [13] |
| LDCM0392 | CL121 | HEK-293T | C174(1.01) | LDD1596 | [13] |
| LDCM0395 | CL124 | HEK-293T | C174(0.91) | LDD1599 | [13] |
| LDCM0396 | CL125 | HEK-293T | C174(1.12) | LDD1600 | [13] |
| LDCM0399 | CL128 | HEK-293T | C174(0.91) | LDD1603 | [13] |
| LDCM0400 | CL13 | HEK-293T | C174(1.01) | LDD1604 | [13] |
| LDCM0403 | CL16 | HEK-293T | C174(1.14) | LDD1607 | [13] |
| LDCM0410 | CL22 | HEK-293T | C174(0.97) | LDD1614 | [13] |
| LDCM0413 | CL25 | HEK-293T | C174(1.10) | LDD1617 | [13] |
| LDCM0416 | CL28 | HEK-293T | C174(1.14) | LDD1620 | [13] |
| LDCM0423 | CL34 | HEK-293T | C174(1.10) | LDD1627 | [13] |
| LDCM0426 | CL37 | HEK-293T | C174(1.19) | LDD1630 | [13] |
| LDCM0429 | CL4 | HEK-293T | C174(0.93) | LDD1633 | [13] |
| LDCM0430 | CL40 | HEK-293T | C174(0.94) | LDD1634 | [13] |
| LDCM0436 | CL46 | HEK-293T | C174(1.24) | LDD1640 | [13] |
| LDCM0439 | CL49 | HEK-293T | C174(1.21) | LDD1643 | [13] |
| LDCM0443 | CL52 | HEK-293T | C174(1.13) | LDD1646 | [13] |
| LDCM0449 | CL58 | HEK-293T | C174(0.84) | LDD1652 | [13] |
| LDCM0453 | CL61 | HEK-293T | C174(1.03) | LDD1656 | [13] |
| LDCM0456 | CL64 | HEK-293T | C174(1.01) | LDD1659 | [13] |
| LDCM0463 | CL70 | HEK-293T | C174(1.33) | LDD1666 | [13] |
| LDCM0466 | CL73 | HEK-293T | C174(0.99) | LDD1669 | [13] |
| LDCM0469 | CL76 | HEK-293T | C174(1.45) | LDD1672 | [13] |
| LDCM0476 | CL82 | HEK-293T | C174(0.98) | LDD1679 | [13] |
| LDCM0479 | CL85 | HEK-293T | C174(1.17) | LDD1682 | [13] |
| LDCM0482 | CL88 | HEK-293T | C174(0.84) | LDD1685 | [13] |
| LDCM0489 | CL94 | HEK-293T | C174(0.92) | LDD1692 | [13] |
| LDCM0492 | CL97 | HEK-293T | C174(1.08) | LDD1695 | [13] |
| LDCM0616 | Fragment61 | Jurkat | _(20.00) | LDD1489 | [5] |
| LDCM0615 | Fragment63-R | Jurkat | _(8.63) | LDD1487 | [5] |
| LDCM0617 | Fragment63-S | Jurkat | _(5.96) | LDD1490 | [5] |
| LDCM0569 | Fragment7 | Jurkat | _(5.34) | LDD1485 | [5] |
| LDCM0022 | KB02 | 42-MG-BA | C183(2.58) | LDD2244 | [3] |
| LDCM0023 | KB03 | 769-P | C183(2.57) | LDD2663 | [3] |
| LDCM0024 | KB05 | G361 | C183(1.57) | LDD3311 | [3] |
| LDCM0014 | Panhematin | K562 | 3.26 | LDD0063 | [6] |
| LDCM0131 | RA190 | MM1.R | 4.71 | LDD0299 | [7] |
| LDCM0016 | Ranjitkar_cp1 | MDA-MB-231 | 3.33 | LDD0123 | [4] |
References
