Details of the Target
General Information of Target
| Target ID | LDTP09054 | |||||
|---|---|---|---|---|---|---|
| Target Name | Golgi membrane protein 1 (GOLM1) | |||||
| Gene Name | GOLM1 | |||||
| Gene ID | 51280 | |||||
| Synonyms |
C9orf155; GOLPH2; Golgi membrane protein 1; Golgi membrane protein GP73; Golgi phosphoprotein 2 |
|||||
| 3D Structure | ||||||
| Sequence |
MMGLGNGRRSMKSPPLVLAALVACIIVLGFNYWIASSRSVDLQTRIMELEGRVRRAAAER
GAVELKKNEFQGELEKQREQLDKIQSSHNFQLESVNKLYQDEKAVLVNNITTGERLIRVL QDQLKTLQRNYGRLQQDVLQFQKNQTNLERKFSYDLSQCINQMKEVKEQCEERIEEVTKK GNEAVASRDLSENNDQRQQLQALSEPQPRLQAAGLPHTEVPQGKGNVLGNSKSQTPAPSS EVVLDSKRQVEKEETNEIQVVNEEPQRDRLPQEPGREQVVEDRPVGGRGFGGAGELGQTP QVQAALSVSQENPEMEGPERDQLVIPDGQEEEQEAAGEGRNQQKLRGEDDYNMDENEAES ETDKQAALAGNDRNIDVFNVEDQKRDTINLLDQREKRNHTL |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
GOLM family
|
|||||
| Subcellular location |
Golgi apparatus, cis-Golgi network membrane
|
|||||
| Function | Unknown. Cellular response protein to viral infection. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| AGS | SNV: p.M163R | DBIA Probe Info | |||
| FTC133 | SNV: p.C170R | . | |||
| HL60 | SNV: p.L42V | . | |||
| ICC3 | SNV: p.L401H | . | |||
| MEWO | SNV: p.P271A | . | |||
| MFE319 | SNV: p.M1? | . | |||
| NCIH1155 | SNV: p.G61S | . | |||
| NCIH1793 | SNV: p.A62S | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FBPP2 Probe Info |
 |
36.55 | LDD0318 | [1] | |
|
STPyne Probe Info |
 |
K125(7.65); K67(7.67) | LDD0277 | [2] | |
|
ONAyne Probe Info |
 |
K66(7.14) | LDD0275 | [2] | |
|
DBIA Probe Info |
 |
C159(2.13) | LDD3310 | [3] | |
|
N1 Probe Info |
 |
N.A. | LDD0245 | [4] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0162 | [5] | |
|
IPM Probe Info |
 |
N.A. | LDD2156 | [6] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [7] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C073 Probe Info |
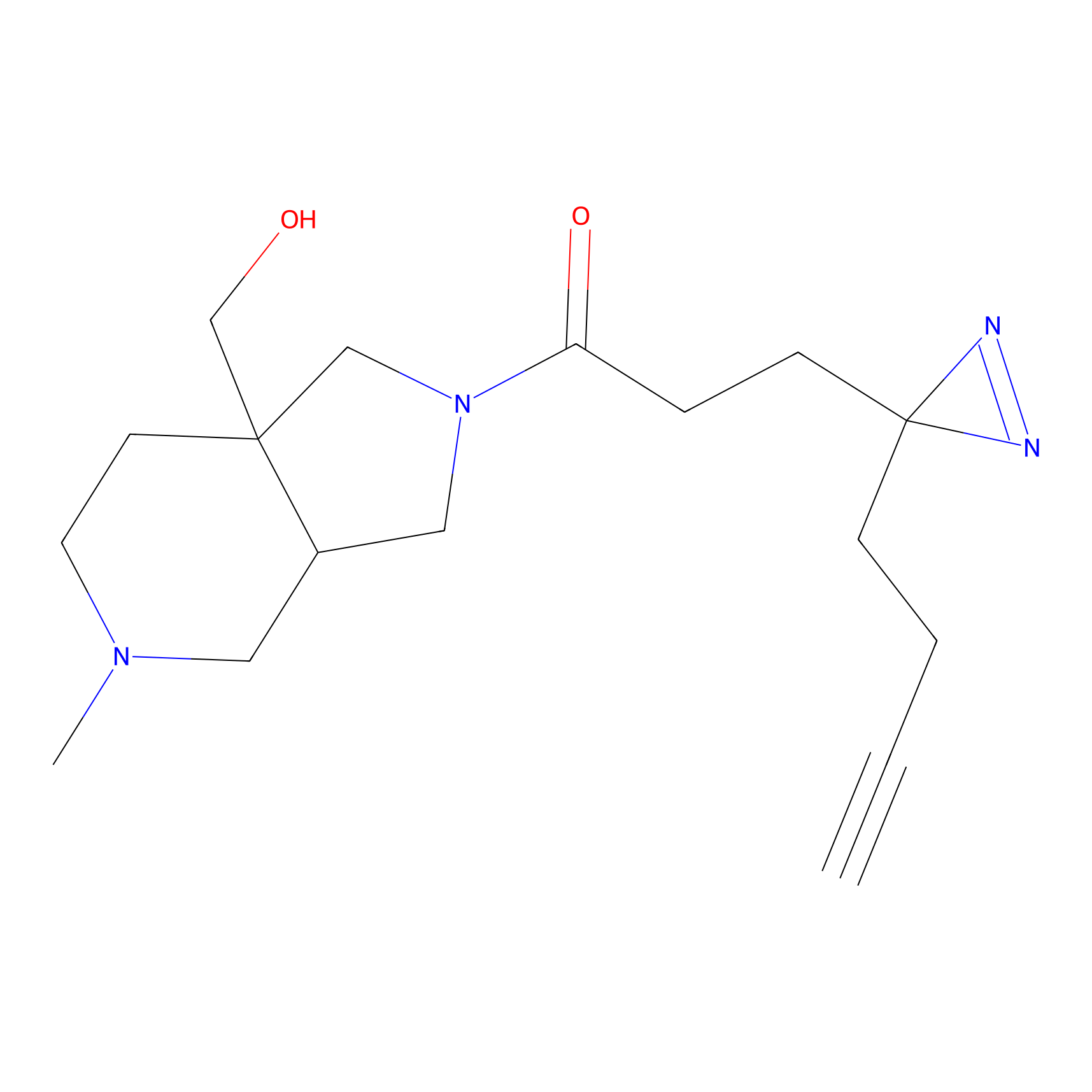 |
4.96 | LDD1769 | [8] | |
|
C170 Probe Info |
 |
16.56 | LDD1850 | [8] | |
|
C221 Probe Info |
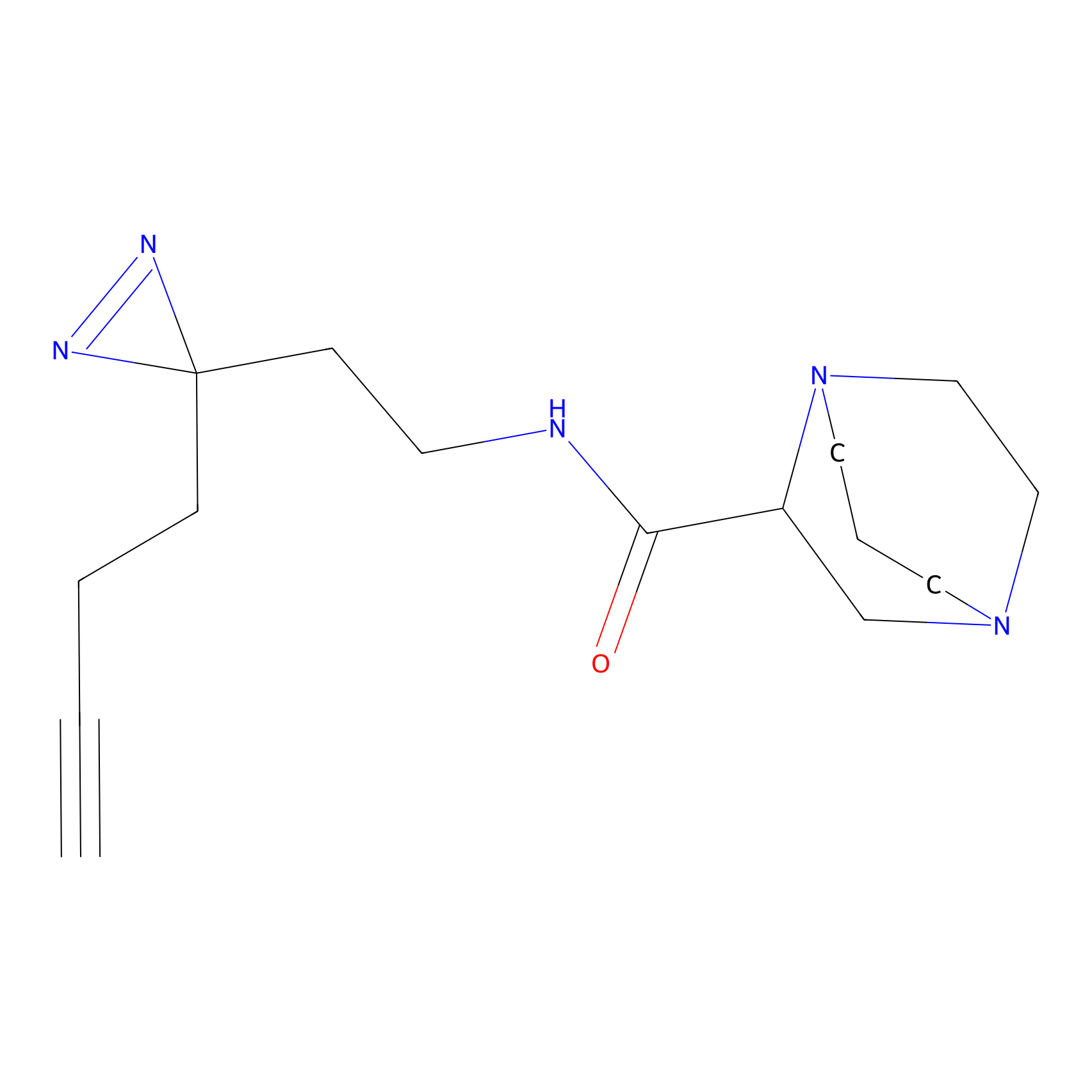 |
6.19 | LDD1895 | [8] | |
|
C237 Probe Info |
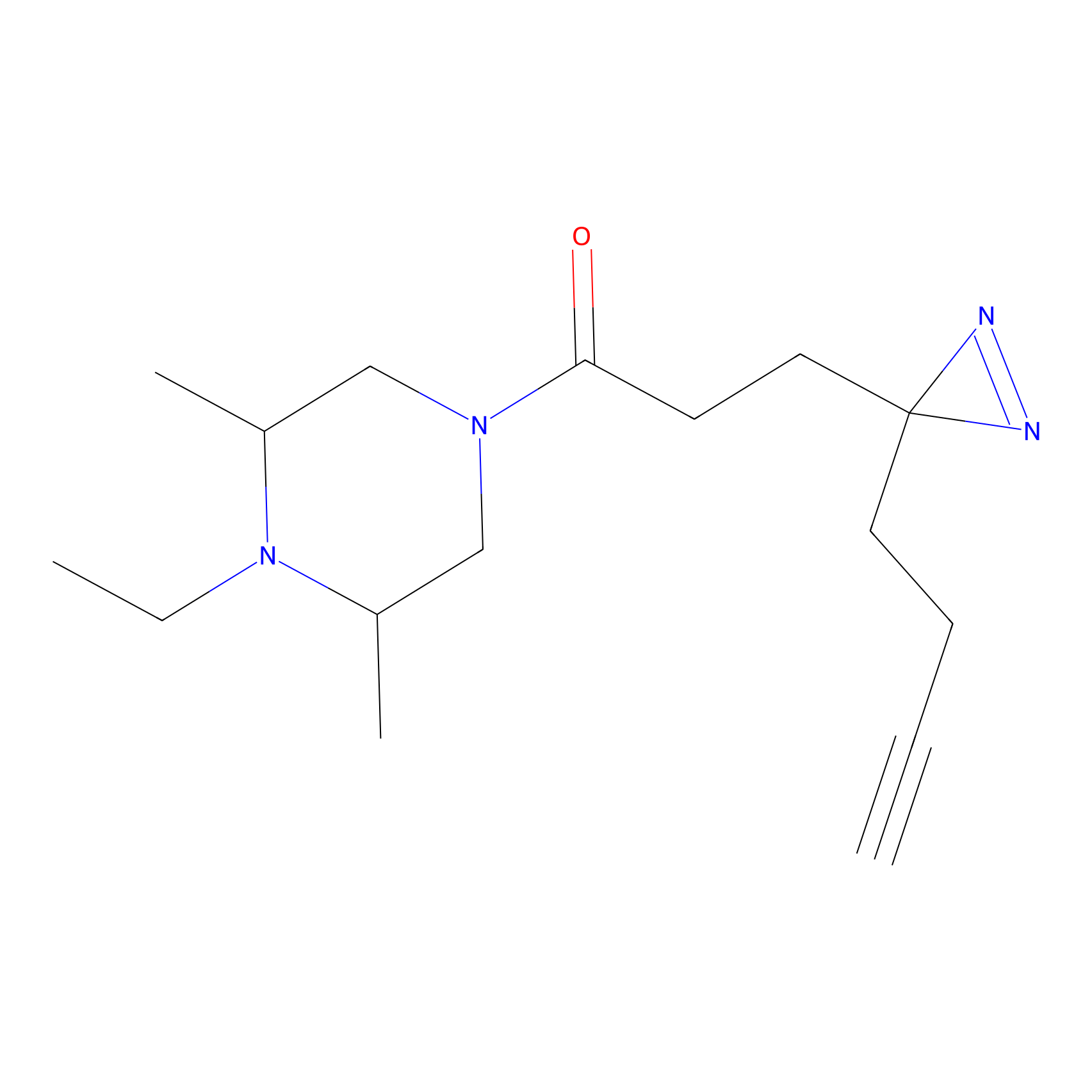 |
7.84 | LDD1910 | [8] | |
|
C310 Probe Info |
 |
19.97 | LDD1977 | [8] | |
|
C346 Probe Info |
 |
10.48 | LDD2007 | [8] | |
|
C376 Probe Info |
 |
7.67 | LDD2036 | [8] | |
|
C425 Probe Info |
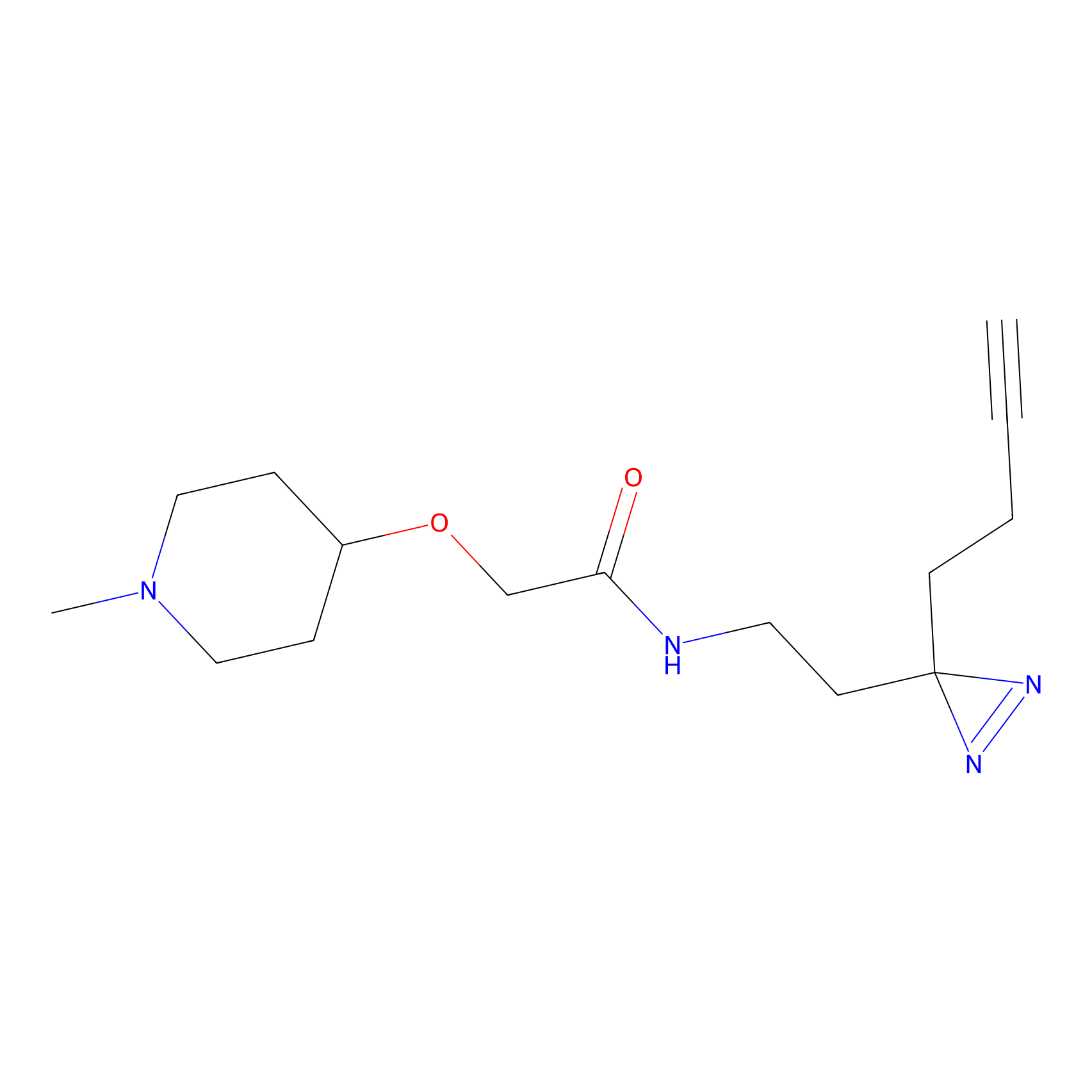 |
5.98 | LDD2080 | [8] | |
|
FFF probe12 Probe Info |
 |
20.00 | LDD0473 | [9] | |
|
FFF probe9 Probe Info |
 |
20.00 | LDD0470 | [9] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0226 | AC11 | HEK-293T | C159(1.34) | LDD1509 | [10] |
| LDCM0278 | AC19 | HEK-293T | C159(2.18) | LDD1517 | [10] |
| LDCM0281 | AC21 | HEK-293T | C159(1.03) | LDD1520 | [10] |
| LDCM0284 | AC24 | HEK-293T | C159(1.08) | LDD1523 | [10] |
| LDCM0287 | AC27 | HEK-293T | C159(1.08) | LDD1526 | [10] |
| LDCM0289 | AC29 | HEK-293T | C159(0.96) | LDD1528 | [10] |
| LDCM0290 | AC3 | HEK-293T | C159(1.24) | LDD1529 | [10] |
| LDCM0293 | AC32 | HEK-293T | C159(0.78) | LDD1532 | [10] |
| LDCM0296 | AC35 | HEK-293T | C159(1.07) | LDD1535 | [10] |
| LDCM0298 | AC37 | HEK-293T | C159(0.93) | LDD1537 | [10] |
| LDCM0302 | AC40 | HEK-293T | C159(1.07) | LDD1541 | [10] |
| LDCM0305 | AC43 | HEK-293T | C159(1.05) | LDD1544 | [10] |
| LDCM0307 | AC45 | HEK-293T | C159(1.14) | LDD1546 | [10] |
| LDCM0310 | AC48 | HEK-293T | C159(1.00) | LDD1549 | [10] |
| LDCM0312 | AC5 | HEK-293T | C159(1.02) | LDD1551 | [10] |
| LDCM0314 | AC51 | HEK-293T | C159(1.39) | LDD1553 | [10] |
| LDCM0316 | AC53 | HEK-293T | C159(1.13) | LDD1555 | [10] |
| LDCM0319 | AC56 | HEK-293T | C159(1.00) | LDD1558 | [10] |
| LDCM0322 | AC59 | HEK-293T | C159(1.10) | LDD1561 | [10] |
| LDCM0325 | AC61 | HEK-293T | C159(0.97) | LDD1564 | [10] |
| LDCM0328 | AC64 | HEK-293T | C159(0.89) | LDD1567 | [10] |
| LDCM0345 | AC8 | HEK-293T | C159(1.06) | LDD1569 | [10] |
| LDCM0248 | AKOS034007472 | HEK-293T | C159(1.16) | LDD1511 | [10] |
| LDCM0275 | AKOS034007705 | HEK-293T | C159(1.14) | LDD1514 | [10] |
| LDCM0367 | CL1 | HEK-293T | C159(0.97) | LDD1571 | [10] |
| LDCM0370 | CL101 | HEK-293T | C159(1.13) | LDD1574 | [10] |
| LDCM0374 | CL105 | HEK-293T | C159(1.13) | LDD1578 | [10] |
| LDCM0378 | CL109 | HEK-293T | C159(0.98) | LDD1582 | [10] |
| LDCM0383 | CL113 | HEK-293T | C159(0.95) | LDD1587 | [10] |
| LDCM0387 | CL117 | HEK-293T | C159(1.06) | LDD1591 | [10] |
| LDCM0390 | CL12 | HEK-293T | C159(1.27) | LDD1594 | [10] |
| LDCM0392 | CL121 | HEK-293T | C159(1.05) | LDD1596 | [10] |
| LDCM0396 | CL125 | HEK-293T | C159(0.92) | LDD1600 | [10] |
| LDCM0400 | CL13 | HEK-293T | C159(1.18) | LDD1604 | [10] |
| LDCM0406 | CL19 | HEK-293T | C159(1.16) | LDD1610 | [10] |
| LDCM0409 | CL21 | HEK-293T | C159(0.86) | LDD1613 | [10] |
| LDCM0412 | CL24 | HEK-293T | C159(0.95) | LDD1616 | [10] |
| LDCM0413 | CL25 | HEK-293T | C159(0.94) | LDD1617 | [10] |
| LDCM0420 | CL31 | HEK-293T | C159(1.04) | LDD1624 | [10] |
| LDCM0422 | CL33 | HEK-293T | C159(1.37) | LDD1626 | [10] |
| LDCM0425 | CL36 | HEK-293T | C159(1.11) | LDD1629 | [10] |
| LDCM0426 | CL37 | HEK-293T | C159(1.07) | LDD1630 | [10] |
| LDCM0433 | CL43 | HEK-293T | C159(1.02) | LDD1637 | [10] |
| LDCM0435 | CL45 | HEK-293T | C159(1.37) | LDD1639 | [10] |
| LDCM0438 | CL48 | HEK-293T | C159(1.14) | LDD1642 | [10] |
| LDCM0439 | CL49 | HEK-293T | C159(1.00) | LDD1643 | [10] |
| LDCM0446 | CL55 | HEK-293T | C159(1.06) | LDD1649 | [10] |
| LDCM0448 | CL57 | HEK-293T | C159(1.18) | LDD1651 | [10] |
| LDCM0452 | CL60 | HEK-293T | C159(1.21) | LDD1655 | [10] |
| LDCM0453 | CL61 | HEK-293T | C159(1.03) | LDD1656 | [10] |
| LDCM0459 | CL67 | HEK-293T | C159(1.01) | LDD1662 | [10] |
| LDCM0461 | CL69 | HEK-293T | C159(1.08) | LDD1664 | [10] |
| LDCM0462 | CL7 | HEK-293T | C159(1.26) | LDD1665 | [10] |
| LDCM0465 | CL72 | HEK-293T | C159(1.37) | LDD1668 | [10] |
| LDCM0466 | CL73 | HEK-293T | C159(1.01) | LDD1669 | [10] |
| LDCM0472 | CL79 | HEK-293T | C159(1.38) | LDD1675 | [10] |
| LDCM0475 | CL81 | HEK-293T | C159(0.99) | LDD1678 | [10] |
| LDCM0478 | CL84 | HEK-293T | C159(1.21) | LDD1681 | [10] |
| LDCM0479 | CL85 | HEK-293T | C159(1.02) | LDD1682 | [10] |
| LDCM0484 | CL9 | HEK-293T | C159(0.88) | LDD1687 | [10] |
| LDCM0486 | CL91 | HEK-293T | C159(1.32) | LDD1689 | [10] |
| LDCM0488 | CL93 | HEK-293T | C159(1.20) | LDD1691 | [10] |
| LDCM0491 | CL96 | HEK-293T | C159(1.38) | LDD1694 | [10] |
| LDCM0492 | CL97 | HEK-293T | C159(1.00) | LDD1695 | [10] |
| LDCM0022 | KB02 | HEK-293T | C159(0.97) | LDD1492 | [10] |
| LDCM0023 | KB03 | HEK-293T | C159(1.09) | LDD1497 | [10] |
| LDCM0024 | KB05 | COLO792 | C159(2.13) | LDD3310 | [3] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Other
References
