Details of the Target
General Information of Target
| Target ID | LDTP04107 | |||||
|---|---|---|---|---|---|---|
| Target Name | Large ribosomal subunit protein eL21 (RPL21) | |||||
| Gene Name | RPL21 | |||||
| Gene ID | 6144 | |||||
| Synonyms |
Large ribosomal subunit protein eL21; 60S ribosomal protein L21 |
|||||
| 3D Structure | ||||||
| Sequence |
MTNTKGKRRGTRYMFSRPFRKHGVVPLATYMRIYKKGDIVDIKGMGTVQKGMPHKCYHGK
TGRVYNVTQHAVGIVVNKQVKGKILAKRINVRIEHIKHSKSRDSFLKRVKENDQKKKEAK EKGTWVQLKRQPAPPREAHFVRTNGKEPELLEPIPYEFMA |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Eukaryotic ribosomal protein eL21 family
|
|||||
| Subcellular location |
Cytoplasm, cytosol
|
|||||
| Function | Component of the large ribosomal subunit. The ribosome is a large ribonucleoprotein complex responsible for the synthesis of proteins in the cell. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
14.39 | LDD0402 | [1] | |
|
A-EBA Probe Info |
 |
2.48 | LDD0215 | [2] | |
|
C-Sul Probe Info |
 |
3.24 | LDD0066 | [3] | |
|
TH211 Probe Info |
 |
Y65(20.00) | LDD0257 | [4] | |
|
TH214 Probe Info |
 |
Y65(11.48) | LDD0258 | [4] | |
|
TH216 Probe Info |
 |
Y30(20.00); Y65(20.00) | LDD0259 | [4] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [5] | |
|
ONAyne Probe Info |
 |
K36(1.00) | LDD0274 | [6] | |
|
STPyne Probe Info |
 |
K122(5.53); K21(1.21); K36(0.85); K97(10.00) | LDD0277 | [6] | |
|
BTD Probe Info |
 |
C56(1.06) | LDD2117 | [7] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [8] | |
|
HHS-482 Probe Info |
 |
Y156(1.24); Y30(0.87); Y65(0.85) | LDD0285 | [9] | |
|
HHS-465 Probe Info |
 |
Y156(0.73); Y65(10.00) | LDD2237 | [10] | |
|
DBIA Probe Info |
 |
C56(1.04) | LDD1510 | [11] | |
|
ATP probe Probe Info |
 |
K122(0.00); K78(0.00); K81(0.00); K36(0.00) | LDD0199 | [12] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0165 | [13] | |
|
ATP probe Probe Info |
 |
K43(0.00); K78(0.00) | LDD0035 | [14] | |
|
IPM Probe Info |
 |
N.A. | LDD0147 | [15] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [16] | |
|
OSF Probe Info |
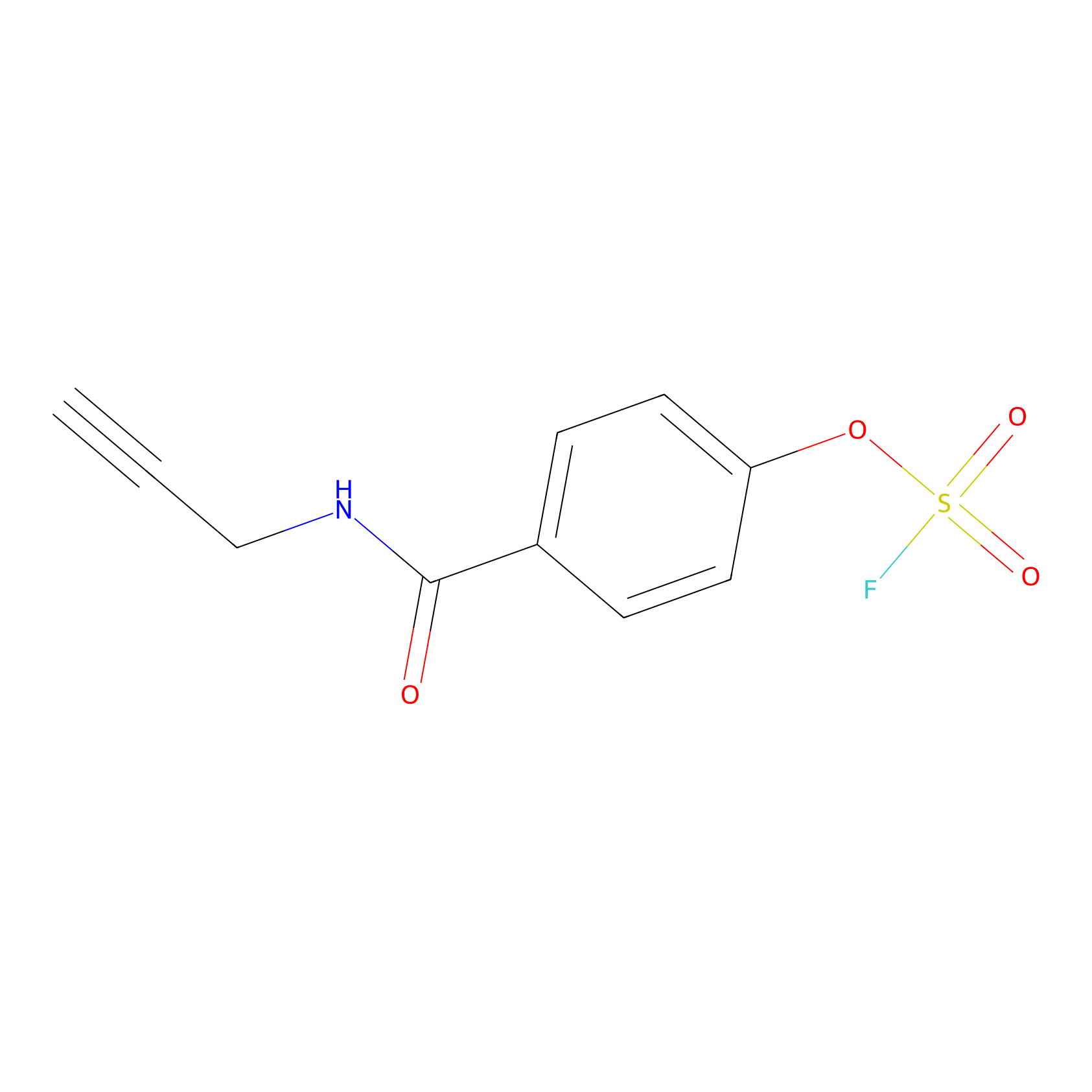 |
H70(0.00); Y65(0.00) | LDD0029 | [17] | |
|
SF Probe Info |
 |
K78(0.00); K97(0.00); Y156(0.00); Y30(0.00) | LDD0028 | [17] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [15] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [18] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [19] | |
|
1c-yne Probe Info |
 |
K43(0.00); K21(0.00) | LDD0228 | [20] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0357 | [20] | |
|
Acrolein Probe Info |
 |
H70(0.00); H139(0.00) | LDD0217 | [21] | |
|
Crotonaldehyde Probe Info |
 |
H70(0.00); H98(0.00); H139(0.00) | LDD0219 | [21] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [21] | |
|
AOyne Probe Info |
 |
12.90 | LDD0443 | [22] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [23] | |
|
HHS-475 Probe Info |
 |
Y65(1.22) | LDD2238 | [10] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-1 Probe Info |
 |
N.A. | LDD0137 | [24] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [24] | |
|
Dasatinib-CA-3PAP Probe Info |
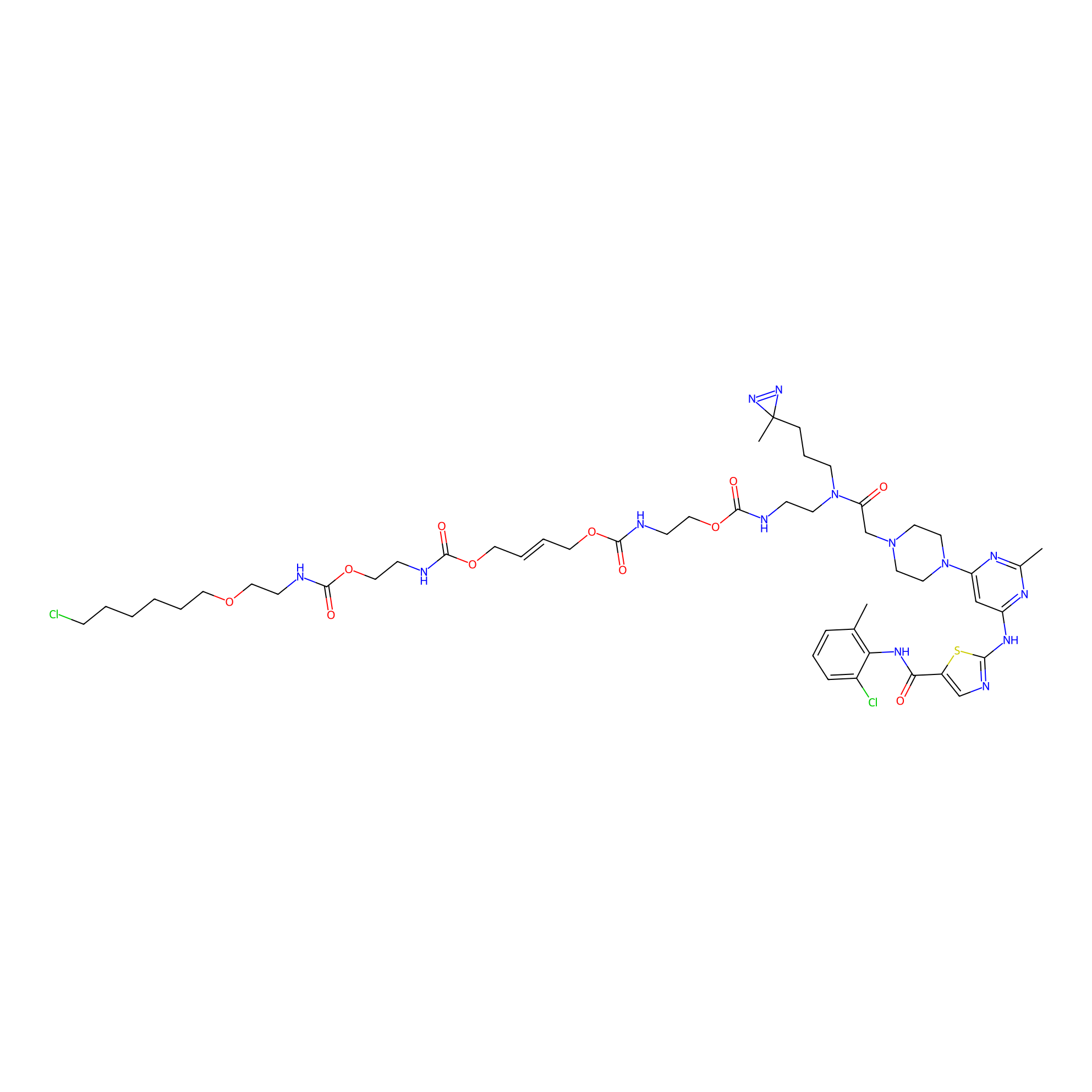 |
6.50 | LDD0365 | [25] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0071 | [26] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C56(1.06) | LDD2117 | [7] |
| LDCM0237 | AC12 | HEK-293T | C56(1.04) | LDD1510 | [11] |
| LDCM0280 | AC20 | HEK-293T | C56(1.09) | LDD1519 | [11] |
| LDCM0288 | AC28 | HEK-293T | C56(1.02) | LDD1527 | [11] |
| LDCM0297 | AC36 | HEK-293T | C56(1.12) | LDD1536 | [11] |
| LDCM0301 | AC4 | HEK-293T | C56(1.11) | LDD1540 | [11] |
| LDCM0306 | AC44 | HEK-293T | C56(1.07) | LDD1545 | [11] |
| LDCM0315 | AC52 | HEK-293T | C56(1.00) | LDD1554 | [11] |
| LDCM0324 | AC60 | HEK-293T | C56(0.92) | LDD1563 | [11] |
| LDCM0156 | Aniline | NCI-H1299 | 12.30 | LDD0403 | [1] |
| LDCM0108 | Chloroacetamide | HeLa | H70(0.00); H98(0.00); H139(0.00) | LDD0222 | [21] |
| LDCM0632 | CL-Sc | Hep-G2 | C56(0.42) | LDD2227 | [23] |
| LDCM0372 | CL103 | HEK-293T | C56(1.19) | LDD1576 | [11] |
| LDCM0376 | CL107 | HEK-293T | C56(1.06) | LDD1580 | [11] |
| LDCM0381 | CL111 | HEK-293T | C56(1.11) | LDD1585 | [11] |
| LDCM0385 | CL115 | HEK-293T | C56(1.13) | LDD1589 | [11] |
| LDCM0389 | CL119 | HEK-293T | C56(1.09) | LDD1593 | [11] |
| LDCM0394 | CL123 | HEK-293T | C56(1.10) | LDD1598 | [11] |
| LDCM0398 | CL127 | HEK-293T | C56(0.87) | LDD1602 | [11] |
| LDCM0402 | CL15 | HEK-293T | C56(1.05) | LDD1606 | [11] |
| LDCM0408 | CL20 | HEK-293T | C56(0.69) | LDD1612 | [11] |
| LDCM0415 | CL27 | HEK-293T | C56(1.06) | LDD1619 | [11] |
| LDCM0418 | CL3 | HEK-293T | C56(0.87) | LDD1622 | [11] |
| LDCM0421 | CL32 | HEK-293T | C56(0.70) | LDD1625 | [11] |
| LDCM0428 | CL39 | HEK-293T | C56(1.25) | LDD1632 | [11] |
| LDCM0434 | CL44 | HEK-293T | C56(0.74) | LDD1638 | [11] |
| LDCM0447 | CL56 | HEK-293T | C56(0.80) | LDD1650 | [11] |
| LDCM0455 | CL63 | HEK-293T | C56(1.09) | LDD1658 | [11] |
| LDCM0460 | CL68 | HEK-293T | C56(0.68) | LDD1663 | [11] |
| LDCM0473 | CL8 | HEK-293T | C56(0.65) | LDD1676 | [11] |
| LDCM0474 | CL80 | HEK-293T | C56(0.72) | LDD1677 | [11] |
| LDCM0481 | CL87 | HEK-293T | C56(0.96) | LDD1684 | [11] |
| LDCM0487 | CL92 | HEK-293T | C56(0.79) | LDD1690 | [11] |
| LDCM0494 | CL99 | HEK-293T | C56(0.99) | LDD1697 | [11] |
| LDCM0097 | Dasatinib | K562 | 6.50 | LDD0365 | [25] |
| LDCM0495 | E2913 | HEK-293T | C56(1.25) | LDD1698 | [11] |
| LDCM0175 | Ethacrynic acid | HeLa | N.A. | LDD0440 | [8] |
| LDCM0573 | Fragment11 | Ramos | C56(20.00) | LDD2190 | [27] |
| LDCM0576 | Fragment14 | Ramos | C56(0.29) | LDD2193 | [27] |
| LDCM0586 | Fragment28 | Ramos | C56(0.69) | LDD2198 | [27] |
| LDCM0468 | Fragment33 | HEK-293T | C56(1.18) | LDD1671 | [11] |
| LDCM0566 | Fragment4 | Ramos | C56(0.64) | LDD2184 | [27] |
| LDCM0569 | Fragment7 | Ramos | C56(0.43) | LDD2186 | [27] |
| LDCM0107 | IAA | HeLa | H70(0.00); H98(0.00); H139(0.00) | LDD0221 | [21] |
| LDCM0123 | JWB131 | DM93 | Y156(1.24); Y30(0.87); Y65(0.85) | LDD0285 | [9] |
| LDCM0124 | JWB142 | DM93 | Y156(0.76); Y30(0.47); Y65(0.73) | LDD0286 | [9] |
| LDCM0125 | JWB146 | DM93 | Y156(1.03); Y30(0.53); Y65(0.93) | LDD0287 | [9] |
| LDCM0126 | JWB150 | DM93 | Y156(2.61); Y30(1.13); Y65(2.95) | LDD0288 | [9] |
| LDCM0127 | JWB152 | DM93 | Y156(1.24); Y30(0.81); Y65(1.91) | LDD0289 | [9] |
| LDCM0128 | JWB198 | DM93 | Y156(1.05); Y30(0.47); Y65(1.34) | LDD0290 | [9] |
| LDCM0129 | JWB202 | DM93 | Y156(0.44); Y30(0.28); Y65(0.44) | LDD0291 | [9] |
| LDCM0130 | JWB211 | DM93 | Y156(1.42); Y30(0.45); Y65(0.87) | LDD0292 | [9] |
| LDCM0022 | KB02 | Ramos | C56(0.54) | LDD2182 | [27] |
| LDCM0023 | KB03 | Ramos | C56(0.62) | LDD2183 | [27] |
| LDCM0024 | KB05 | Ramos | C56(0.38) | LDD2185 | [27] |
| LDCM0109 | NEM | HeLa | H70(0.00); H139(0.00); H98(0.00) | LDD0223 | [21] |
References
