Details of the Target
General Information of Target
| Target ID | LDTP11331 | |||||
|---|---|---|---|---|---|---|
| Target Name | Sideroflexin-3 (SFXN3) | |||||
| Gene Name | SFXN3 | |||||
| Gene ID | 81855 | |||||
| Synonyms |
Sideroflexin-3 |
|||||
| 3D Structure | ||||||
| Sequence |
MASGSNWLSGVNVVLVMAYGSLVFVLLFIFVKRQIMRFAMKSRRGPHVPVGHNAPKDLKE
EIDIRLSRVQDIKYEPQLLADDDARLLQLETQGNQSCYNYLYRMKALDAIRTSEIPFHSE GRHPRSLMGKNFRSYLLDLRNTSTPFKGVRKALIDTLLDGYETARYGTGVFGQNEYLRYQ EALSELATAVKARIGSSQRHHQSAAKDLTQSPEVSPTTIQVTYLPSSQKSKRAKHFLELK SFKDNYNTLESTL |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Sideroflexin family
|
|||||
| Subcellular location |
Mitochondrion membrane
|
|||||
| Function |
Mitochondrial serine transporter that mediates transport of serine into mitochondria, an important step of the one-carbon metabolism pathway. Mitochondrial serine is converted to glycine and formate, which then exits to the cytosol where it is used to generate the charged folates that serve as one-carbon donors.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
10.41 | LDD0402 | [1] | |
|
STPyne Probe Info |
 |
K169(4.51); K319(10.00); K71(7.49); K85(5.00) | LDD0277 | [2] | |
|
DBIA Probe Info |
 |
C234(2.11) | LDD3310 | [3] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [4] | |
|
Acrolein Probe Info |
 |
H26(0.00); H293(0.00) | LDD0223 | [5] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [6] | |
|
IA-alkyne Probe Info |
 |
C234(0.00); C189(0.00) | LDD0162 | [7] | |
|
TFBX Probe Info |
 |
C234(0.00); C189(0.00) | LDD0027 | [8] | |
|
IPM Probe Info |
 |
N.A. | LDD2156 | [9] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [5] | |
|
AOyne Probe Info |
 |
12.70 | LDD0443 | [10] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [11] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-1 Probe Info |
 |
N.A. | LDD0137 | [12] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [12] | |
|
Staurosporine capture compound Probe Info |
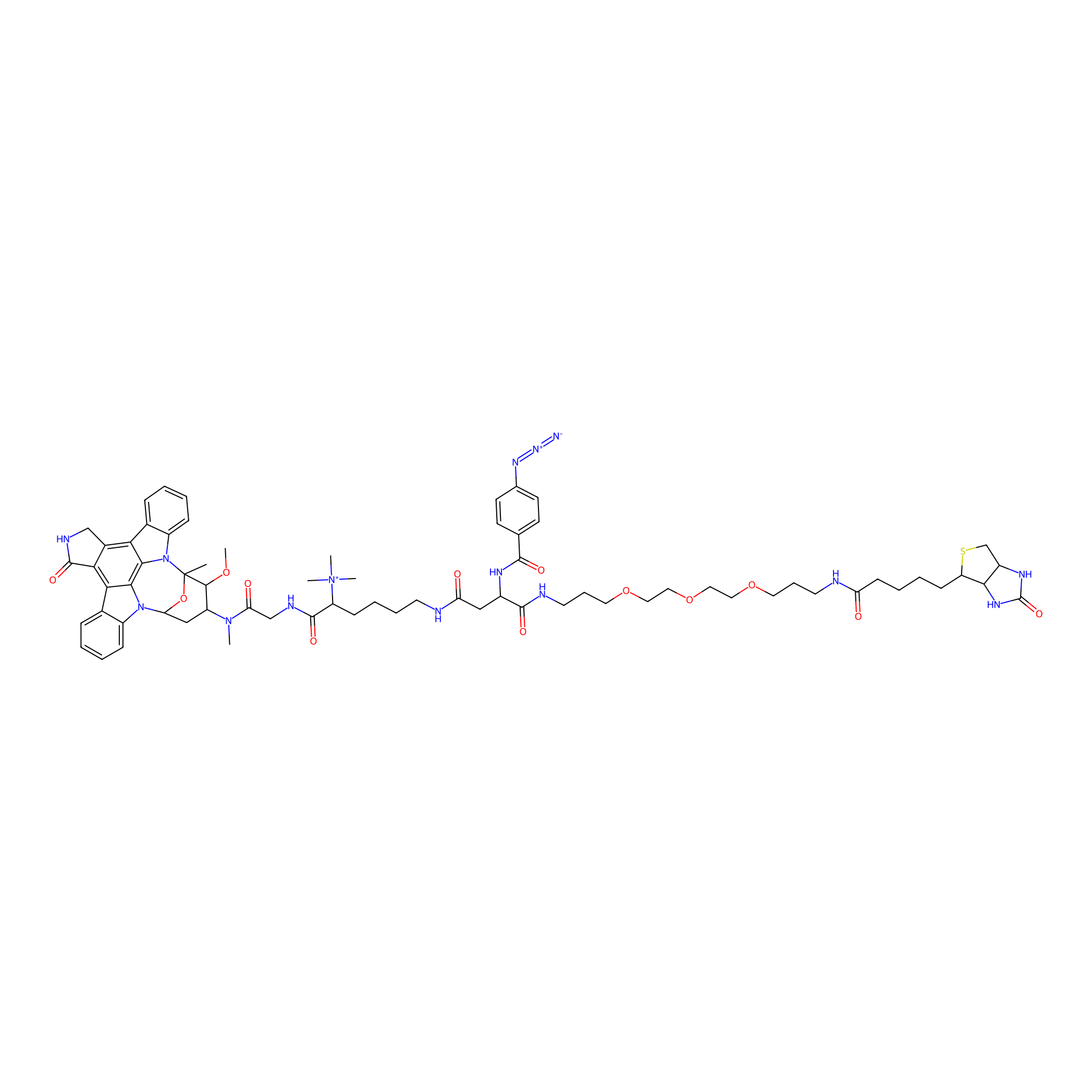 |
N.A. | LDD0083 | [13] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0561 | Abegg_cp(-)-10 | HeLa | C189(4.80) | LDD0312 | [8] |
| LDCM0214 | AC1 | PaTu 8988t | C109(0.28) | LDD1093 | [14] |
| LDCM0215 | AC10 | PaTu 8988t | C109(0.67); C193(0.53) | LDD1094 | [14] |
| LDCM0226 | AC11 | PaTu 8988t | C109(0.84); C193(0.81) | LDD1105 | [14] |
| LDCM0237 | AC12 | PaTu 8988t | C109(0.66); C193(0.71) | LDD1116 | [14] |
| LDCM0246 | AC128 | PaTu 8988t | C193(0.94) | LDD1125 | [14] |
| LDCM0247 | AC129 | PaTu 8988t | C193(1.26) | LDD1126 | [14] |
| LDCM0249 | AC130 | PaTu 8988t | C193(1.21) | LDD1128 | [14] |
| LDCM0250 | AC131 | PaTu 8988t | C193(1.22) | LDD1129 | [14] |
| LDCM0251 | AC132 | PaTu 8988t | C193(0.66) | LDD1130 | [14] |
| LDCM0252 | AC133 | PaTu 8988t | C193(0.63) | LDD1131 | [14] |
| LDCM0253 | AC134 | PaTu 8988t | C193(0.87) | LDD1132 | [14] |
| LDCM0254 | AC135 | PaTu 8988t | C193(0.89) | LDD1133 | [14] |
| LDCM0255 | AC136 | PaTu 8988t | C193(0.62) | LDD1134 | [14] |
| LDCM0256 | AC137 | PaTu 8988t | C193(0.70) | LDD1135 | [14] |
| LDCM0257 | AC138 | PaTu 8988t | C193(1.16) | LDD1136 | [14] |
| LDCM0258 | AC139 | PaTu 8988t | C193(0.88) | LDD1137 | [14] |
| LDCM0259 | AC14 | PaTu 8988t | C109(0.88); C193(0.73) | LDD1138 | [14] |
| LDCM0260 | AC140 | PaTu 8988t | C193(0.91) | LDD1139 | [14] |
| LDCM0261 | AC141 | PaTu 8988t | C193(1.38) | LDD1140 | [14] |
| LDCM0262 | AC142 | PaTu 8988t | C193(1.15) | LDD1141 | [14] |
| LDCM0263 | AC143 | PaTu 8988t | C109(0.67); C193(0.70) | LDD1142 | [14] |
| LDCM0264 | AC144 | PaTu 8988t | C109(0.58); C193(0.54) | LDD1143 | [14] |
| LDCM0265 | AC145 | PaTu 8988t | C109(0.79); C193(0.61) | LDD1144 | [14] |
| LDCM0266 | AC146 | PaTu 8988t | C109(0.35); C193(0.76) | LDD1145 | [14] |
| LDCM0267 | AC147 | PaTu 8988t | C109(0.63); C193(0.85) | LDD1146 | [14] |
| LDCM0268 | AC148 | PaTu 8988t | C109(0.49); C193(0.52) | LDD1147 | [14] |
| LDCM0269 | AC149 | PaTu 8988t | C109(0.50); C193(0.49) | LDD1148 | [14] |
| LDCM0270 | AC15 | PaTu 8988t | C109(0.62); C193(0.65) | LDD1149 | [14] |
| LDCM0271 | AC150 | PaTu 8988t | C109(0.60); C193(0.82) | LDD1150 | [14] |
| LDCM0272 | AC151 | PaTu 8988t | C109(0.60); C193(0.64) | LDD1151 | [14] |
| LDCM0273 | AC152 | PaTu 8988t | C109(0.50); C193(0.55) | LDD1152 | [14] |
| LDCM0274 | AC153 | PaTu 8988t | C109(0.56); C193(0.47) | LDD1153 | [14] |
| LDCM0621 | AC154 | PaTu 8988t | C109(0.50); C193(0.53) | LDD2166 | [14] |
| LDCM0622 | AC155 | PaTu 8988t | C109(0.53); C193(0.55) | LDD2167 | [14] |
| LDCM0623 | AC156 | PaTu 8988t | C109(0.35); C193(0.56) | LDD2168 | [14] |
| LDCM0624 | AC157 | PaTu 8988t | C109(0.53); C193(0.68) | LDD2169 | [14] |
| LDCM0279 | AC2 | PaTu 8988t | C109(0.24) | LDD1158 | [14] |
| LDCM0285 | AC25 | PaTu 8988t | C109(0.96); C193(0.98) | LDD1164 | [14] |
| LDCM0286 | AC26 | PaTu 8988t | C109(0.66); C193(0.74) | LDD1165 | [14] |
| LDCM0287 | AC27 | PaTu 8988t | C109(0.62); C193(0.70) | LDD1166 | [14] |
| LDCM0288 | AC28 | PaTu 8988t | C109(0.67); C193(0.60) | LDD1167 | [14] |
| LDCM0289 | AC29 | PaTu 8988t | C109(0.60); C193(0.60) | LDD1168 | [14] |
| LDCM0290 | AC3 | PaTu 8988t | C109(0.23) | LDD1169 | [14] |
| LDCM0291 | AC30 | PaTu 8988t | C109(1.24); C193(1.03) | LDD1170 | [14] |
| LDCM0292 | AC31 | PaTu 8988t | C109(1.40); C193(0.99) | LDD1171 | [14] |
| LDCM0293 | AC32 | PaTu 8988t | C109(1.19); C193(1.00) | LDD1172 | [14] |
| LDCM0294 | AC33 | PaTu 8988t | C109(0.75); C193(0.67) | LDD1173 | [14] |
| LDCM0295 | AC34 | PaTu 8988t | C109(0.95); C193(1.06) | LDD1174 | [14] |
| LDCM0296 | AC35 | PaTu 8988t | C109(1.24); C193(0.55) | LDD1175 | [14] |
| LDCM0297 | AC36 | PaTu 8988t | C109(1.02); C193(0.45) | LDD1176 | [14] |
| LDCM0298 | AC37 | PaTu 8988t | C109(0.68); C193(0.76) | LDD1177 | [14] |
| LDCM0299 | AC38 | PaTu 8988t | C109(0.74); C193(0.63) | LDD1178 | [14] |
| LDCM0300 | AC39 | PaTu 8988t | C109(0.84); C193(0.56) | LDD1179 | [14] |
| LDCM0301 | AC4 | PaTu 8988t | C109(0.26) | LDD1180 | [14] |
| LDCM0302 | AC40 | PaTu 8988t | C109(0.73); C193(0.60) | LDD1181 | [14] |
| LDCM0303 | AC41 | PaTu 8988t | C109(0.79); C193(0.74) | LDD1182 | [14] |
| LDCM0304 | AC42 | PaTu 8988t | C109(0.55); C193(0.73) | LDD1183 | [14] |
| LDCM0305 | AC43 | PaTu 8988t | C109(0.98); C193(0.76) | LDD1184 | [14] |
| LDCM0306 | AC44 | PaTu 8988t | C109(0.59); C193(0.67) | LDD1185 | [14] |
| LDCM0307 | AC45 | PaTu 8988t | C109(0.51); C193(0.83) | LDD1186 | [14] |
| LDCM0308 | AC46 | PaTu 8988t | C109(1.00) | LDD1187 | [14] |
| LDCM0309 | AC47 | PaTu 8988t | C109(0.84) | LDD1188 | [14] |
| LDCM0310 | AC48 | PaTu 8988t | C109(0.74) | LDD1189 | [14] |
| LDCM0311 | AC49 | PaTu 8988t | C109(0.98) | LDD1190 | [14] |
| LDCM0312 | AC5 | PaTu 8988t | C109(0.26) | LDD1191 | [14] |
| LDCM0313 | AC50 | PaTu 8988t | C109(0.80) | LDD1192 | [14] |
| LDCM0314 | AC51 | PaTu 8988t | C109(0.93) | LDD1193 | [14] |
| LDCM0315 | AC52 | PaTu 8988t | C109(0.94) | LDD1194 | [14] |
| LDCM0316 | AC53 | PaTu 8988t | C109(0.82) | LDD1195 | [14] |
| LDCM0317 | AC54 | PaTu 8988t | C109(0.89) | LDD1196 | [14] |
| LDCM0318 | AC55 | PaTu 8988t | C109(0.96) | LDD1197 | [14] |
| LDCM0319 | AC56 | PaTu 8988t | C109(0.86) | LDD1198 | [14] |
| LDCM0320 | AC57 | PaTu 8988t | C109(1.04); C193(1.02) | LDD1199 | [14] |
| LDCM0321 | AC58 | PaTu 8988t | C109(1.07); C193(0.94) | LDD1200 | [14] |
| LDCM0322 | AC59 | PaTu 8988t | C109(1.29); C193(1.14) | LDD1201 | [14] |
| LDCM0323 | AC6 | PaTu 8988t | C109(0.78); C193(0.58) | LDD1202 | [14] |
| LDCM0324 | AC60 | PaTu 8988t | C109(1.16); C193(1.00) | LDD1203 | [14] |
| LDCM0325 | AC61 | PaTu 8988t | C109(1.20); C193(0.98) | LDD1204 | [14] |
| LDCM0326 | AC62 | PaTu 8988t | C109(1.16); C193(0.98) | LDD1205 | [14] |
| LDCM0327 | AC63 | PaTu 8988t | C109(1.06); C193(0.88) | LDD1206 | [14] |
| LDCM0328 | AC64 | PaTu 8988t | C109(1.01); C193(1.04) | LDD1207 | [14] |
| LDCM0329 | AC65 | PaTu 8988t | C109(1.32); C193(1.12) | LDD1208 | [14] |
| LDCM0330 | AC66 | PaTu 8988t | C109(1.26); C193(0.95) | LDD1209 | [14] |
| LDCM0331 | AC67 | PaTu 8988t | C109(1.08); C193(1.04) | LDD1210 | [14] |
| LDCM0332 | AC68 | PaTu 8988t | C109(1.16); C193(1.22) | LDD1211 | [14] |
| LDCM0333 | AC69 | PaTu 8988t | C109(0.78); C193(1.05) | LDD1212 | [14] |
| LDCM0334 | AC7 | PaTu 8988t | C109(1.10); C193(0.77) | LDD1213 | [14] |
| LDCM0335 | AC70 | PaTu 8988t | C109(1.13); C193(0.98) | LDD1214 | [14] |
| LDCM0336 | AC71 | PaTu 8988t | C109(1.10); C193(0.83) | LDD1215 | [14] |
| LDCM0337 | AC72 | PaTu 8988t | C109(0.90); C193(1.31) | LDD1216 | [14] |
| LDCM0338 | AC73 | PaTu 8988t | C109(0.83); C193(1.23) | LDD1217 | [14] |
| LDCM0339 | AC74 | PaTu 8988t | C109(0.75); C193(1.08) | LDD1218 | [14] |
| LDCM0340 | AC75 | PaTu 8988t | C109(0.77); C193(0.80) | LDD1219 | [14] |
| LDCM0341 | AC76 | PaTu 8988t | C109(1.00); C193(1.04) | LDD1220 | [14] |
| LDCM0342 | AC77 | PaTu 8988t | C109(0.79); C193(0.78) | LDD1221 | [14] |
| LDCM0343 | AC78 | PaTu 8988t | C109(1.10); C193(1.12) | LDD1222 | [14] |
| LDCM0344 | AC79 | PaTu 8988t | C109(0.84); C193(0.70) | LDD1223 | [14] |
| LDCM0345 | AC8 | PaTu 8988t | C109(0.71); C193(0.74) | LDD1224 | [14] |
| LDCM0346 | AC80 | PaTu 8988t | C109(1.03); C193(0.93) | LDD1225 | [14] |
| LDCM0347 | AC81 | PaTu 8988t | C109(1.02); C193(1.34) | LDD1226 | [14] |
| LDCM0348 | AC82 | PaTu 8988t | C109(0.65); C193(1.10) | LDD1227 | [14] |
| LDCM0248 | AKOS034007472 | PaTu 8988t | C109(0.63); C193(0.73) | LDD1127 | [14] |
| LDCM0356 | AKOS034007680 | PaTu 8988t | C109(0.66); C193(0.59) | LDD1235 | [14] |
| LDCM0275 | AKOS034007705 | PaTu 8988t | C109(0.56); C193(0.51) | LDD1154 | [14] |
| LDCM0367 | CL1 | PaTu 8988t | C193(0.64) | LDD1246 | [14] |
| LDCM0368 | CL10 | PaTu 8988t | C193(0.42) | LDD1247 | [14] |
| LDCM0369 | CL100 | PaTu 8988t | C109(0.23) | LDD1248 | [14] |
| LDCM0370 | CL101 | PaTu 8988t | C109(0.86); C193(0.90) | LDD1249 | [14] |
| LDCM0371 | CL102 | PaTu 8988t | C109(0.84); C193(0.56) | LDD1250 | [14] |
| LDCM0372 | CL103 | PaTu 8988t | C109(0.67); C193(0.54) | LDD1251 | [14] |
| LDCM0373 | CL104 | PaTu 8988t | C109(0.78); C193(0.62) | LDD1252 | [14] |
| LDCM0379 | CL11 | PaTu 8988t | C193(0.55) | LDD1258 | [14] |
| LDCM0382 | CL112 | PaTu 8988t | C109(0.97); C193(0.84) | LDD1261 | [14] |
| LDCM0383 | CL113 | PaTu 8988t | C109(1.02); C193(0.97) | LDD1262 | [14] |
| LDCM0384 | CL114 | PaTu 8988t | C109(0.78); C193(0.84) | LDD1263 | [14] |
| LDCM0385 | CL115 | PaTu 8988t | C109(0.77); C193(0.99) | LDD1264 | [14] |
| LDCM0386 | CL116 | PaTu 8988t | C109(0.60); C193(0.77) | LDD1265 | [14] |
| LDCM0387 | CL117 | PaTu 8988t | C109(0.62); C193(0.83) | LDD1266 | [14] |
| LDCM0388 | CL118 | PaTu 8988t | C109(0.42); C193(1.03) | LDD1267 | [14] |
| LDCM0389 | CL119 | PaTu 8988t | C109(0.59); C193(0.67) | LDD1268 | [14] |
| LDCM0390 | CL12 | PaTu 8988t | C193(1.04) | LDD1269 | [14] |
| LDCM0391 | CL120 | PaTu 8988t | C109(1.03); C193(0.51) | LDD1270 | [14] |
| LDCM0392 | CL121 | PaTu 8988t | C109(0.78) | LDD1271 | [14] |
| LDCM0393 | CL122 | PaTu 8988t | C109(1.20) | LDD1272 | [14] |
| LDCM0394 | CL123 | PaTu 8988t | C109(0.82) | LDD1273 | [14] |
| LDCM0395 | CL124 | PaTu 8988t | C109(0.88) | LDD1274 | [14] |
| LDCM0396 | CL125 | PaTu 8988t | C109(1.21); C193(1.10) | LDD1275 | [14] |
| LDCM0397 | CL126 | PaTu 8988t | C109(1.23); C193(1.19) | LDD1276 | [14] |
| LDCM0398 | CL127 | PaTu 8988t | C109(1.20); C193(1.01) | LDD1277 | [14] |
| LDCM0399 | CL128 | PaTu 8988t | C109(1.00); C193(1.03) | LDD1278 | [14] |
| LDCM0400 | CL13 | PaTu 8988t | C193(0.53) | LDD1279 | [14] |
| LDCM0401 | CL14 | PaTu 8988t | C193(1.22) | LDD1280 | [14] |
| LDCM0402 | CL15 | PaTu 8988t | C193(0.56) | LDD1281 | [14] |
| LDCM0407 | CL2 | PaTu 8988t | C193(0.87) | LDD1286 | [14] |
| LDCM0418 | CL3 | PaTu 8988t | C193(0.51) | LDD1297 | [14] |
| LDCM0420 | CL31 | PaTu 8988t | C193(0.53) | LDD1299 | [14] |
| LDCM0421 | CL32 | PaTu 8988t | C193(0.33) | LDD1300 | [14] |
| LDCM0422 | CL33 | PaTu 8988t | C193(0.46) | LDD1301 | [14] |
| LDCM0423 | CL34 | PaTu 8988t | C193(0.33) | LDD1302 | [14] |
| LDCM0424 | CL35 | PaTu 8988t | C193(0.48) | LDD1303 | [14] |
| LDCM0425 | CL36 | PaTu 8988t | C193(0.53) | LDD1304 | [14] |
| LDCM0426 | CL37 | PaTu 8988t | C193(0.24) | LDD1305 | [14] |
| LDCM0428 | CL39 | PaTu 8988t | C193(0.48) | LDD1307 | [14] |
| LDCM0429 | CL4 | PaTu 8988t | C193(0.72) | LDD1308 | [14] |
| LDCM0430 | CL40 | PaTu 8988t | C193(0.96) | LDD1309 | [14] |
| LDCM0431 | CL41 | PaTu 8988t | C193(0.33) | LDD1310 | [14] |
| LDCM0432 | CL42 | PaTu 8988t | C193(0.48) | LDD1311 | [14] |
| LDCM0433 | CL43 | PaTu 8988t | C193(0.33) | LDD1312 | [14] |
| LDCM0434 | CL44 | PaTu 8988t | C193(0.42) | LDD1313 | [14] |
| LDCM0435 | CL45 | PaTu 8988t | C193(0.64) | LDD1314 | [14] |
| LDCM0436 | CL46 | PaTu 8988t | C109(1.11) | LDD1315 | [14] |
| LDCM0437 | CL47 | PaTu 8988t | C109(0.86) | LDD1316 | [14] |
| LDCM0438 | CL48 | PaTu 8988t | C109(0.84) | LDD1317 | [14] |
| LDCM0439 | CL49 | PaTu 8988t | C109(0.92) | LDD1318 | [14] |
| LDCM0440 | CL5 | PaTu 8988t | C193(0.64) | LDD1319 | [14] |
| LDCM0441 | CL50 | PaTu 8988t | C109(0.83) | LDD1320 | [14] |
| LDCM0442 | CL51 | PaTu 8988t | C109(0.73) | LDD1321 | [14] |
| LDCM0443 | CL52 | PaTu 8988t | C109(0.94) | LDD1322 | [14] |
| LDCM0444 | CL53 | PaTu 8988t | C109(0.60) | LDD1323 | [14] |
| LDCM0445 | CL54 | PaTu 8988t | C109(1.00) | LDD1324 | [14] |
| LDCM0446 | CL55 | PaTu 8988t | C109(0.97) | LDD1325 | [14] |
| LDCM0447 | CL56 | PaTu 8988t | C109(0.98) | LDD1326 | [14] |
| LDCM0448 | CL57 | PaTu 8988t | C109(0.82) | LDD1327 | [14] |
| LDCM0449 | CL58 | PaTu 8988t | C109(0.80) | LDD1328 | [14] |
| LDCM0450 | CL59 | PaTu 8988t | C109(1.16) | LDD1329 | [14] |
| LDCM0451 | CL6 | PaTu 8988t | C193(0.93) | LDD1330 | [14] |
| LDCM0452 | CL60 | PaTu 8988t | C109(1.09) | LDD1331 | [14] |
| LDCM0453 | CL61 | PaTu 8988t | C193(1.51) | LDD1332 | [14] |
| LDCM0454 | CL62 | PaTu 8988t | C193(1.60) | LDD1333 | [14] |
| LDCM0455 | CL63 | PaTu 8988t | C193(0.82) | LDD1334 | [14] |
| LDCM0456 | CL64 | PaTu 8988t | C193(0.70) | LDD1335 | [14] |
| LDCM0457 | CL65 | PaTu 8988t | C193(0.96) | LDD1336 | [14] |
| LDCM0458 | CL66 | PaTu 8988t | C193(0.59) | LDD1337 | [14] |
| LDCM0459 | CL67 | PaTu 8988t | C193(0.78) | LDD1338 | [14] |
| LDCM0460 | CL68 | PaTu 8988t | C193(1.02) | LDD1339 | [14] |
| LDCM0461 | CL69 | PaTu 8988t | C193(0.76) | LDD1340 | [14] |
| LDCM0462 | CL7 | PaTu 8988t | C193(0.63) | LDD1341 | [14] |
| LDCM0463 | CL70 | PaTu 8988t | C193(0.68) | LDD1342 | [14] |
| LDCM0464 | CL71 | PaTu 8988t | C193(1.09) | LDD1343 | [14] |
| LDCM0465 | CL72 | PaTu 8988t | C193(0.73) | LDD1344 | [14] |
| LDCM0466 | CL73 | PaTu 8988t | C193(0.80) | LDD1345 | [14] |
| LDCM0467 | CL74 | PaTu 8988t | C193(0.86) | LDD1346 | [14] |
| LDCM0469 | CL76 | PaTu 8988t | C193(0.70) | LDD1348 | [14] |
| LDCM0470 | CL77 | PaTu 8988t | C193(0.63) | LDD1349 | [14] |
| LDCM0471 | CL78 | PaTu 8988t | C193(0.48) | LDD1350 | [14] |
| LDCM0472 | CL79 | PaTu 8988t | C193(0.51) | LDD1351 | [14] |
| LDCM0473 | CL8 | PaTu 8988t | C193(0.37) | LDD1352 | [14] |
| LDCM0474 | CL80 | PaTu 8988t | C193(0.44) | LDD1353 | [14] |
| LDCM0475 | CL81 | PaTu 8988t | C193(0.48) | LDD1354 | [14] |
| LDCM0476 | CL82 | PaTu 8988t | C193(0.23) | LDD1355 | [14] |
| LDCM0477 | CL83 | PaTu 8988t | C193(0.31) | LDD1356 | [14] |
| LDCM0478 | CL84 | PaTu 8988t | C193(0.90) | LDD1357 | [14] |
| LDCM0479 | CL85 | PaTu 8988t | C193(0.84) | LDD1358 | [14] |
| LDCM0480 | CL86 | PaTu 8988t | C193(0.54) | LDD1359 | [14] |
| LDCM0481 | CL87 | PaTu 8988t | C193(0.70) | LDD1360 | [14] |
| LDCM0482 | CL88 | PaTu 8988t | C193(0.60) | LDD1361 | [14] |
| LDCM0483 | CL89 | PaTu 8988t | C193(1.10) | LDD1362 | [14] |
| LDCM0484 | CL9 | PaTu 8988t | C193(0.82) | LDD1363 | [14] |
| LDCM0485 | CL90 | PaTu 8988t | C193(0.62) | LDD1364 | [14] |
| LDCM0486 | CL91 | PaTu 8988t | C109(0.27) | LDD1365 | [14] |
| LDCM0487 | CL92 | PaTu 8988t | C109(0.39) | LDD1366 | [14] |
| LDCM0488 | CL93 | PaTu 8988t | C109(0.21) | LDD1367 | [14] |
| LDCM0489 | CL94 | PaTu 8988t | C109(0.49) | LDD1368 | [14] |
| LDCM0490 | CL95 | PaTu 8988t | C109(0.22) | LDD1369 | [14] |
| LDCM0491 | CL96 | PaTu 8988t | C109(0.31) | LDD1370 | [14] |
| LDCM0492 | CL97 | PaTu 8988t | C109(0.25) | LDD1371 | [14] |
| LDCM0493 | CL98 | PaTu 8988t | C109(0.29) | LDD1372 | [14] |
| LDCM0494 | CL99 | PaTu 8988t | C109(0.56) | LDD1373 | [14] |
| LDCM0175 | Ethacrynic acid | HeLa | N.A. | LDD0440 | [4] |
| LDCM0468 | Fragment33 | PaTu 8988t | C193(0.76) | LDD1347 | [14] |
| LDCM0427 | Fragment51 | PaTu 8988t | C193(0.51) | LDD1306 | [14] |
| LDCM0015 | HNE | MDA-MB-231 | C234(1.03); C189(0.91) | LDD0346 | [15] |
| LDCM0022 | KB02 | 22RV1 | C190(2.13); C234(1.81) | LDD2243 | [3] |
| LDCM0023 | KB03 | 22RV1 | C190(3.75); C234(3.12) | LDD2660 | [3] |
| LDCM0024 | KB05 | COLO792 | C234(2.11) | LDD3310 | [3] |
| LDCM0109 | NEM | HeLa | H26(0.00); H293(0.00) | LDD0223 | [5] |
| LDCM0019 | Staurosporine | Hep-G2 | N.A. | LDD0083 | [13] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Other
References
