Details of the Target
General Information of Target
| Target ID | LDTP05977 | |||||
|---|---|---|---|---|---|---|
| Target Name | Nascent polypeptide-associated complex subunit alpha (NACA) | |||||
| Gene Name | NACA | |||||
| Gene ID | 4666 | |||||
| Synonyms |
Nascent polypeptide-associated complex subunit alpha; NAC-alpha; Alpha-NAC; allergen Hom s 2 |
|||||
| 3D Structure | ||||||
| Sequence |
MPGEATETVPATEQELPQPQAETGSGTESDSDESVPELEEQDSTQATTQQAQLAAAAEID
EEPVSKAKQSRSEKKARKAMSKLGLRQVTGVTRVTIRKSKNILFVITKPDVYKSPASDTY IVFGEAKIEDLSQQAQLAAAEKFKVQGEAVSNIQENTQTPTVQEESEEEEVDETGVEVKD IELVMSQANVSRAKAVRALKNNSNDIVNAIMELTM |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
NAC-alpha family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Prevents inappropriate targeting of non-secretory polypeptides to the endoplasmic reticulum (ER). Binds to nascent polypeptide chains as they emerge from the ribosome and blocks their interaction with the signal recognition particle (SRP), which normally targets nascent secretory peptides to the ER. Also reduces the inherent affinity of ribosomes for protein translocation sites in the ER membrane (M sites). May act as a specific coactivator for JUN, binding to DNA and stabilizing the interaction of JUN homodimers with target gene promoters.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
AZ-9 Probe Info |
 |
8.05 | LDD0393 | [1] | |
|
C-Sul Probe Info |
 |
8.27 | LDD0066 | [2] | |
|
TH211 Probe Info |
 |
Y112(20.00) | LDD0257 | [3] | |
|
TH216 Probe Info |
 |
Y112(20.00) | LDD0259 | [3] | |
|
OPA-S-S-alkyne Probe Info |
 |
K82(3.23) | LDD3494 | [4] | |
|
Probe 1 Probe Info |
 |
Y120(33.59) | LDD3495 | [5] | |
|
P11 Probe Info |
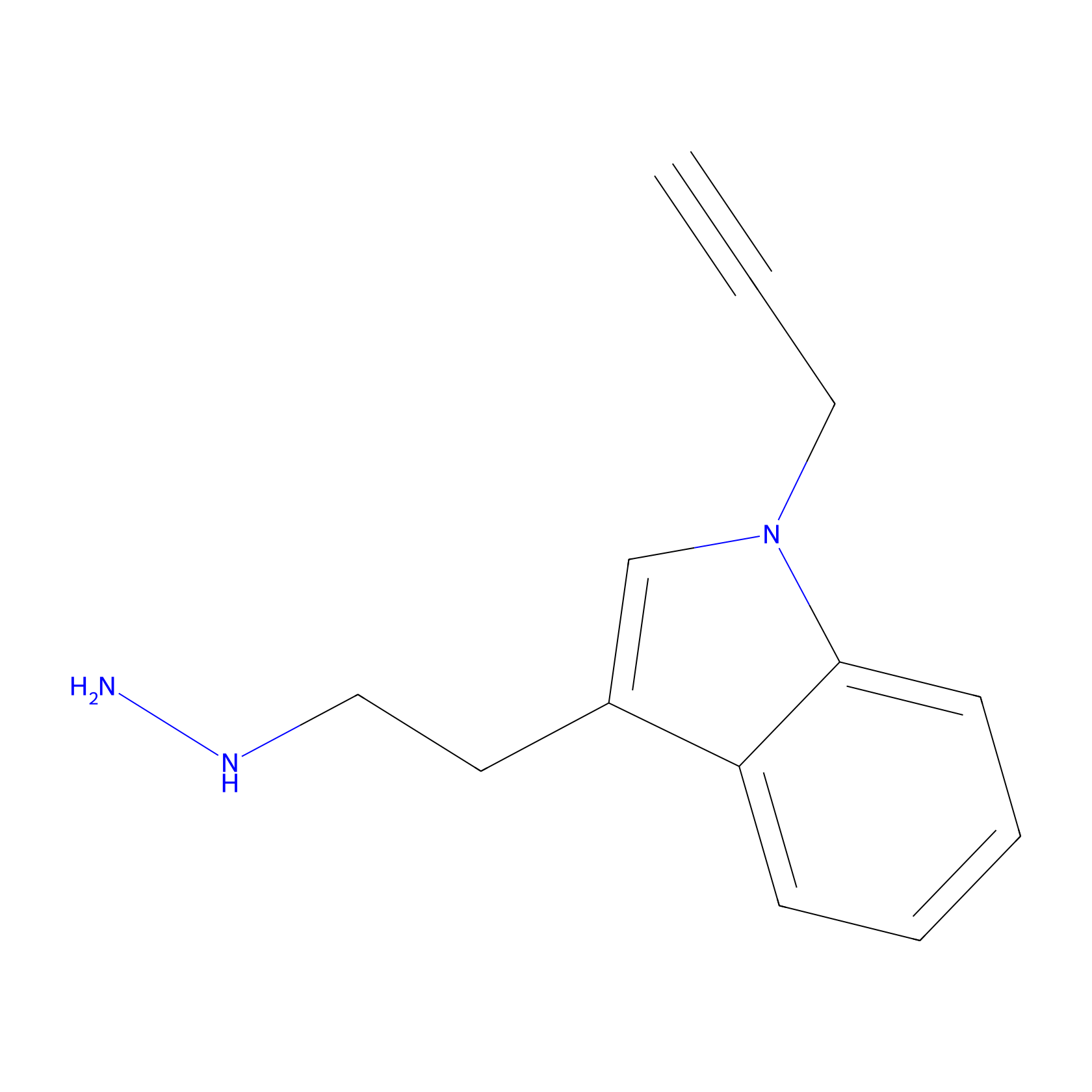 |
5.69 | LDD0201 | [6] | |
|
HHS-482 Probe Info |
 |
Y120(0.29) | LDD0285 | [7] | |
|
HHS-475 Probe Info |
 |
Y112(0.84); Y120(0.94) | LDD0264 | [8] | |
|
HHS-465 Probe Info |
 |
Y120(10.00) | LDD2237 | [9] | |
|
ATP probe Probe Info |
 |
K142(0.00); K144(0.00); K82(0.00); K100(0.00) | LDD0199 | [10] | |
|
NHS Probe Info |
 |
K108(0.00); K142(0.00); K100(0.00) | LDD0010 | [11] | |
|
SF Probe Info |
 |
Y112(0.00); K100(0.00); Y120(0.00); K113(0.00) | LDD0028 | [12] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [11] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe3 Probe Info |
 |
13.09 | LDD0464 | [13] | |
|
PARPYnD Probe Info |
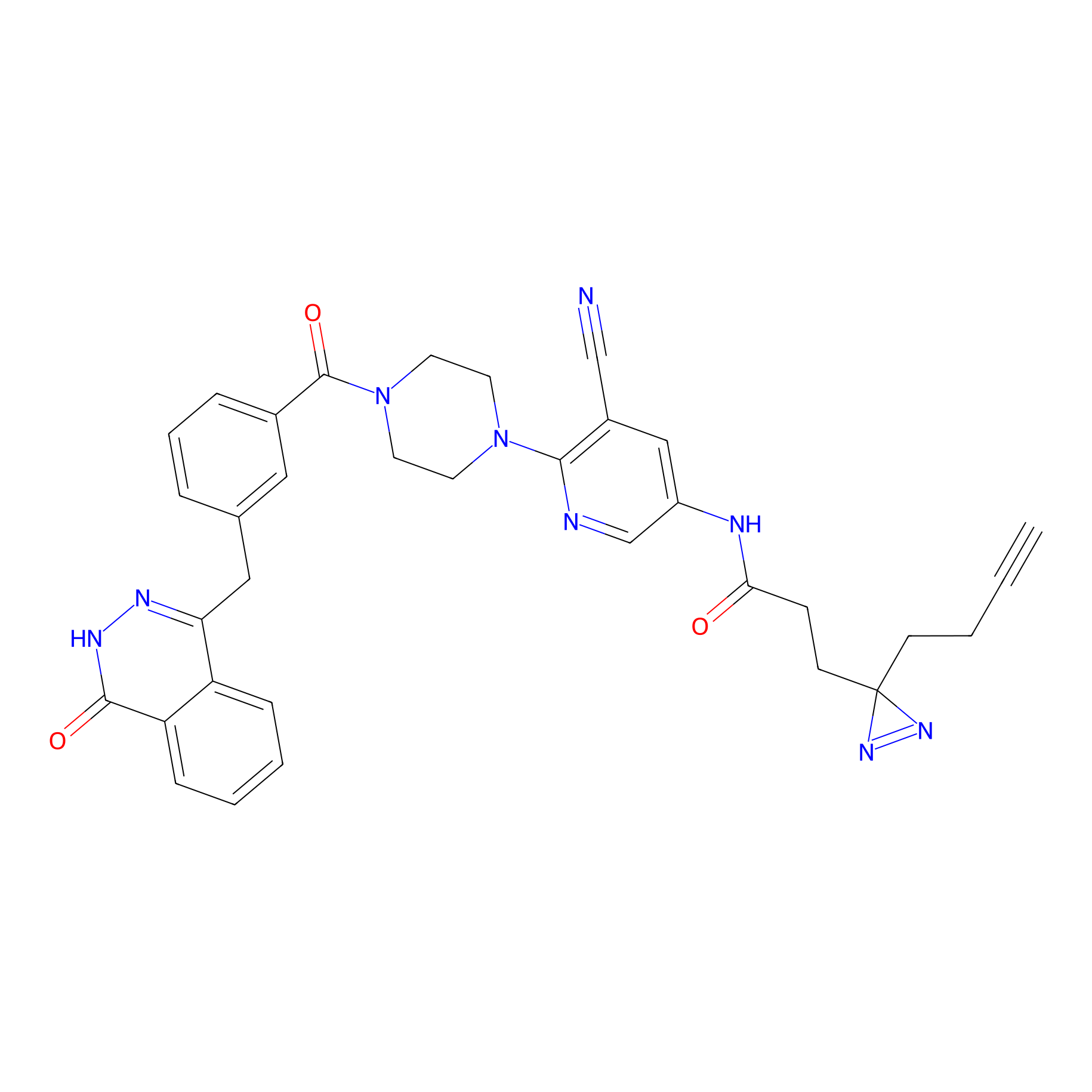 |
0.56 | LDD0376 | [14] | |
|
Diazir Probe Info |
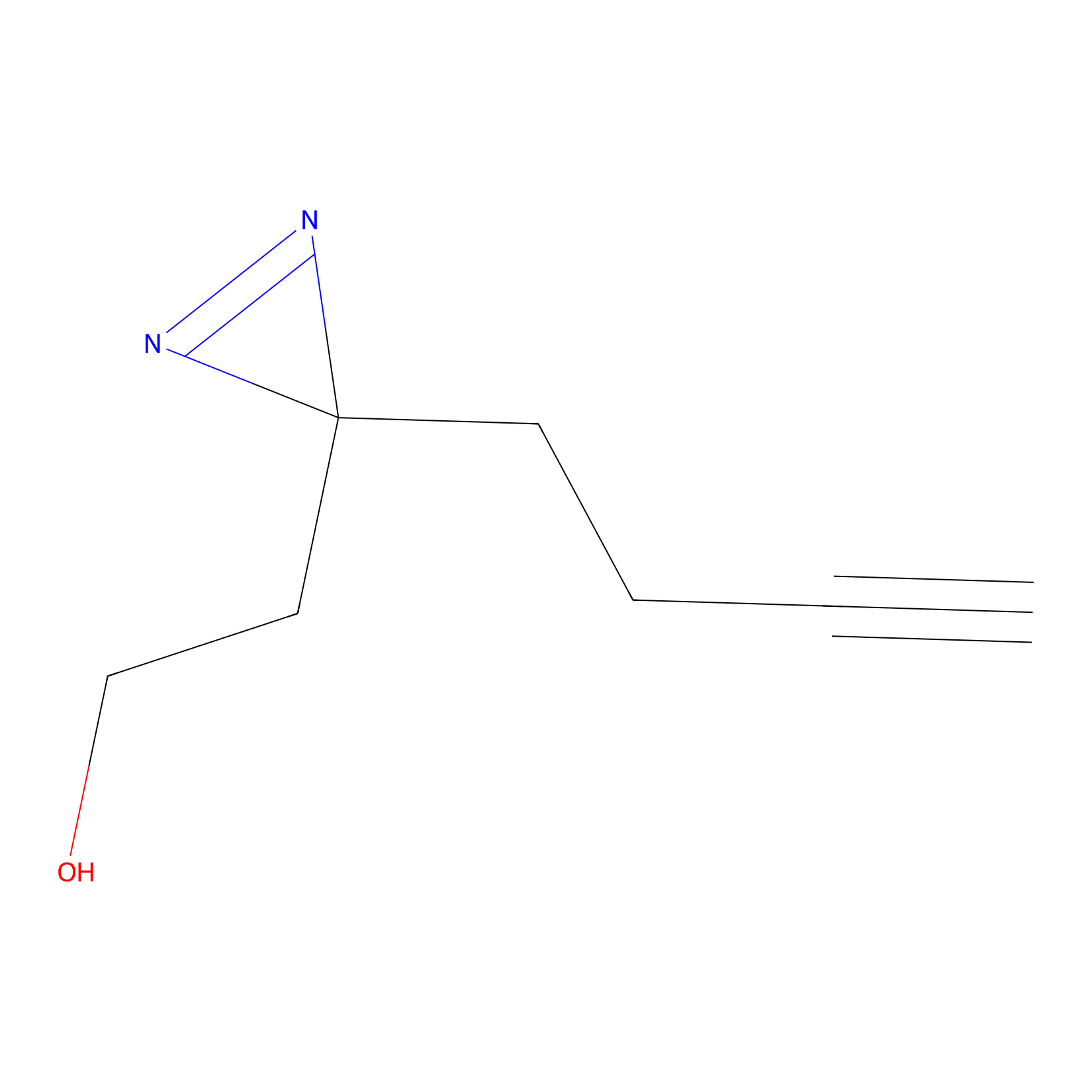 |
N.A. | LDD0011 | [11] | |
|
LD-F Probe Info |
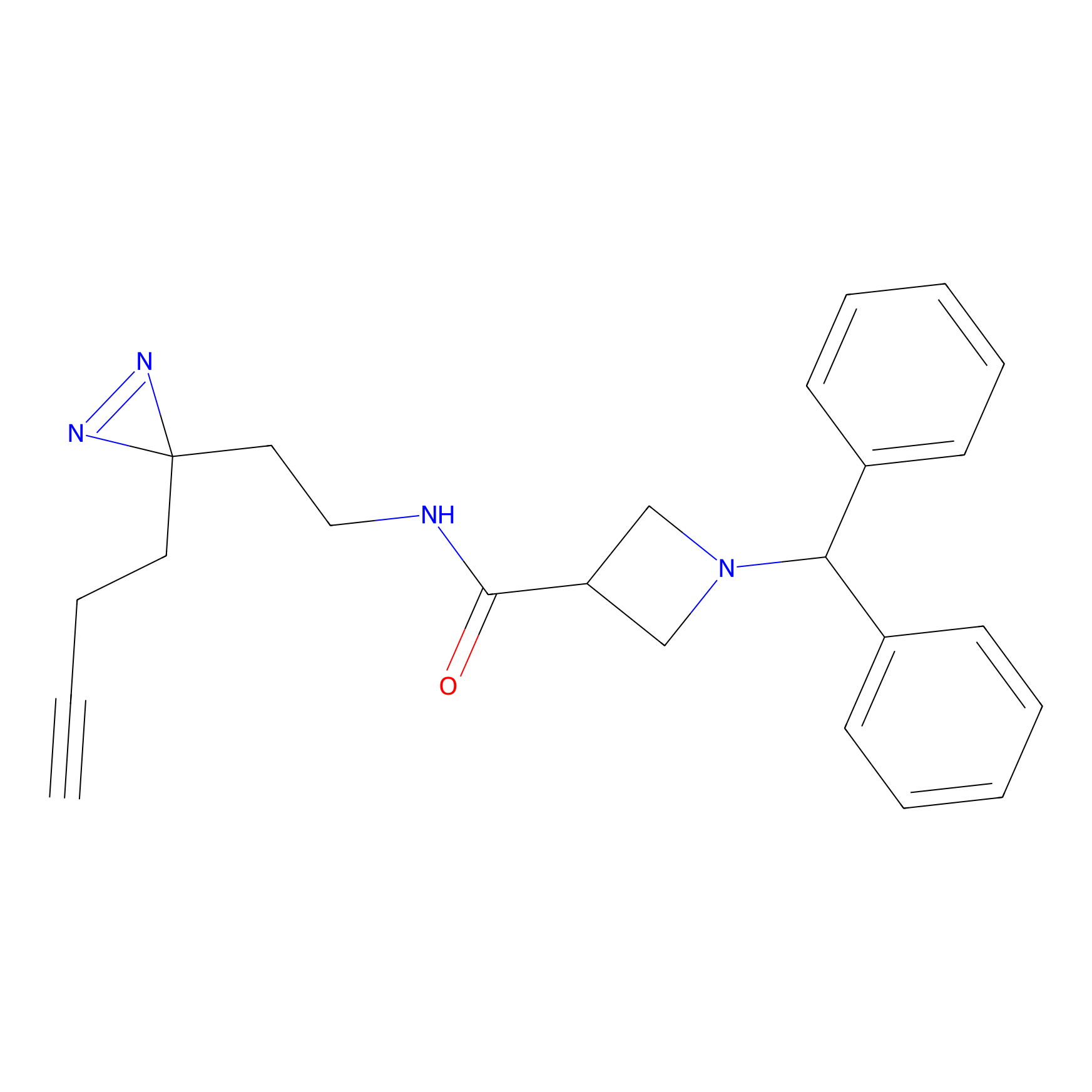 |
S132(0.00); D130(0.00); L131(0.00) | LDD0015 | [15] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0073 | [16] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0147 | AZ0108 | MDA-MB-468 | 0.56 | LDD0376 | [14] |
| LDCM0088 | C45 | HEK-293T | 5.69 | LDD0201 | [6] |
| LDCM0116 | HHS-0101 | DM93 | Y112(0.84); Y120(0.94) | LDD0264 | [8] |
| LDCM0117 | HHS-0201 | DM93 | Y112(0.71); Y120(1.06) | LDD0265 | [8] |
| LDCM0118 | HHS-0301 | DM93 | Y112(0.90); Y120(1.32) | LDD0266 | [8] |
| LDCM0119 | HHS-0401 | DM93 | Y112(0.84); Y120(1.29) | LDD0267 | [8] |
| LDCM0120 | HHS-0701 | DM93 | Y112(0.74); Y120(0.98) | LDD0268 | [8] |
| LDCM0123 | JWB131 | DM93 | Y120(0.29) | LDD0285 | [7] |
| LDCM0124 | JWB142 | DM93 | Y120(0.86) | LDD0286 | [7] |
| LDCM0125 | JWB146 | DM93 | Y112(0.09); Y120(0.85) | LDD0287 | [7] |
| LDCM0126 | JWB150 | DM93 | Y112(0.14); Y120(2.36) | LDD0288 | [7] |
| LDCM0127 | JWB152 | DM93 | Y112(0.26); Y120(1.10) | LDD0289 | [7] |
| LDCM0128 | JWB198 | DM93 | Y112(0.07); Y120(0.34) | LDD0290 | [7] |
| LDCM0129 | JWB202 | DM93 | Y112(0.09) | LDD0291 | [7] |
| LDCM0130 | JWB211 | DM93 | Y112(0.18) | LDD0292 | [7] |
The Interaction Atlas With This Target
References
