Details of the Target
General Information of Target
| Target ID | LDTP05746 | |||||
|---|---|---|---|---|---|---|
| Target Name | COP9 signalosome complex subunit 1 (GPS1) | |||||
| Gene Name | GPS1 | |||||
| Gene ID | 2873 | |||||
| Synonyms |
COPS1; CSN1; COP9 signalosome complex subunit 1; SGN1; Signalosome subunit 1; G protein pathway suppressor 1; GPS-1; JAB1-containing signalosome subunit 1; Protein MFH |
|||||
| 3D Structure | ||||||
| Sequence |
MPLPVQVFNLQGAVEPMQIDVDPQEDPQNAPDVNYVVENPSLDLEQYAASYSGLMRIERL
QFIADHCPTLRVEALKMALSFVQRTFNVDMYEEIHRKLSEATRSSLRELQNAPDAIPESG VEPPALDTAWVEATRKKALLKLEKLDTDLKNYKGNSIKESIRRGHDDLGDHYLDCGDLSN ALKCYSRARDYCTSAKHVINMCLNVIKVSVYLQNWSHVLSYVSKAESTPEIAEQRGERDS QTQAILTKLKCAAGLAELAARKYKQAAKCLLLASFDHCDFPELLSPSNVAIYGGLCALAT FDRQELQRNVISSSSFKLFLELEPQVRDIIFKFYESKYASCLKMLDEMKDNLLLDMYLAP HVRTLYTQIRNRALIQYFSPYVSADMHRMAAAFNTTVAALEDELTQLILEGLISARVDSH SKILYARDVDQRSTTFEKSLLMGKEFQRRAKAMMLRAAVLRNQIHVKSPPREGSQGELTP ANSQSRMSTNM |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
CSN1 family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Essential component of the COP9 signalosome complex (CSN), a complex involved in various cellular and developmental processes. The CSN complex is an essential regulator of the ubiquitin (Ubl) conjugation pathway by mediating the deneddylation of the cullin subunits of SCF-type E3 ligase complexes, leading to decrease the Ubl ligase activity of SCF-type complexes such as SCF, CSA or DDB2. The complex is also involved in phosphorylation of p53/TP53, c-jun/JUN, IkappaBalpha/NFKBIA, ITPK1 and IRF8/ICSBP, possibly via its association with CK2 and PKD kinases. CSN-dependent phosphorylation of TP53 and JUN promotes and protects degradation by the Ubl system, respectively. Suppresses G-protein- and mitogen-activated protein kinase-mediated signal transduction.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
7.30 | LDD0402 | [1] | |
|
DBIA Probe Info |
 |
C228(5.27); C238(1.87); C67(2.33) | LDD3311 | [2] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [3] | |
|
BTD Probe Info |
 |
C341(2.34) | LDD1700 | [4] | |
|
AHL-Pu-1 Probe Info |
 |
C251(6.56); C67(2.07) | LDD0169 | [5] | |
|
HHS-475 Probe Info |
 |
Y381(0.69) | LDD0264 | [6] | |
|
HHS-465 Probe Info |
 |
Y381(2.67) | LDD2237 | [7] | |
|
ATP probe Probe Info |
 |
K337(0.00); K183(0.00) | LDD0199 | [8] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0358 | [9] | |
|
CY-1 Probe Info |
 |
K150(0.00); Y152(0.00) | LDD0246 | [10] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C192(0.00); C251(0.00); C175(0.00); C202(0.00) | LDD0038 | [11] | |
|
IA-alkyne Probe Info |
 |
C67(0.00); C251(0.00); C175(0.00) | LDD0032 | [12] | |
|
Lodoacetamide azide Probe Info |
 |
C251(0.00); C192(0.00); C175(0.00); C202(0.00) | LDD0037 | [11] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [13] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [14] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [15] | |
|
IPM Probe Info |
 |
C251(0.00); C67(0.00) | LDD0147 | [13] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [14] | |
|
NPM Probe Info |
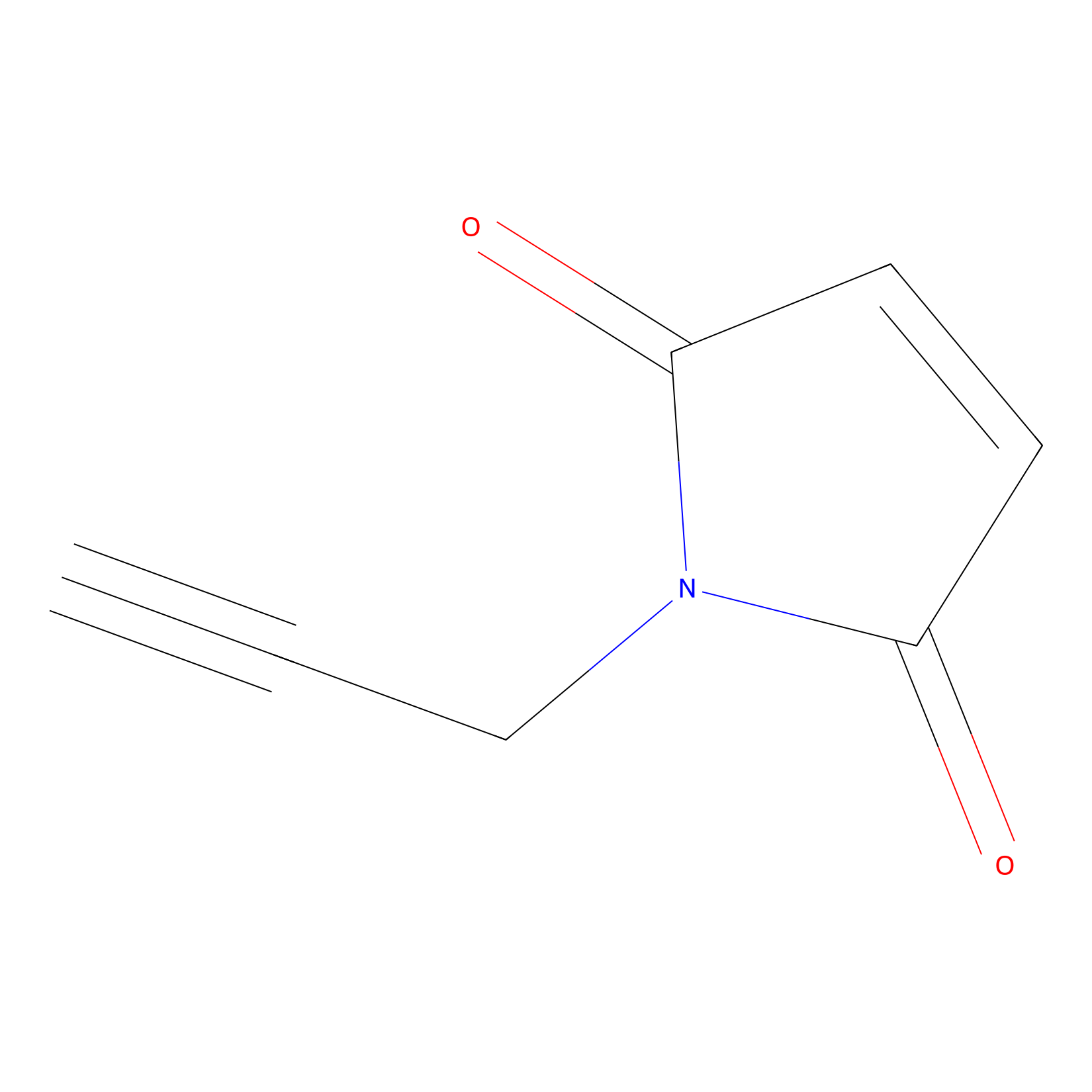 |
N.A. | LDD0016 | [14] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [14] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [14] | |
|
1c-yne Probe Info |
 |
K76(0.00); K349(0.00); K224(0.00) | LDD0228 | [9] | |
|
Acrolein Probe Info |
 |
C175(0.00); C67(0.00); H95(0.00); C192(0.00) | LDD0217 | [16] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [16] | |
|
Methacrolein Probe Info |
 |
C67(0.00); C341(0.00) | LDD0218 | [16] | |
|
W1 Probe Info |
 |
C175(0.00); C67(0.00) | LDD0236 | [17] | |
|
NAIA_5 Probe Info |
 |
C192(0.00); C251(0.00); C341(0.00); C175(0.00) | LDD2223 | [18] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C341(0.56); C251(0.51) | LDD2142 | [4] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C341(0.72) | LDD2112 | [4] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C192(0.94); C341(1.13); C251(1.36); C67(0.70) | LDD2117 | [4] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C341(0.95); C251(0.88) | LDD2152 | [4] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C341(1.54); C251(0.79) | LDD2103 | [4] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C341(0.53) | LDD2131 | [4] |
| LDCM0025 | 4SU-RNA | DM93 | C67(3.31) | LDD0170 | [5] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C251(6.56); C67(2.07) | LDD0169 | [5] |
| LDCM0214 | AC1 | HEK-293T | C192(0.92); C202(1.02); C175(0.96); C67(0.86) | LDD1507 | [19] |
| LDCM0215 | AC10 | HEK-293T | C192(1.05); C202(1.01); C175(1.07); C67(0.92) | LDD1508 | [19] |
| LDCM0226 | AC11 | HEK-293T | C192(0.91); C202(0.94); C175(1.03); C67(0.97) | LDD1509 | [19] |
| LDCM0237 | AC12 | HEK-293T | C192(0.94); C202(0.96); C175(1.12); C67(0.93) | LDD1510 | [19] |
| LDCM0259 | AC14 | HEK-293T | C192(1.06); C175(1.03); C67(1.08) | LDD1512 | [19] |
| LDCM0270 | AC15 | HEK-293T | C192(1.09); C202(1.14); C175(1.17); C67(0.94) | LDD1513 | [19] |
| LDCM0276 | AC17 | HEK-293T | C192(1.07); C202(0.95); C175(1.07); C67(0.92) | LDD1515 | [19] |
| LDCM0277 | AC18 | HEK-293T | C192(1.06); C202(1.00); C175(1.02); C67(1.06) | LDD1516 | [19] |
| LDCM0278 | AC19 | HEK-293T | C192(0.90); C202(0.87); C175(1.15); C67(1.03) | LDD1517 | [19] |
| LDCM0279 | AC2 | HEK-293T | C192(1.05); C202(1.03); C175(1.23); C67(1.01) | LDD1518 | [19] |
| LDCM0280 | AC20 | HEK-293T | C192(0.96); C202(0.95); C175(1.19); C67(0.98) | LDD1519 | [19] |
| LDCM0281 | AC21 | HEK-293T | C192(1.05); C202(1.03); C175(1.09); C67(0.98) | LDD1520 | [19] |
| LDCM0282 | AC22 | HEK-293T | C192(1.00); C175(1.07); C67(1.25) | LDD1521 | [19] |
| LDCM0283 | AC23 | HEK-293T | C192(1.04); C202(1.15); C175(1.12); C67(1.11) | LDD1522 | [19] |
| LDCM0284 | AC24 | HEK-293T | C192(1.04); C175(1.06); C67(1.03) | LDD1523 | [19] |
| LDCM0285 | AC25 | HEK-293T | C192(1.00); C202(0.94); C175(1.03); C67(0.83) | LDD1524 | [19] |
| LDCM0286 | AC26 | HEK-293T | C192(1.06); C202(1.00); C175(1.03); C67(0.92) | LDD1525 | [19] |
| LDCM0287 | AC27 | HEK-293T | C192(1.04); C202(0.94); C175(1.08); C67(1.12) | LDD1526 | [19] |
| LDCM0288 | AC28 | HEK-293T | C192(0.95); C202(0.99); C175(1.08); C67(1.01) | LDD1527 | [19] |
| LDCM0289 | AC29 | HEK-293T | C192(1.02); C202(1.08); C175(0.99); C67(0.87) | LDD1528 | [19] |
| LDCM0290 | AC3 | HEK-293T | C192(0.94); C202(1.01); C175(1.17); C67(0.89) | LDD1529 | [19] |
| LDCM0291 | AC30 | HEK-293T | C192(1.02); C175(1.06); C67(1.00) | LDD1530 | [19] |
| LDCM0292 | AC31 | HEK-293T | C192(1.06); C202(1.03); C175(1.07); C67(0.97) | LDD1531 | [19] |
| LDCM0293 | AC32 | HEK-293T | C192(1.02); C175(1.05); C67(0.98) | LDD1532 | [19] |
| LDCM0294 | AC33 | HEK-293T | C192(0.93); C202(0.96); C175(1.00); C67(0.88) | LDD1533 | [19] |
| LDCM0295 | AC34 | HEK-293T | C192(1.10); C202(0.99); C175(1.07); C67(0.95) | LDD1534 | [19] |
| LDCM0296 | AC35 | HEK-293T | C192(0.92); C202(0.90); C175(1.07); C67(1.12) | LDD1535 | [19] |
| LDCM0297 | AC36 | HEK-293T | C192(0.92); C202(0.92); C175(0.98); C67(0.88) | LDD1536 | [19] |
| LDCM0298 | AC37 | HEK-293T | C192(1.02); C202(1.05); C175(0.99); C67(0.90) | LDD1537 | [19] |
| LDCM0299 | AC38 | HEK-293T | C192(1.04); C175(1.13); C67(1.14) | LDD1538 | [19] |
| LDCM0300 | AC39 | HEK-293T | C192(1.12); C202(1.00); C175(1.05); C67(0.96) | LDD1539 | [19] |
| LDCM0301 | AC4 | HEK-293T | C192(0.94); C202(0.88); C175(1.11); C67(1.01) | LDD1540 | [19] |
| LDCM0302 | AC40 | HEK-293T | C192(1.12); C175(1.00); C67(0.96) | LDD1541 | [19] |
| LDCM0303 | AC41 | HEK-293T | C192(0.99); C202(0.87); C175(0.96); C67(0.94) | LDD1542 | [19] |
| LDCM0304 | AC42 | HEK-293T | C192(1.09); C202(0.94); C175(1.10); C67(0.94) | LDD1543 | [19] |
| LDCM0305 | AC43 | HEK-293T | C192(0.87); C202(0.94); C175(1.12); C67(1.03) | LDD1544 | [19] |
| LDCM0306 | AC44 | HEK-293T | C192(0.90); C202(0.98); C175(1.03); C67(0.82) | LDD1545 | [19] |
| LDCM0307 | AC45 | HEK-293T | C192(0.96); C202(0.94); C175(1.05); C67(0.88) | LDD1546 | [19] |
| LDCM0308 | AC46 | HEK-293T | C192(1.01); C175(1.15); C67(1.18) | LDD1547 | [19] |
| LDCM0309 | AC47 | HEK-293T | C192(1.06); C202(1.15); C175(0.97); C67(1.00) | LDD1548 | [19] |
| LDCM0310 | AC48 | HEK-293T | C192(1.14); C175(1.05); C67(0.96) | LDD1549 | [19] |
| LDCM0311 | AC49 | HEK-293T | C192(0.95); C202(0.91); C175(1.01); C67(0.85) | LDD1550 | [19] |
| LDCM0312 | AC5 | HEK-293T | C192(1.03); C202(1.00); C175(1.06); C67(0.87) | LDD1551 | [19] |
| LDCM0313 | AC50 | HEK-293T | C192(1.06); C202(0.94); C175(1.09); C67(1.03) | LDD1552 | [19] |
| LDCM0314 | AC51 | HEK-293T | C192(0.96); C202(0.98); C175(1.06); C67(1.04) | LDD1553 | [19] |
| LDCM0315 | AC52 | HEK-293T | C192(0.94); C202(0.93); C175(1.09); C67(0.96) | LDD1554 | [19] |
| LDCM0316 | AC53 | HEK-293T | C192(0.98); C202(1.06); C175(1.10); C67(0.99) | LDD1555 | [19] |
| LDCM0317 | AC54 | HEK-293T | C192(0.99); C175(1.01); C67(1.04) | LDD1556 | [19] |
| LDCM0318 | AC55 | HEK-293T | C192(1.06); C202(1.17); C175(1.07); C67(1.00) | LDD1557 | [19] |
| LDCM0319 | AC56 | HEK-293T | C192(1.13); C175(1.19); C67(0.95) | LDD1558 | [19] |
| LDCM0320 | AC57 | HEK-293T | C192(0.95); C202(0.98); C175(0.91); C67(1.00) | LDD1559 | [19] |
| LDCM0321 | AC58 | HEK-293T | C192(1.12); C202(0.96); C175(1.09); C67(1.03) | LDD1560 | [19] |
| LDCM0322 | AC59 | HEK-293T | C192(0.91); C202(0.92); C175(1.29); C67(1.06) | LDD1561 | [19] |
| LDCM0323 | AC6 | HEK-293T | C192(0.97); C175(1.11); C67(0.94) | LDD1562 | [19] |
| LDCM0324 | AC60 | HEK-293T | C192(0.97); C202(0.98); C175(1.10); C67(0.92) | LDD1563 | [19] |
| LDCM0325 | AC61 | HEK-293T | C192(0.98); C202(1.05); C175(1.04); C67(0.93) | LDD1564 | [19] |
| LDCM0326 | AC62 | HEK-293T | C192(1.05); C175(1.05); C67(1.11) | LDD1565 | [19] |
| LDCM0327 | AC63 | HEK-293T | C192(1.11); C202(1.05); C175(1.02); C67(0.99) | LDD1566 | [19] |
| LDCM0328 | AC64 | HEK-293T | C192(1.14); C175(1.06); C67(1.00) | LDD1567 | [19] |
| LDCM0334 | AC7 | HEK-293T | C192(1.06); C202(1.07); C175(1.09); C67(1.02) | LDD1568 | [19] |
| LDCM0345 | AC8 | HEK-293T | C192(1.16); C175(1.17); C67(0.90) | LDD1569 | [19] |
| LDCM0545 | Acetamide | MDA-MB-231 | C341(0.54) | LDD2138 | [4] |
| LDCM0248 | AKOS034007472 | HEK-293T | C192(1.01); C202(1.01); C175(1.04); C67(0.95) | LDD1511 | [19] |
| LDCM0356 | AKOS034007680 | HEK-293T | C192(1.00); C202(0.92); C175(1.07); C67(0.92) | LDD1570 | [19] |
| LDCM0275 | AKOS034007705 | HEK-293T | C192(1.01); C175(1.12); C67(0.95) | LDD1514 | [19] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C341(0.50) | LDD2091 | [4] |
| LDCM0108 | Chloroacetamide | HeLa | C67(0.00); C192(0.00) | LDD0222 | [16] |
| LDCM0632 | CL-Sc | Hep-G2 | C251(20.00); C175(1.07); C251(1.05); C67(0.90) | LDD2227 | [18] |
| LDCM0367 | CL1 | HEK-293T | C192(1.14); C202(1.04); C175(1.09) | LDD1571 | [19] |
| LDCM0368 | CL10 | HEK-293T | C192(1.34); C175(1.31); C67(1.09) | LDD1572 | [19] |
| LDCM0369 | CL100 | HEK-293T | C192(0.92); C202(0.92); C175(1.07); C67(1.02) | LDD1573 | [19] |
| LDCM0370 | CL101 | HEK-293T | C192(1.07); C202(0.90); C175(1.04) | LDD1574 | [19] |
| LDCM0371 | CL102 | HEK-293T | C192(0.95); C202(1.02); C175(1.01); C67(0.92) | LDD1575 | [19] |
| LDCM0372 | CL103 | HEK-293T | C192(1.03); C202(1.00); C175(1.11); C67(0.96) | LDD1576 | [19] |
| LDCM0373 | CL104 | HEK-293T | C192(1.12); C202(1.02); C175(1.10); C67(1.01) | LDD1577 | [19] |
| LDCM0374 | CL105 | HEK-293T | C192(1.01); C202(0.90); C175(1.07) | LDD1578 | [19] |
| LDCM0375 | CL106 | HEK-293T | C192(1.02); C202(1.04); C175(1.01); C67(1.01) | LDD1579 | [19] |
| LDCM0376 | CL107 | HEK-293T | C192(0.98); C202(1.05); C175(1.07); C67(1.06) | LDD1580 | [19] |
| LDCM0377 | CL108 | HEK-293T | C192(1.07); C202(0.87); C175(1.13); C67(1.06) | LDD1581 | [19] |
| LDCM0378 | CL109 | HEK-293T | C192(1.04); C202(1.01); C175(0.96) | LDD1582 | [19] |
| LDCM0379 | CL11 | HEK-293T | C192(1.16); C202(1.24); C175(1.21); C67(0.94) | LDD1583 | [19] |
| LDCM0380 | CL110 | HEK-293T | C192(1.06); C202(1.02); C175(1.04); C67(0.96) | LDD1584 | [19] |
| LDCM0381 | CL111 | HEK-293T | C192(1.02); C202(0.98); C175(1.04); C67(1.00) | LDD1585 | [19] |
| LDCM0382 | CL112 | HEK-293T | C192(0.96); C202(0.89); C175(1.12); C67(0.91) | LDD1586 | [19] |
| LDCM0383 | CL113 | HEK-293T | C192(1.08); C202(0.99); C175(0.95) | LDD1587 | [19] |
| LDCM0384 | CL114 | HEK-293T | C192(1.07); C202(1.04); C175(1.05); C67(0.94) | LDD1588 | [19] |
| LDCM0385 | CL115 | HEK-293T | C192(1.02); C202(1.03); C175(1.06); C67(0.98) | LDD1589 | [19] |
| LDCM0386 | CL116 | HEK-293T | C192(0.99); C202(0.92); C175(1.12); C67(0.90) | LDD1590 | [19] |
| LDCM0387 | CL117 | HEK-293T | C192(1.05); C202(0.99); C175(1.03) | LDD1591 | [19] |
| LDCM0388 | CL118 | HEK-293T | C192(0.97); C202(0.97); C175(1.03); C67(0.89) | LDD1592 | [19] |
| LDCM0389 | CL119 | HEK-293T | C192(1.04); C202(1.08); C175(1.22); C67(0.95) | LDD1593 | [19] |
| LDCM0390 | CL12 | HEK-293T | C192(1.23); C175(1.23); C67(1.08) | LDD1594 | [19] |
| LDCM0391 | CL120 | HEK-293T | C192(0.97); C202(1.02); C175(1.12); C67(0.93) | LDD1595 | [19] |
| LDCM0392 | CL121 | HEK-293T | C192(1.08); C202(0.98); C175(1.06) | LDD1596 | [19] |
| LDCM0393 | CL122 | HEK-293T | C192(0.98); C202(1.15); C175(1.00); C67(0.96) | LDD1597 | [19] |
| LDCM0394 | CL123 | HEK-293T | C192(1.20); C202(1.24); C175(1.30); C67(0.92) | LDD1598 | [19] |
| LDCM0395 | CL124 | HEK-293T | C192(1.00); C202(0.99); C175(1.04); C67(1.05) | LDD1599 | [19] |
| LDCM0396 | CL125 | HEK-293T | C192(1.07); C202(1.00); C175(1.01) | LDD1600 | [19] |
| LDCM0397 | CL126 | HEK-293T | C192(1.01); C202(1.03); C175(1.01); C67(1.01) | LDD1601 | [19] |
| LDCM0398 | CL127 | HEK-293T | C192(1.00); C202(1.08); C175(1.09); C67(1.00) | LDD1602 | [19] |
| LDCM0399 | CL128 | HEK-293T | C192(1.06); C202(1.03); C175(1.12); C67(0.98) | LDD1603 | [19] |
| LDCM0400 | CL13 | HEK-293T | C192(0.90); C202(0.92); C175(0.95) | LDD1604 | [19] |
| LDCM0401 | CL14 | HEK-293T | C192(0.90); C202(1.02); C175(1.03); C67(0.91) | LDD1605 | [19] |
| LDCM0402 | CL15 | HEK-293T | C192(1.11); C202(1.12); C175(1.11); C67(1.01) | LDD1606 | [19] |
| LDCM0403 | CL16 | HEK-293T | C192(0.98); C202(0.95); C175(1.16); C67(0.92) | LDD1607 | [19] |
| LDCM0404 | CL17 | HEK-293T | C192(0.95); C202(1.05); C175(1.11); C67(0.79) | LDD1608 | [19] |
| LDCM0405 | CL18 | HEK-293T | C192(1.11); C202(1.00); C175(1.00); C67(0.95) | LDD1609 | [19] |
| LDCM0406 | CL19 | HEK-293T | C192(1.11); C202(1.05); C175(1.07); C67(0.90) | LDD1610 | [19] |
| LDCM0407 | CL2 | HEK-293T | C192(0.91); C202(1.06); C175(1.00); C67(0.85) | LDD1611 | [19] |
| LDCM0408 | CL20 | HEK-293T | C192(0.93); C202(1.02); C175(1.20); C67(0.81) | LDD1612 | [19] |
| LDCM0409 | CL21 | HEK-293T | C192(1.27); C202(1.16); C175(1.12); C67(0.96) | LDD1613 | [19] |
| LDCM0410 | CL22 | HEK-293T | C192(1.09); C175(1.14); C67(0.99) | LDD1614 | [19] |
| LDCM0411 | CL23 | HEK-293T | C192(1.13); C202(1.22); C175(1.17); C67(0.91) | LDD1615 | [19] |
| LDCM0412 | CL24 | HEK-293T | C192(1.19); C175(1.27); C67(1.03) | LDD1616 | [19] |
| LDCM0413 | CL25 | HEK-293T | C192(1.03); C202(0.98); C175(1.06) | LDD1617 | [19] |
| LDCM0414 | CL26 | HEK-293T | C192(0.94); C202(1.06); C175(0.91); C67(0.96) | LDD1618 | [19] |
| LDCM0415 | CL27 | HEK-293T | C192(1.00); C202(1.08); C175(1.09); C67(1.05) | LDD1619 | [19] |
| LDCM0416 | CL28 | HEK-293T | C192(1.06); C202(0.96); C175(1.08); C67(0.95) | LDD1620 | [19] |
| LDCM0417 | CL29 | HEK-293T | C192(1.07); C202(1.05); C175(1.00); C67(0.87) | LDD1621 | [19] |
| LDCM0418 | CL3 | HEK-293T | C192(1.04); C202(1.14); C175(1.23); C67(0.99) | LDD1622 | [19] |
| LDCM0419 | CL30 | HEK-293T | C192(1.08); C202(1.03); C175(1.13); C67(0.96) | LDD1623 | [19] |
| LDCM0420 | CL31 | HEK-293T | C192(1.00); C202(0.97); C175(1.05); C67(0.96) | LDD1624 | [19] |
| LDCM0421 | CL32 | HEK-293T | C192(0.98); C202(1.01); C175(1.26); C67(0.93) | LDD1625 | [19] |
| LDCM0422 | CL33 | HEK-293T | C192(1.30); C202(1.37); C175(1.26); C67(0.95) | LDD1626 | [19] |
| LDCM0423 | CL34 | HEK-293T | C192(1.15); C175(1.34); C67(1.32) | LDD1627 | [19] |
| LDCM0424 | CL35 | HEK-293T | C192(1.18); C202(1.18); C175(1.23); C67(0.98) | LDD1628 | [19] |
| LDCM0425 | CL36 | HEK-293T | C192(1.28); C175(1.13); C67(1.06) | LDD1629 | [19] |
| LDCM0426 | CL37 | HEK-293T | C192(1.04); C202(0.97); C175(0.99) | LDD1630 | [19] |
| LDCM0428 | CL39 | HEK-293T | C192(1.01); C202(1.00); C175(1.15); C67(1.07) | LDD1632 | [19] |
| LDCM0429 | CL4 | HEK-293T | C192(1.09); C202(1.06); C175(1.20); C67(0.93) | LDD1633 | [19] |
| LDCM0430 | CL40 | HEK-293T | C192(1.16); C202(0.84); C175(1.05); C67(0.99) | LDD1634 | [19] |
| LDCM0431 | CL41 | HEK-293T | C192(1.07); C202(1.05); C175(1.02); C67(0.84) | LDD1635 | [19] |
| LDCM0432 | CL42 | HEK-293T | C192(1.09); C202(1.10); C175(0.97); C67(1.03) | LDD1636 | [19] |
| LDCM0433 | CL43 | HEK-293T | C192(1.19); C202(0.98); C175(1.18); C67(1.00) | LDD1637 | [19] |
| LDCM0434 | CL44 | HEK-293T | C192(1.03); C202(0.98); C175(1.10); C67(0.90) | LDD1638 | [19] |
| LDCM0435 | CL45 | HEK-293T | C192(1.10); C202(1.11); C175(1.23); C67(0.89) | LDD1639 | [19] |
| LDCM0436 | CL46 | HEK-293T | C192(1.16); C175(1.28); C67(1.13) | LDD1640 | [19] |
| LDCM0437 | CL47 | HEK-293T | C192(1.10); C202(1.30); C175(1.14); C67(1.01) | LDD1641 | [19] |
| LDCM0438 | CL48 | HEK-293T | C192(1.20); C175(1.24); C67(1.03) | LDD1642 | [19] |
| LDCM0439 | CL49 | HEK-293T | C192(1.18); C202(1.10); C175(1.05) | LDD1643 | [19] |
| LDCM0440 | CL5 | HEK-293T | C192(0.83); C202(0.99); C175(0.99); C67(0.76) | LDD1644 | [19] |
| LDCM0441 | CL50 | HEK-293T | C192(0.96); C202(1.04); C175(1.07); C67(0.94) | LDD1645 | [19] |
| LDCM0443 | CL52 | HEK-293T | C192(0.96); C202(0.97); C175(1.09); C67(0.81) | LDD1646 | [19] |
| LDCM0444 | CL53 | HEK-293T | C192(1.03); C202(1.10); C175(0.98); C67(0.87) | LDD1647 | [19] |
| LDCM0445 | CL54 | HEK-293T | C192(1.27); C202(1.17); C175(1.37); C67(0.93) | LDD1648 | [19] |
| LDCM0446 | CL55 | HEK-293T | C192(0.98); C202(0.96); C175(1.25); C67(1.14) | LDD1649 | [19] |
| LDCM0447 | CL56 | HEK-293T | C192(0.95); C202(1.09); C175(1.16); C67(1.04) | LDD1650 | [19] |
| LDCM0448 | CL57 | HEK-293T | C192(1.19); C202(1.12); C175(1.05); C67(0.83) | LDD1651 | [19] |
| LDCM0449 | CL58 | HEK-293T | C192(1.23); C175(1.11); C67(1.08) | LDD1652 | [19] |
| LDCM0450 | CL59 | HEK-293T | C192(1.26); C202(1.17); C175(1.18); C67(1.01) | LDD1653 | [19] |
| LDCM0451 | CL6 | HEK-293T | C192(1.16); C202(1.14); C175(1.00); C67(0.94) | LDD1654 | [19] |
| LDCM0452 | CL60 | HEK-293T | C192(1.27); C175(1.13); C67(0.98) | LDD1655 | [19] |
| LDCM0453 | CL61 | HEK-293T | C192(1.05); C202(1.01); C175(1.02) | LDD1656 | [19] |
| LDCM0454 | CL62 | HEK-293T | C192(0.92); C202(1.04); C175(0.97); C67(1.03) | LDD1657 | [19] |
| LDCM0455 | CL63 | HEK-293T | C192(1.00); C202(1.19); C175(1.18); C67(0.97) | LDD1658 | [19] |
| LDCM0456 | CL64 | HEK-293T | C192(1.11); C202(1.12); C175(1.04); C67(0.96) | LDD1659 | [19] |
| LDCM0457 | CL65 | HEK-293T | C192(0.98); C202(1.09); C175(1.06); C67(0.92) | LDD1660 | [19] |
| LDCM0458 | CL66 | HEK-293T | C192(1.03); C202(1.01); C175(1.09); C67(0.91) | LDD1661 | [19] |
| LDCM0459 | CL67 | HEK-293T | C192(0.94); C202(0.95); C175(1.27); C67(0.96) | LDD1662 | [19] |
| LDCM0460 | CL68 | HEK-293T | C192(1.03); C202(1.07); C175(1.19); C67(0.95) | LDD1663 | [19] |
| LDCM0461 | CL69 | HEK-293T | C192(1.11); C202(1.29); C175(1.19); C67(0.86) | LDD1664 | [19] |
| LDCM0462 | CL7 | HEK-293T | C192(1.13); C202(1.15); C175(1.07); C67(0.98) | LDD1665 | [19] |
| LDCM0463 | CL70 | HEK-293T | C192(1.13); C175(1.11); C67(1.01) | LDD1666 | [19] |
| LDCM0464 | CL71 | HEK-293T | C192(1.19); C202(1.16); C175(1.32); C67(0.98) | LDD1667 | [19] |
| LDCM0465 | CL72 | HEK-293T | C192(1.13); C175(1.23); C67(1.11) | LDD1668 | [19] |
| LDCM0466 | CL73 | HEK-293T | C192(1.11); C202(1.05); C175(1.04) | LDD1669 | [19] |
| LDCM0467 | CL74 | HEK-293T | C192(0.99); C202(1.06); C175(1.01); C67(0.95) | LDD1670 | [19] |
| LDCM0469 | CL76 | HEK-293T | C192(0.98); C202(1.08); C175(1.00); C67(0.93) | LDD1672 | [19] |
| LDCM0470 | CL77 | HEK-293T | C192(1.04); C202(0.95); C175(1.06); C67(0.91) | LDD1673 | [19] |
| LDCM0471 | CL78 | HEK-293T | C192(1.06); C202(1.04); C175(1.09); C67(0.93) | LDD1674 | [19] |
| LDCM0472 | CL79 | HEK-293T | C192(1.06); C202(0.95); C175(1.09); C67(1.09) | LDD1675 | [19] |
| LDCM0473 | CL8 | HEK-293T | C192(1.24); C202(1.22); C175(1.47); C67(0.92) | LDD1676 | [19] |
| LDCM0474 | CL80 | HEK-293T | C192(1.02); C202(1.08); C175(1.17); C67(1.00) | LDD1677 | [19] |
| LDCM0475 | CL81 | HEK-293T | C192(1.07); C202(1.24); C175(1.05); C67(0.88) | LDD1678 | [19] |
| LDCM0476 | CL82 | HEK-293T | C192(1.14); C175(1.16); C67(1.20) | LDD1679 | [19] |
| LDCM0477 | CL83 | HEK-293T | C192(1.18); C202(1.26); C175(1.13); C67(1.01) | LDD1680 | [19] |
| LDCM0478 | CL84 | HEK-293T | C192(1.29); C175(1.11); C67(0.97) | LDD1681 | [19] |
| LDCM0479 | CL85 | HEK-293T | C192(1.09); C202(1.06); C175(0.93) | LDD1682 | [19] |
| LDCM0480 | CL86 | HEK-293T | C192(0.95); C202(1.12); C175(1.11); C67(0.93) | LDD1683 | [19] |
| LDCM0481 | CL87 | HEK-293T | C192(1.00); C202(1.07); C175(1.20); C67(1.01) | LDD1684 | [19] |
| LDCM0482 | CL88 | HEK-293T | C192(0.97); C202(1.11); C175(1.11); C67(1.00) | LDD1685 | [19] |
| LDCM0483 | CL89 | HEK-293T | C192(0.98); C202(0.99); C175(1.00); C67(0.84) | LDD1686 | [19] |
| LDCM0484 | CL9 | HEK-293T | C192(1.10); C202(1.21); C175(1.20); C67(0.77) | LDD1687 | [19] |
| LDCM0485 | CL90 | HEK-293T | C192(1.36); C202(1.08); C175(1.56); C67(0.91) | LDD1688 | [19] |
| LDCM0486 | CL91 | HEK-293T | C192(1.07); C202(1.07); C175(1.27); C67(1.02) | LDD1689 | [19] |
| LDCM0487 | CL92 | HEK-293T | C192(1.02); C202(1.05); C175(1.30); C67(0.94) | LDD1690 | [19] |
| LDCM0488 | CL93 | HEK-293T | C192(1.18); C202(1.08); C175(1.14); C67(0.86) | LDD1691 | [19] |
| LDCM0489 | CL94 | HEK-293T | C192(1.06); C175(1.11); C67(1.11) | LDD1692 | [19] |
| LDCM0490 | CL95 | HEK-293T | C192(1.33); C202(1.24); C175(1.32); C67(1.07) | LDD1693 | [19] |
| LDCM0491 | CL96 | HEK-293T | C192(1.23); C175(1.10); C67(0.98) | LDD1694 | [19] |
| LDCM0492 | CL97 | HEK-293T | C192(1.03); C202(0.97); C175(1.01) | LDD1695 | [19] |
| LDCM0493 | CL98 | HEK-293T | C192(0.91); C202(1.00); C175(1.12); C67(0.91) | LDD1696 | [19] |
| LDCM0494 | CL99 | HEK-293T | C192(1.03); C202(1.17); C175(1.04); C67(0.99) | LDD1697 | [19] |
| LDCM0495 | E2913 | HEK-293T | C192(0.95); C202(1.19); C175(1.13); C67(1.00) | LDD1698 | [19] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C251(2.48); C341(1.29) | LDD1702 | [4] |
| LDCM0625 | F8 | Ramos | C67(0.69); C251(1.17) | LDD2187 | [20] |
| LDCM0572 | Fragment10 | Ramos | C67(0.45); C192(0.58); C251(1.24) | LDD2189 | [20] |
| LDCM0573 | Fragment11 | Ramos | C67(0.25); C192(20.00); C251(0.97) | LDD2190 | [20] |
| LDCM0574 | Fragment12 | Ramos | C67(0.63); C192(0.36); C251(0.96) | LDD2191 | [20] |
| LDCM0575 | Fragment13 | Ramos | C67(1.14); C192(0.57); C251(0.90) | LDD2192 | [20] |
| LDCM0576 | Fragment14 | Ramos | C67(0.93); C192(0.90); C251(1.50) | LDD2193 | [20] |
| LDCM0579 | Fragment20 | Ramos | C67(0.46); C192(0.24); C251(1.19) | LDD2194 | [20] |
| LDCM0580 | Fragment21 | Ramos | C67(1.21); C192(0.71); C251(0.77) | LDD2195 | [20] |
| LDCM0582 | Fragment23 | Ramos | C67(0.93); C192(0.57); C251(0.42) | LDD2196 | [20] |
| LDCM0578 | Fragment27 | Ramos | C67(1.12); C192(0.66); C251(0.87) | LDD2197 | [20] |
| LDCM0586 | Fragment28 | Ramos | C67(0.76); C192(0.73); C251(1.03) | LDD2198 | [20] |
| LDCM0588 | Fragment30 | Ramos | C67(1.24); C251(0.93) | LDD2199 | [20] |
| LDCM0589 | Fragment31 | Ramos | C67(1.95); C251(1.71) | LDD2200 | [20] |
| LDCM0590 | Fragment32 | Ramos | C67(0.65); C192(0.50); C251(0.51) | LDD2201 | [20] |
| LDCM0468 | Fragment33 | HEK-293T | C192(0.99); C202(1.19); C175(1.25); C67(1.04) | LDD1671 | [19] |
| LDCM0596 | Fragment38 | Ramos | C67(0.89); C192(0.57); C251(0.89) | LDD2203 | [20] |
| LDCM0566 | Fragment4 | Ramos | C67(1.17); C251(1.11) | LDD2184 | [20] |
| LDCM0427 | Fragment51 | HEK-293T | C192(0.98); C202(1.07); C175(0.98); C67(0.96) | LDD1631 | [19] |
| LDCM0610 | Fragment52 | Ramos | C67(1.13); C251(1.33) | LDD2204 | [20] |
| LDCM0614 | Fragment56 | Ramos | C67(1.36); C192(0.97); C251(1.11) | LDD2205 | [20] |
| LDCM0569 | Fragment7 | Ramos | C67(0.40); C192(0.41); C251(0.68) | LDD2186 | [20] |
| LDCM0571 | Fragment9 | Ramos | C67(0.43); C192(0.36); C251(1.14) | LDD2188 | [20] |
| LDCM0116 | HHS-0101 | DM93 | Y381(0.69) | LDD0264 | [6] |
| LDCM0117 | HHS-0201 | DM93 | Y381(1.09) | LDD0265 | [6] |
| LDCM0118 | HHS-0301 | DM93 | Y381(1.20) | LDD0266 | [6] |
| LDCM0119 | HHS-0401 | DM93 | Y381(1.27) | LDD0267 | [6] |
| LDCM0120 | HHS-0701 | DM93 | Y381(1.14) | LDD0268 | [6] |
| LDCM0107 | IAA | HeLa | H66(0.00); C67(0.00) | LDD0221 | [16] |
| LDCM0179 | JZ128 | PC-3 | N.A. | LDD0462 | [3] |
| LDCM0022 | KB02 | HEK-293T | C192(0.84); C175(1.02) | LDD1492 | [19] |
| LDCM0023 | KB03 | HEK-293T | C192(0.94); C175(0.90) | LDD1497 | [19] |
| LDCM0024 | KB05 | G361 | C228(5.27); C238(1.87); C67(2.33) | LDD3311 | [2] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C251(1.03) | LDD2102 | [4] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C192(1.34) | LDD2121 | [4] |
| LDCM0109 | NEM | HeLa | H95(0.00); H361(0.00) | LDD0223 | [16] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C341(0.62) | LDD2089 | [4] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C341(1.27) | LDD2090 | [4] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C251(1.05) | LDD2093 | [4] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C251(0.23) | LDD2096 | [4] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C251(1.03) | LDD2097 | [4] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C192(0.87); C341(1.20); C251(0.91); C67(1.09) | LDD2099 | [4] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C341(0.60) | LDD2100 | [4] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C341(0.67) | LDD2104 | [4] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C192(0.82); C251(0.46) | LDD2107 | [4] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C192(0.94); C341(0.50); C251(0.62) | LDD2108 | [4] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C341(0.72); C251(0.52) | LDD2109 | [4] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C341(0.87) | LDD2110 | [4] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C251(0.87) | LDD2111 | [4] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C251(0.53) | LDD2115 | [4] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C251(0.41) | LDD2116 | [4] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C341(0.26); C251(0.48) | LDD2118 | [4] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C341(1.90); C251(1.69); C67(1.12) | LDD2119 | [4] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C251(1.03) | LDD2120 | [4] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C341(0.17) | LDD2122 | [4] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C192(0.94); C251(0.65); C67(1.22) | LDD2123 | [4] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C341(0.10); C251(0.29) | LDD2124 | [4] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C192(0.76); C341(1.00); C251(0.73); C67(0.84) | LDD2125 | [4] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C251(0.34) | LDD2126 | [4] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C192(1.45); C251(0.93) | LDD2127 | [4] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C341(1.07); C251(0.99) | LDD2128 | [4] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C192(0.99); C341(0.98) | LDD2129 | [4] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C251(0.74) | LDD2134 | [4] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C251(1.31) | LDD2135 | [4] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C192(1.20); C341(1.21); C251(1.47) | LDD2136 | [4] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C341(0.96); C251(0.86); C67(1.06) | LDD2137 | [4] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C341(2.34) | LDD1700 | [4] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C192(0.73); C341(0.92); C251(0.61) | LDD2140 | [4] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C341(0.65); C251(0.50) | LDD2141 | [4] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C341(0.49); C251(1.00) | LDD2143 | [4] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C192(1.54); C341(1.98) | LDD2144 | [4] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C251(0.14) | LDD2145 | [4] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C251(0.84); C67(1.30) | LDD2146 | [4] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C341(1.05) | LDD2147 | [4] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C341(0.46) | LDD2148 | [4] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C251(0.27) | LDD2149 | [4] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C341(0.45) | LDD2150 | [4] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C251(0.29) | LDD2151 | [4] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C341(1.63) | LDD2153 | [4] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| COP9 signalosome complex subunit 5 (COPS5) | Peptidase M67A family | Q92905 | |||
Other
References
