Details of the Target
General Information of Target
| Target ID | LDTP04914 | |||||
|---|---|---|---|---|---|---|
| Target Name | Large ribosomal subunit protein uL24 (RPL26) | |||||
| Gene Name | RPL26 | |||||
| Gene ID | 6154 | |||||
| Synonyms |
Large ribosomal subunit protein uL24; 60S ribosomal protein L26 |
|||||
| 3D Structure | ||||||
| Sequence |
MKFNPFVTSDRSKNRKRHFNAPSHIRRKIMSSPLSKELRQKYNVRSMPIRKDDEVQVVRG
HYKGQQIGKVVQVYRKKYVIYIERVQREKANGTTVHVGIHPSKVVITRLKLDKDRKKILE RKAKSRQVGKEKGKYKEETIEKMQE |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Universal ribosomal protein uL24 family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function | Component of the large ribosomal subunit. The ribosome is a large ribonucleoprotein complex responsible for the synthesis of proteins in the cell. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
A-EBA Probe Info |
 |
3.21 | LDD0215 | [1] | |
|
AZ-9 Probe Info |
 |
4.39 | LDD0393 | [2] | |
|
C-Sul Probe Info |
 |
4.15 | LDD0066 | [3] | |
|
TH211 Probe Info |
 |
Y74(20.00); Y78(13.67); Y135(10.70) | LDD0257 | [4] | |
|
TH216 Probe Info |
 |
Y78(17.86); Y74(16.24); Y135(13.38) | LDD0259 | [4] | |
|
STPyne Probe Info |
 |
K110(4.40); K134(10.00); K136(6.67); K2(8.45) | LDD0277 | [5] | |
|
ONAyne Probe Info |
 |
K28(0.00); K36(0.00); K41(0.00); K51(0.00) | LDD0273 | [5] | |
|
OPA-S-S-alkyne Probe Info |
 |
K36(1.79); K51(2.85) | LDD3494 | [6] | |
|
HHS-475 Probe Info |
 |
Y135(0.77) | LDD0264 | [7] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [8] | |
|
AMP probe Probe Info |
 |
N.A. | LDD0200 | [9] | |
|
ATP probe Probe Info |
 |
K36(0.00); K28(0.00); K51(0.00); K134(0.00) | LDD0199 | [9] | |
|
m-APA Probe Info |
 |
N.A. | LDD2231 | [8] | |
|
ATP probe Probe Info |
 |
K134(0.00); K77(0.00); K103(0.00); K63(0.00) | LDD0035 | [10] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [11] | |
|
OSF Probe Info |
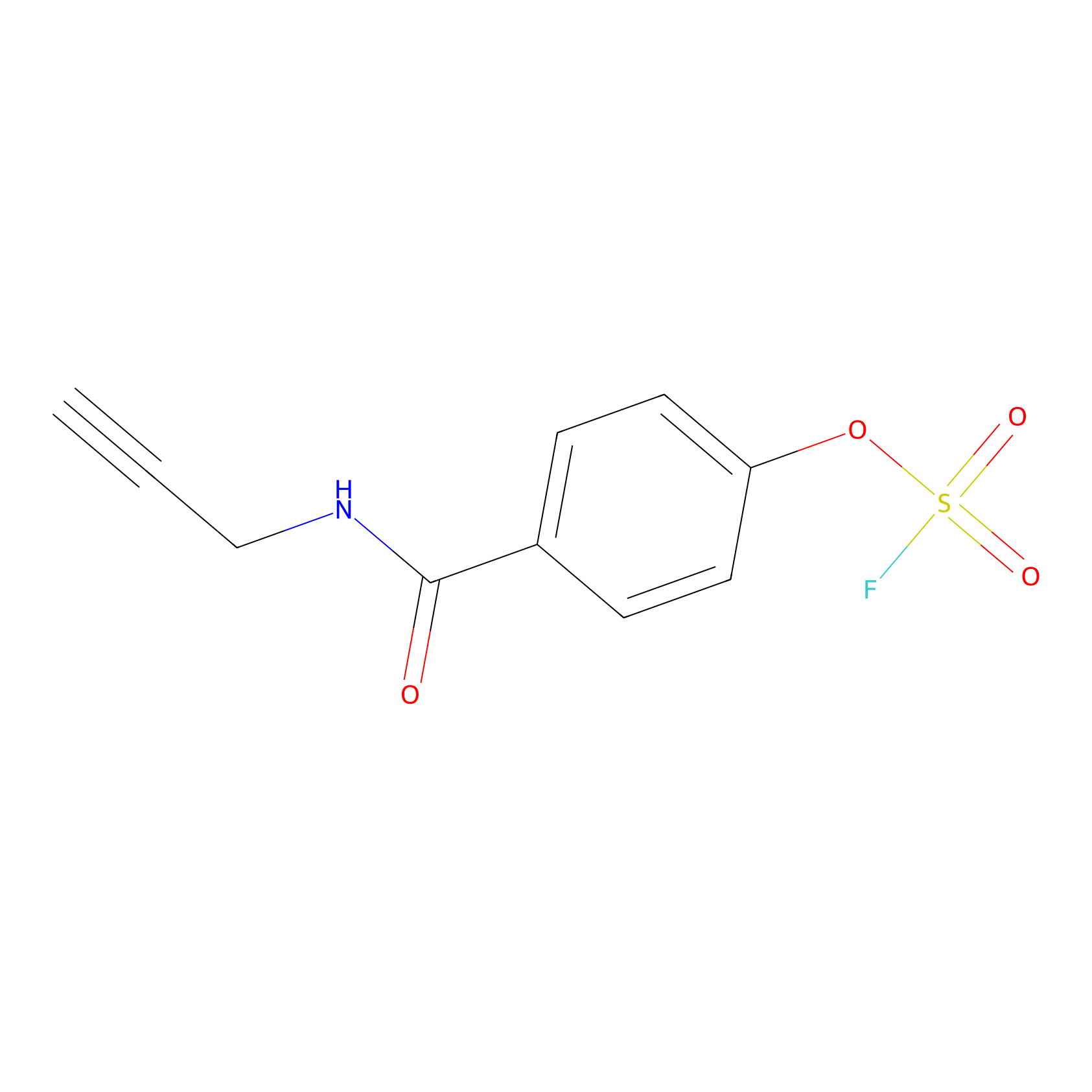 |
Y74(0.00); H96(0.00) | LDD0029 | [12] | |
|
SF Probe Info |
 |
Y81(0.00); Y135(0.00); Y78(0.00); Y74(0.00) | LDD0028 | [12] | |
|
Acrolein Probe Info |
 |
H24(0.00); H18(0.00) | LDD0217 | [13] | |
|
Crotonaldehyde Probe Info |
 |
H24(0.00); H18(0.00) | LDD0219 | [13] | |
|
Methacrolein Probe Info |
 |
H24(0.00); H18(0.00) | LDD0218 | [13] | |
|
HHS-482 Probe Info |
 |
Y135(1.28) | LDD2239 | [14] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C187 Probe Info |
 |
18.13 | LDD1865 | [15] | |
|
C191 Probe Info |
 |
14.32 | LDD1868 | [15] | |
|
C193 Probe Info |
 |
7.06 | LDD1869 | [15] | |
|
C405 Probe Info |
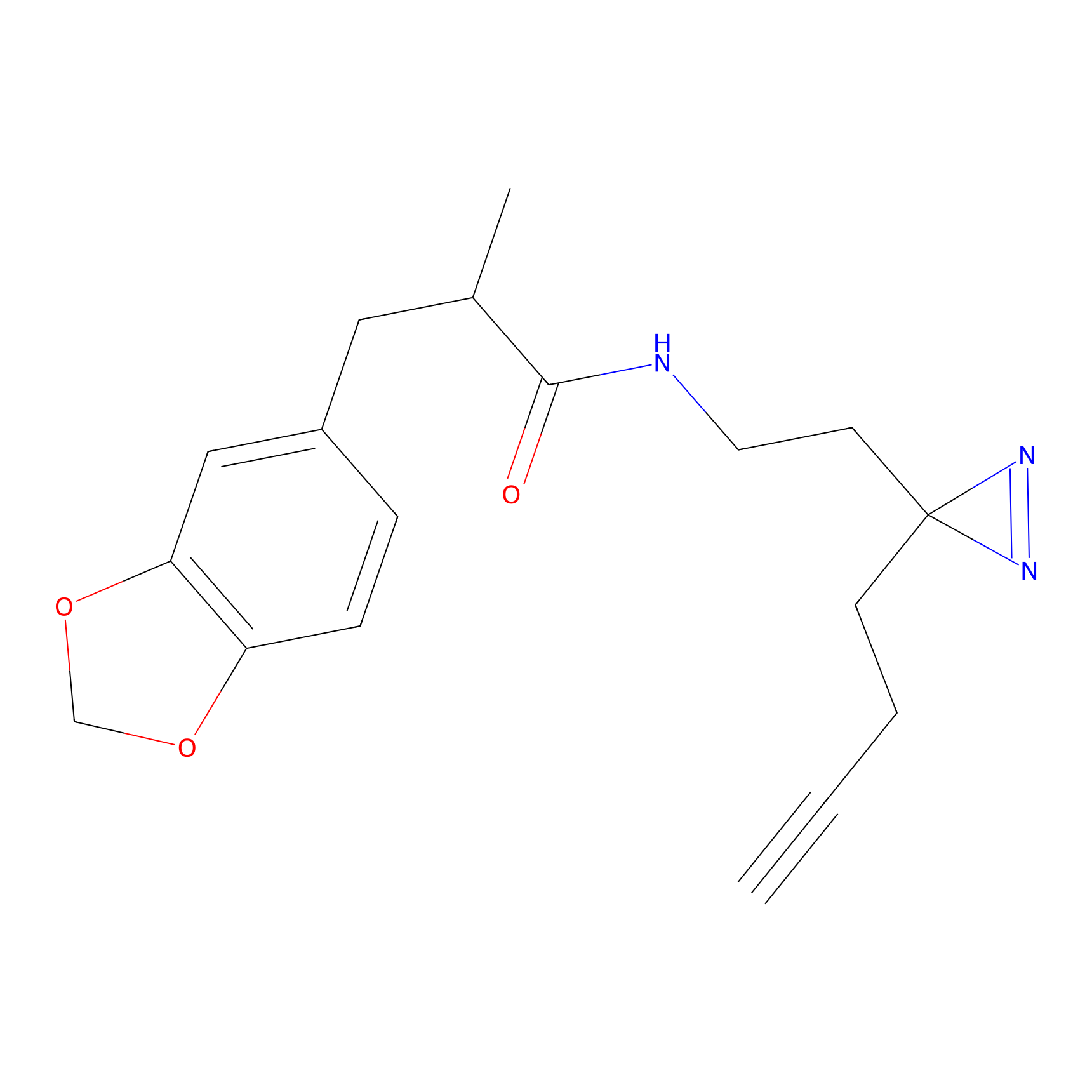 |
5.70 | LDD2063 | [15] | |
|
C407 Probe Info |
 |
19.43 | LDD2064 | [15] | |
|
C408 Probe Info |
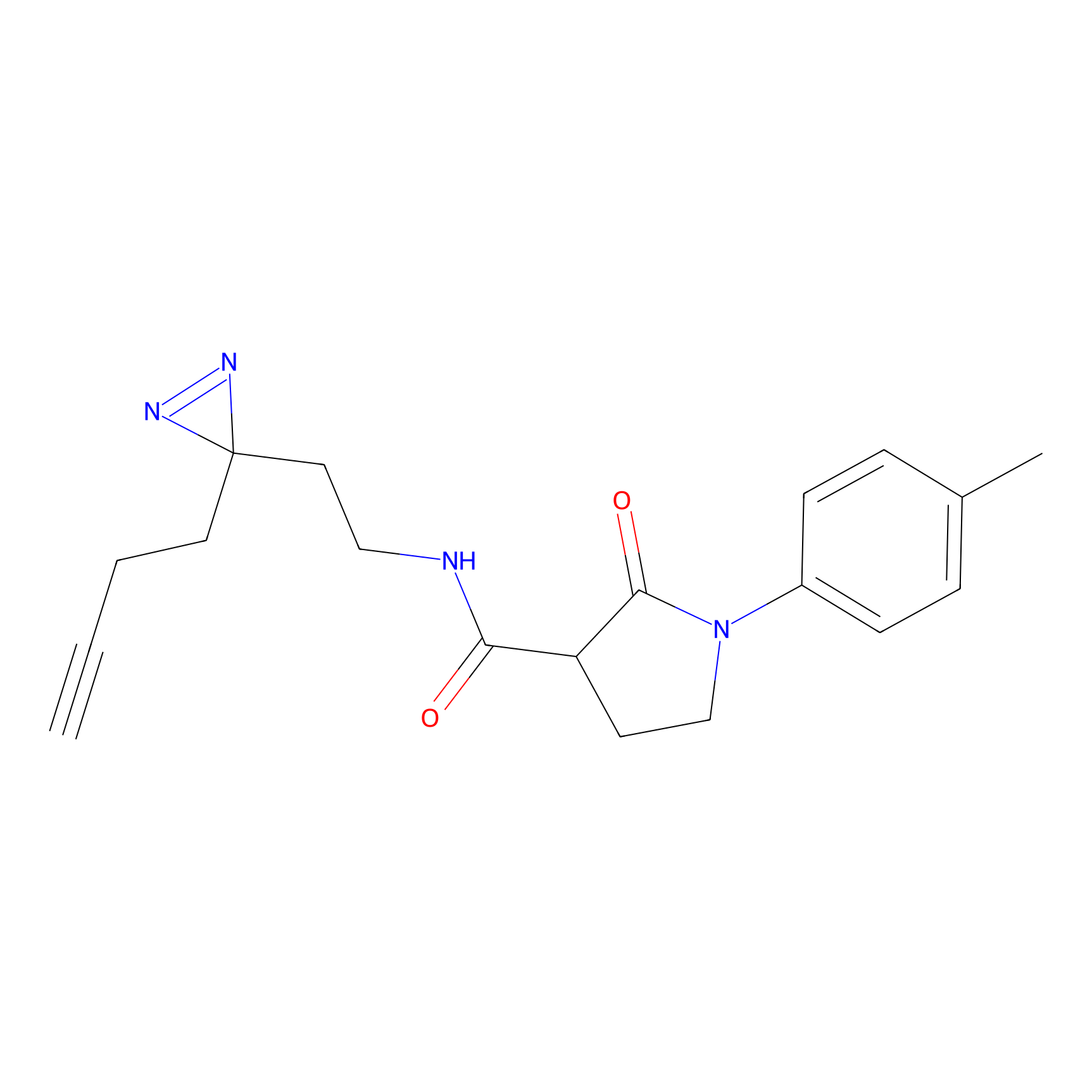 |
6.06 | LDD2065 | [15] | |
|
C409 Probe Info |
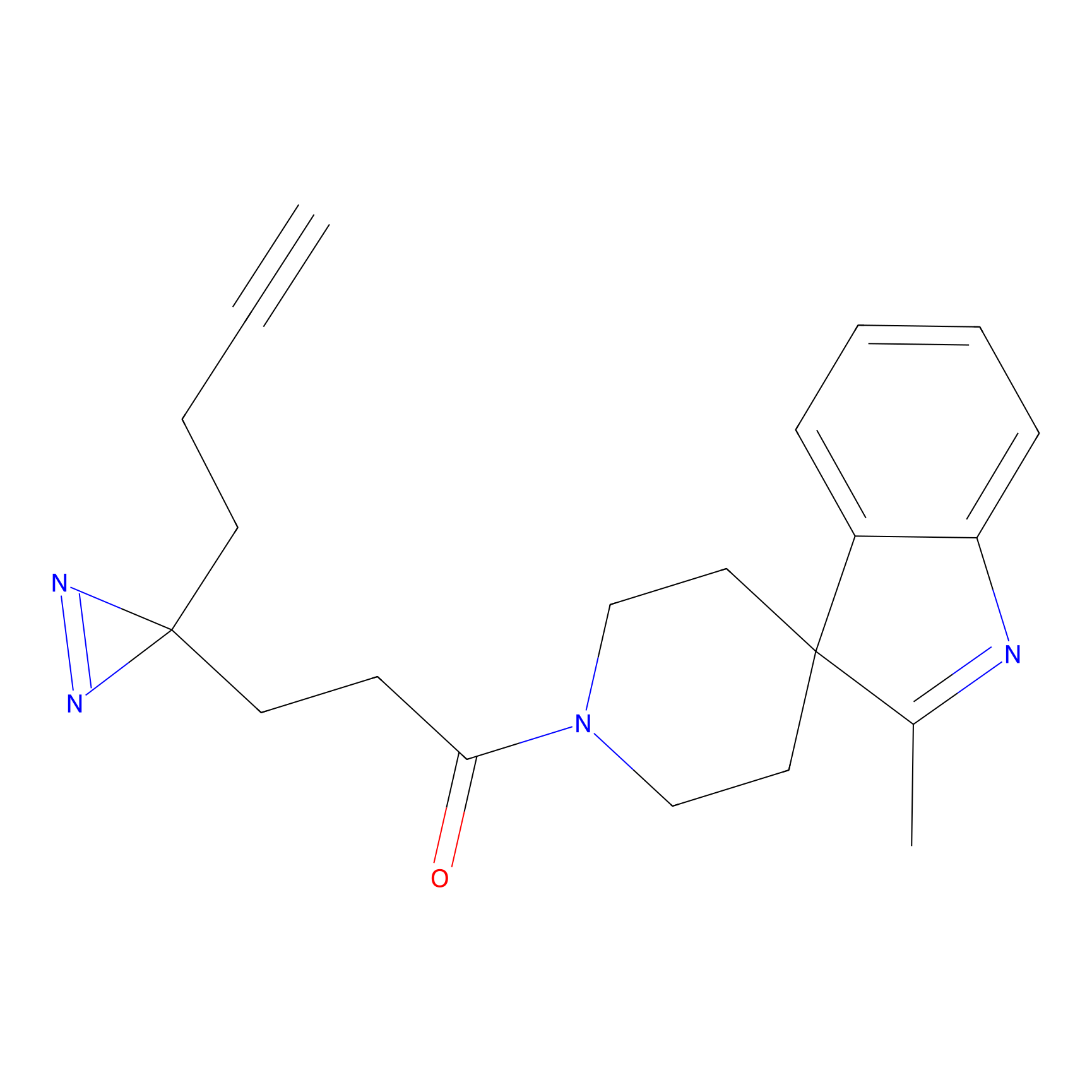 |
7.26 | LDD2066 | [15] | |
|
FFF probe4 Probe Info |
 |
6.61 | LDD0466 | [16] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0137 | [17] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | H24(0.00); H18(0.00) | LDD0222 | [13] |
| LDCM0116 | HHS-0101 | DM93 | Y135(0.77) | LDD0264 | [7] |
| LDCM0117 | HHS-0201 | DM93 | Y135(0.83) | LDD0265 | [7] |
| LDCM0118 | HHS-0301 | DM93 | Y135(1.60) | LDD0266 | [7] |
| LDCM0119 | HHS-0401 | DM93 | Y135(0.95) | LDD0267 | [7] |
| LDCM0120 | HHS-0701 | DM93 | Y135(0.64) | LDD0268 | [7] |
| LDCM0107 | IAA | HeLa | H24(0.00); H18(0.00) | LDD0221 | [13] |
| LDCM0109 | NEM | HeLa | H24(0.00); H18(0.00) | LDD0223 | [13] |
References
