Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Alkylaryl probe 3 Probe Info |
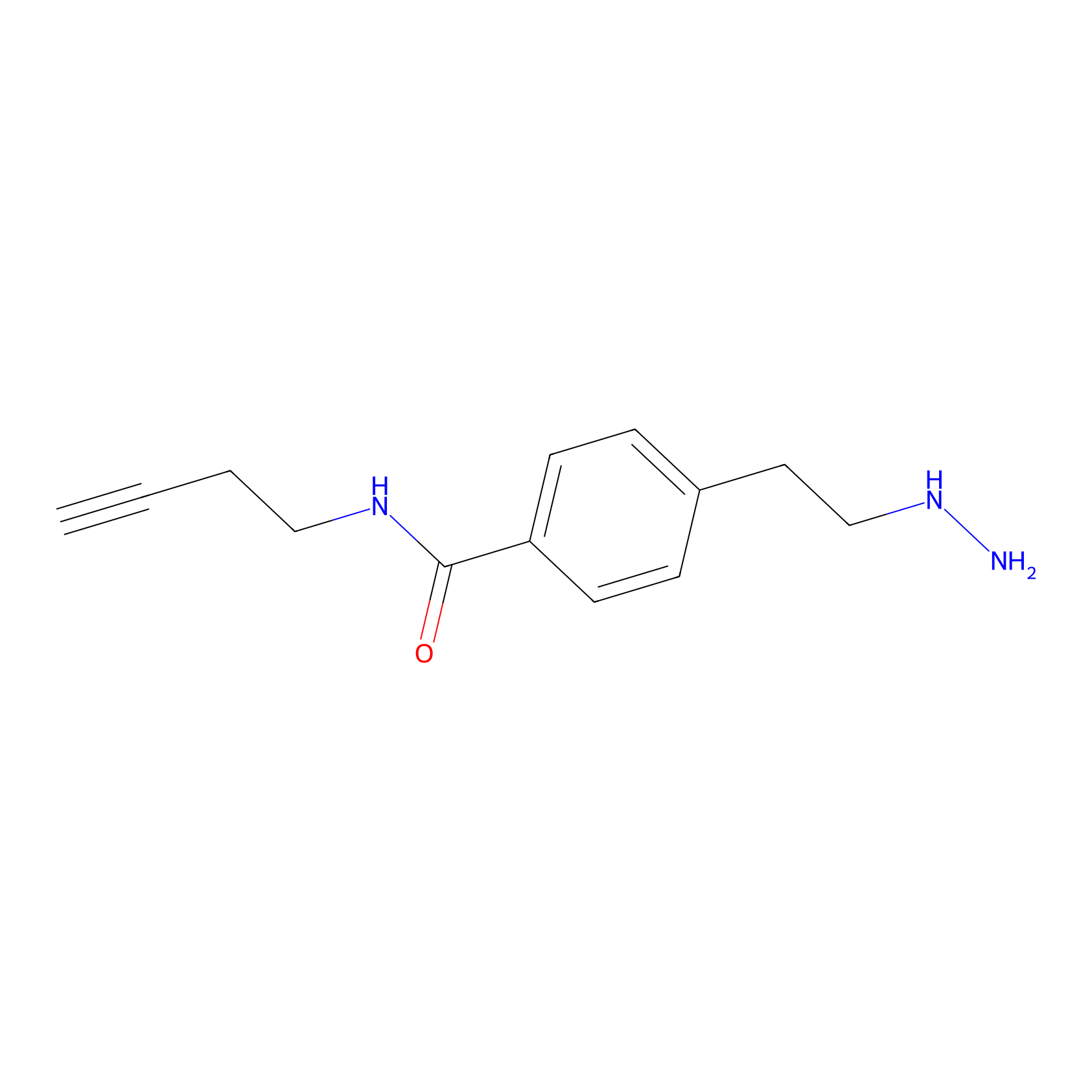 |
20.00 | LDD0381 | [1] | |
|
AZ-9 Probe Info |
 |
4.46 | LDD0393 | [2] | |
|
FBPP2 Probe Info |
 |
25.67 | LDD0318 | [3] | |
|
TH211 Probe Info |
 |
Y48(20.00) | LDD0257 | [4] | |
|
TH216 Probe Info |
 |
Y48(20.00) | LDD0259 | [4] | |
|
STPyne Probe Info |
 |
K173(7.32) | LDD0277 | [5] | |
|
HHS-475 Probe Info |
 |
Y48(1.04) | LDD0264 | [6] | |
|
DBIA Probe Info |
 |
C104(0.72) | LDD1575 | [7] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [8] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [9] | |
|
ATP probe Probe Info |
 |
K23(0.00); K38(0.00) | LDD0035 | [10] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C041 Probe Info |
 |
5.94 | LDD1741 | [11] | |
|
C087 Probe Info |
 |
7.16 | LDD1779 | [11] | |
|
C193 Probe Info |
 |
5.21 | LDD1869 | [11] | |
|
C269 Probe Info |
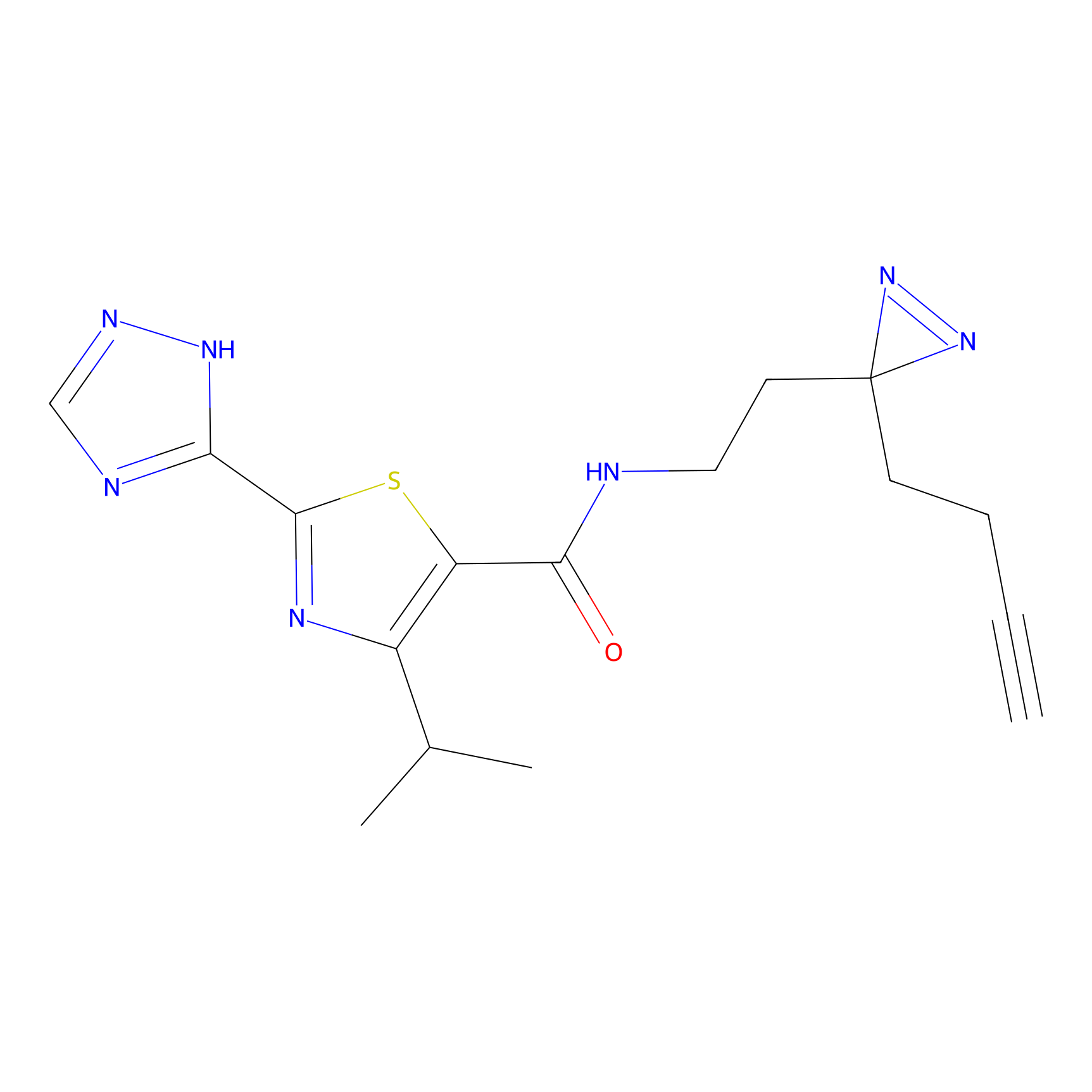 |
6.50 | LDD1939 | [11] | |
|
C390 Probe Info |
 |
43.41 | LDD2049 | [11] | |
|
C391 Probe Info |
 |
24.59 | LDD2050 | [11] | |
|
Kambe_31 Probe Info |
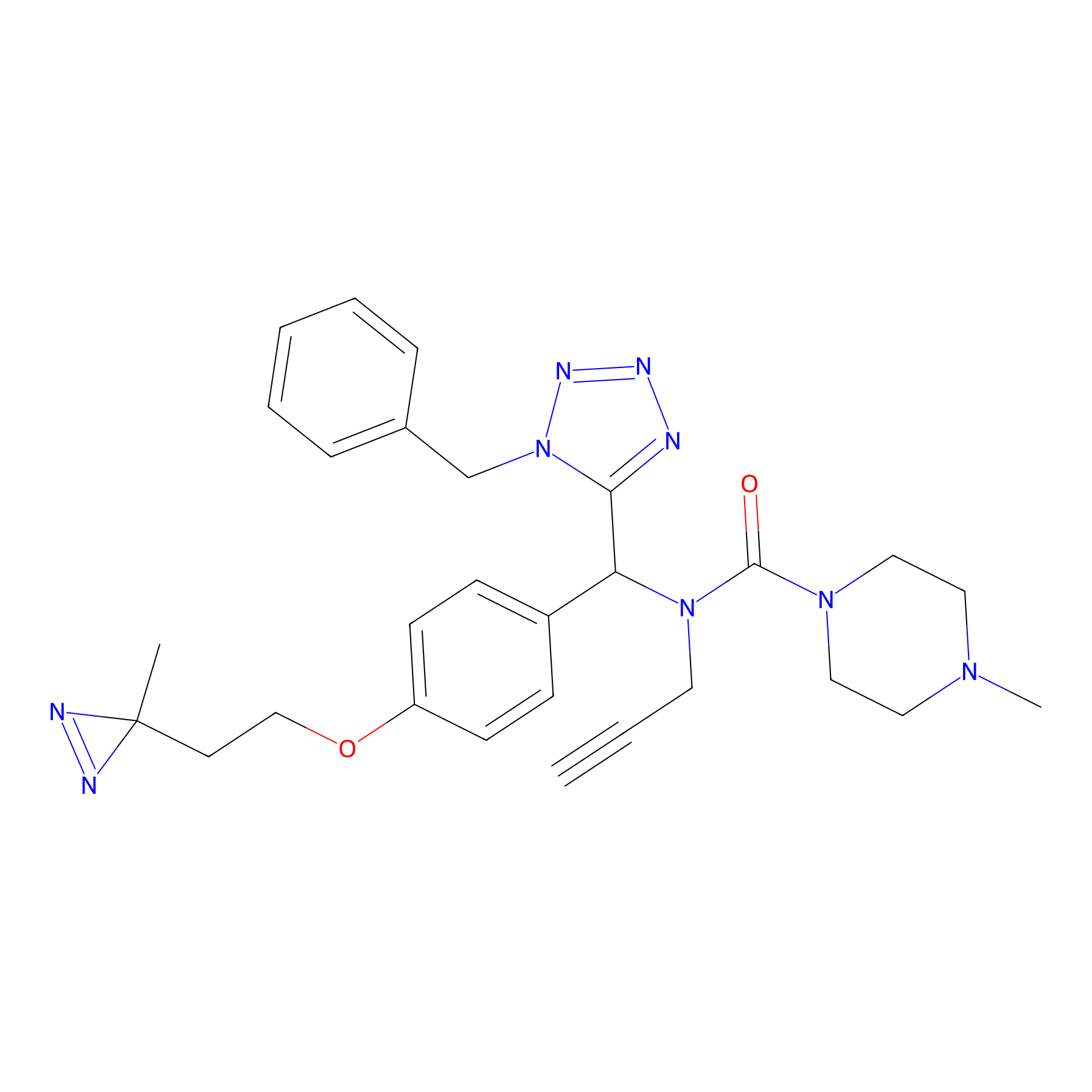 |
9.36 | LDD0128 | [12] | |
|
Kambe_8 Probe Info |
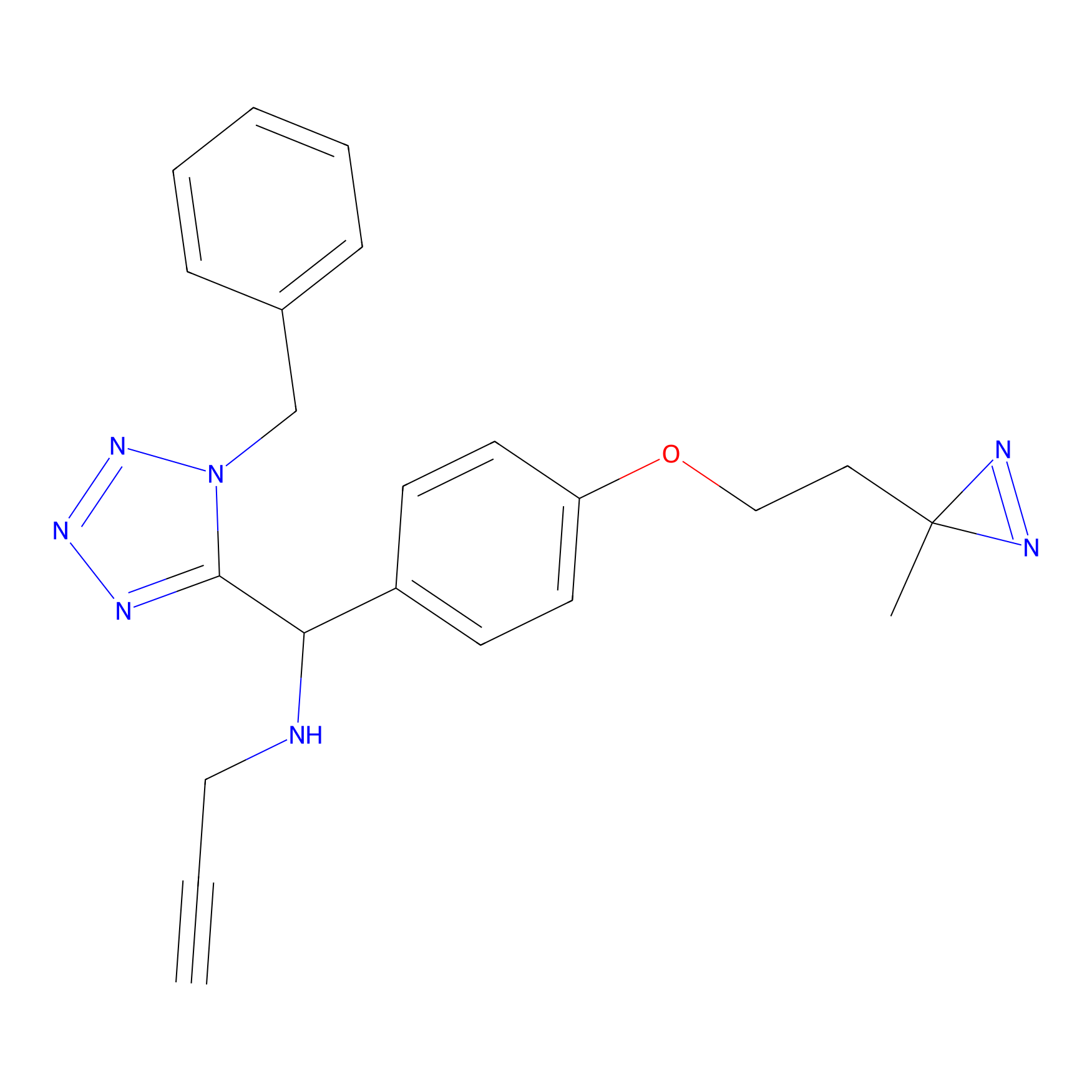 |
2.13 | LDD0131 | [12] | |
|
OEA-DA Probe Info |
 |
20.00 | LDD0046 | [13] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0371 | CL102 | HEK-293T | C104(0.72) | LDD1575 | [7] |
| LDCM0375 | CL106 | HEK-293T | C104(1.09) | LDD1579 | [7] |
| LDCM0380 | CL110 | HEK-293T | C104(0.71) | LDD1584 | [7] |
| LDCM0384 | CL114 | HEK-293T | C104(0.63) | LDD1588 | [7] |
| LDCM0388 | CL118 | HEK-293T | C104(0.86) | LDD1592 | [7] |
| LDCM0393 | CL122 | HEK-293T | C104(0.91) | LDD1597 | [7] |
| LDCM0397 | CL126 | HEK-293T | C104(0.79) | LDD1601 | [7] |
| LDCM0401 | CL14 | HEK-293T | C104(1.04) | LDD1605 | [7] |
| LDCM0407 | CL2 | HEK-293T | C104(1.19) | LDD1611 | [7] |
| LDCM0414 | CL26 | HEK-293T | C104(1.05) | LDD1618 | [7] |
| LDCM0441 | CL50 | HEK-293T | C104(0.81) | LDD1645 | [7] |
| LDCM0454 | CL62 | HEK-293T | C104(0.78) | LDD1657 | [7] |
| LDCM0467 | CL74 | HEK-293T | C104(0.69) | LDD1670 | [7] |
| LDCM0480 | CL86 | HEK-293T | C104(0.56) | LDD1683 | [7] |
| LDCM0493 | CL98 | HEK-293T | C104(0.91) | LDD1696 | [7] |
| LDCM0427 | Fragment51 | HEK-293T | C104(1.11) | LDD1631 | [7] |
| LDCM0116 | HHS-0101 | DM93 | Y48(1.04) | LDD0264 | [6] |
| LDCM0117 | HHS-0201 | DM93 | Y48(0.71) | LDD0265 | [6] |
| LDCM0118 | HHS-0301 | DM93 | Y48(0.74) | LDD0266 | [6] |
| LDCM0119 | HHS-0401 | DM93 | Y48(0.78) | LDD0267 | [6] |
| LDCM0120 | HHS-0701 | DM93 | Y48(1.44) | LDD0268 | [6] |
| LDCM0073 | Kambe_cp3 | PC-3 | 9.36 | LDD0128 | [12] |
| LDCM0075 | kambe_cp62 | PC-3 | 2.13 | LDD0131 | [12] |
| LDCM0079 | kambe_cp64 | PC-3 | 2.30 | LDD0135 | [12] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Huntingtin (HTT) | Huntingtin family | P42858 | |||
| Huntingtin-interacting protein 1 (HIP1) | SLA2 family | O00291 | |||
| Wolframin (WFS1) | . | O76024 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Heat shock protein beta-1 (HSPB1) | Small heat shock protein (HSP20) family | P04792 | |||
| RING finger protein 11 (RNF11) | . | Q9Y3C5 | |||
References
