Details of the Target
General Information of Target
| Target ID | LDTP15132 | |||||
|---|---|---|---|---|---|---|
| Target Name | 2-oxoglutarate and iron-dependent oxygenase domain-containing protein 3 (OGFOD3) | |||||
| Gene Name | OGFOD3 | |||||
| Gene ID | 79701 | |||||
| Synonyms |
C17orf101; 2-oxoglutarate and iron-dependent oxygenase domain-containing protein 3; EC 1.14.11.- |
|||||
| 3D Structure | ||||||
| Sequence |
MDLKLKDCEFWYSLHGQVPGLLDWDMRNELFLPCTTDQCSLAEQILAKYRVGVMKPPEMP
QKRRPSPDGDGPPCEPNLWMWVDPNILCPLGSQEAPKPSGKEDLTNISPFPQPPQKDEGS NCSEDKVVESLPSSSSEQSPLQKQGIHSPSDFELTEEEAEEPDDNSLQSPEMKCYQSQKL WQINNQEKSWQRPPLNCSHLIALALRNNPHCGLSVQEIYNFTRQHFPFFWTAPDGWKSTI HYNLCFLDSFEKVPDSLKDEDNARPRSCLWKLTKEGHRRFWEETRVLAFAQRERIQECMS QPELLTSLFDL |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
OGFOD3 family
|
|||||
| Subcellular location |
Membrane
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| A431 | SNV: p.T118I | . | |||
| ETK1 | SNV: p.D310Y | . | |||
| MOLT4 | SNV: p.R185W | IA-alkyne Probe Info | |||
| NCIH1048 | SNV: p.I303V | . | |||
| RKO | SNV: p.V184M | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DBIA Probe Info |
 |
C103(1.57) | LDD3333 | [1] | |
|
IA-alkyne Probe Info |
 |
C103(0.00); C90(0.00) | LDD0162 | [2] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [3] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C056 Probe Info |
 |
19.03 | LDD1753 | [4] | |
|
C161 Probe Info |
 |
11.47 | LDD1841 | [4] | |
|
C169 Probe Info |
 |
85.04 | LDD1849 | [4] | |
|
C236 Probe Info |
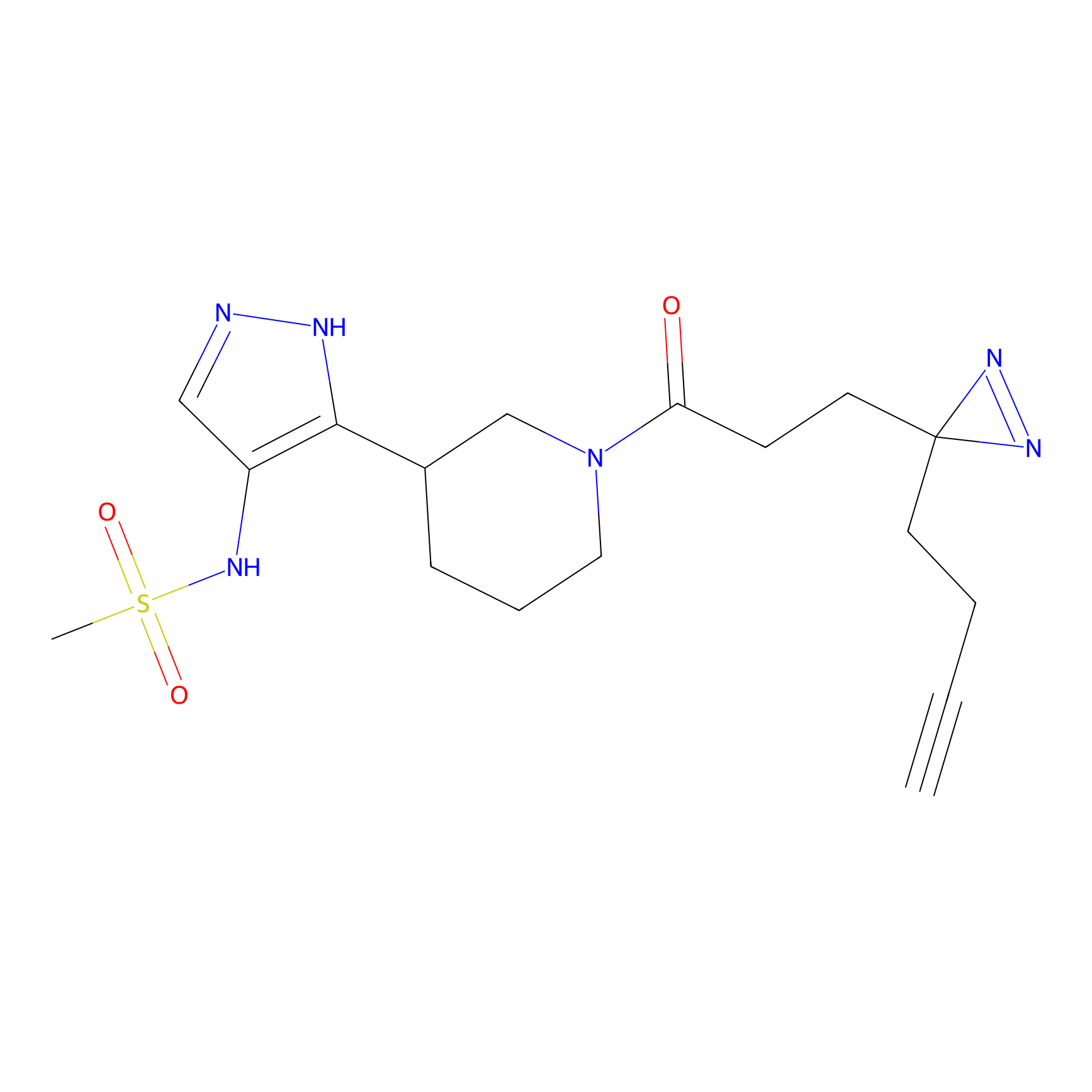 |
34.30 | LDD1909 | [4] | |
|
C238 Probe Info |
 |
17.03 | LDD1911 | [4] | |
|
C240 Probe Info |
 |
8.22 | LDD1913 | [4] | |
|
C251 Probe Info |
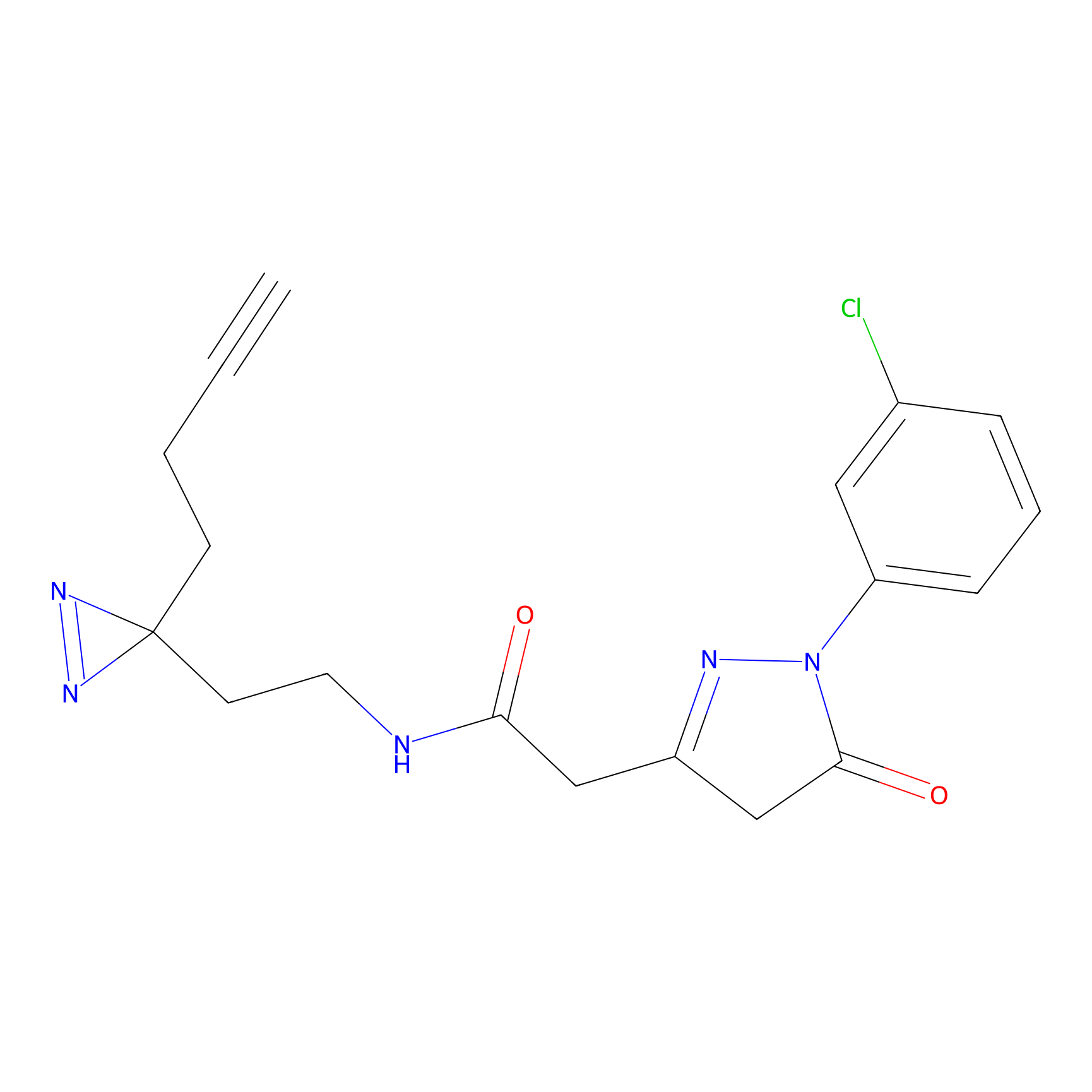 |
27.47 | LDD1924 | [4] | |
|
C269 Probe Info |
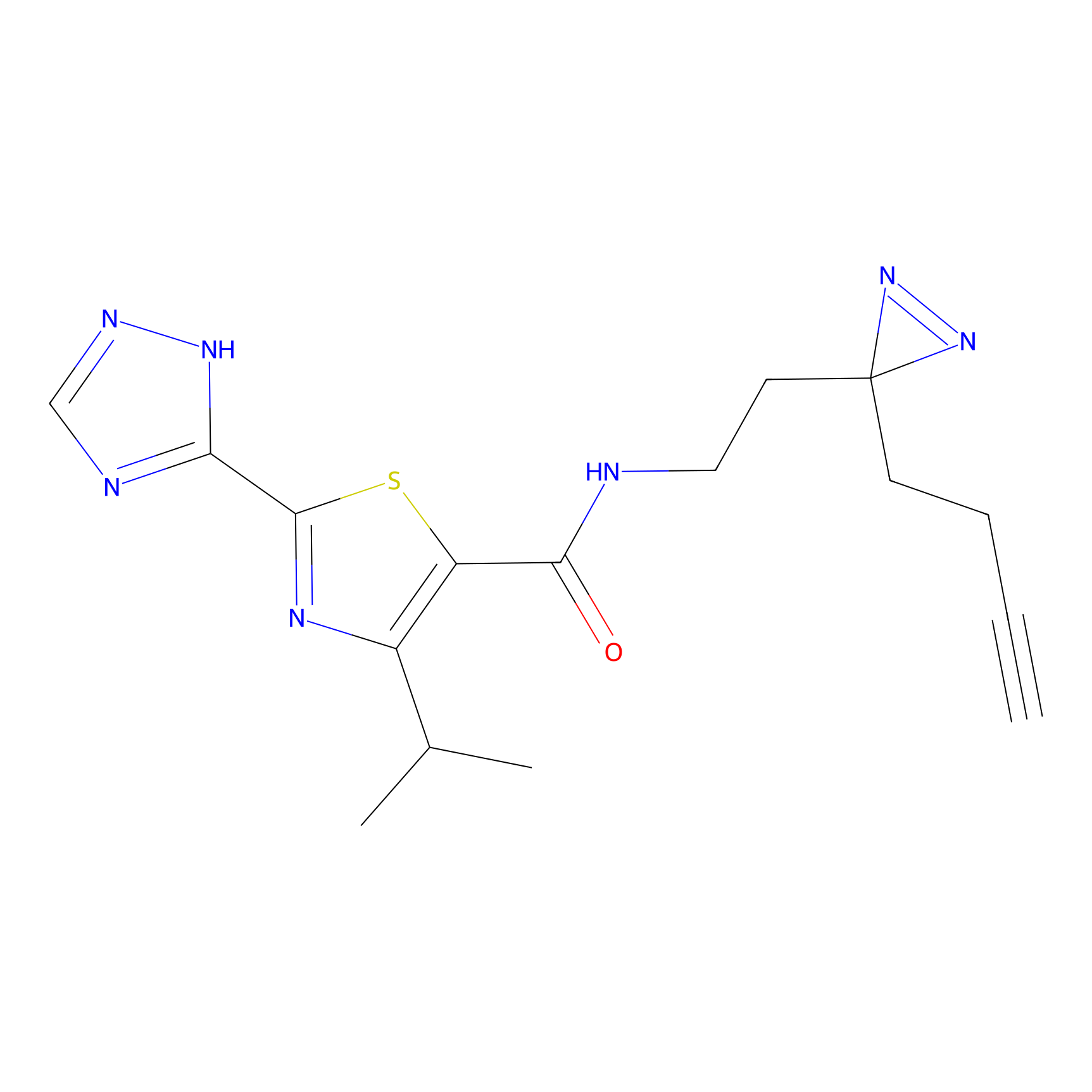 |
6.15 | LDD1939 | [4] | |
|
C388 Probe Info |
 |
64.89 | LDD2047 | [4] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C103(1.06) | LDD1507 | [5] |
| LDCM0215 | AC10 | HEK-293T | C103(1.02) | LDD1508 | [5] |
| LDCM0226 | AC11 | HEK-293T | C103(1.05) | LDD1509 | [5] |
| LDCM0237 | AC12 | HEK-293T | C103(1.06) | LDD1510 | [5] |
| LDCM0276 | AC17 | HEK-293T | C103(1.10) | LDD1515 | [5] |
| LDCM0277 | AC18 | HEK-293T | C103(1.03) | LDD1516 | [5] |
| LDCM0278 | AC19 | HEK-293T | C103(1.23) | LDD1517 | [5] |
| LDCM0279 | AC2 | HEK-293T | C103(1.17) | LDD1518 | [5] |
| LDCM0280 | AC20 | HEK-293T | C103(1.07) | LDD1519 | [5] |
| LDCM0285 | AC25 | HEK-293T | C103(0.95) | LDD1524 | [5] |
| LDCM0286 | AC26 | HEK-293T | C103(1.18) | LDD1525 | [5] |
| LDCM0287 | AC27 | HEK-293T | C103(1.04) | LDD1526 | [5] |
| LDCM0288 | AC28 | HEK-293T | C103(1.02) | LDD1527 | [5] |
| LDCM0290 | AC3 | HEK-293T | C103(1.02) | LDD1529 | [5] |
| LDCM0294 | AC33 | HEK-293T | C103(1.06) | LDD1533 | [5] |
| LDCM0295 | AC34 | HEK-293T | C103(1.06) | LDD1534 | [5] |
| LDCM0296 | AC35 | HEK-293T | C103(1.14) | LDD1535 | [5] |
| LDCM0297 | AC36 | HEK-293T | C103(1.03) | LDD1536 | [5] |
| LDCM0301 | AC4 | HEK-293T | C103(1.01) | LDD1540 | [5] |
| LDCM0303 | AC41 | HEK-293T | C103(1.04) | LDD1542 | [5] |
| LDCM0304 | AC42 | HEK-293T | C103(1.00) | LDD1543 | [5] |
| LDCM0305 | AC43 | HEK-293T | C103(1.07) | LDD1544 | [5] |
| LDCM0306 | AC44 | HEK-293T | C103(1.10) | LDD1545 | [5] |
| LDCM0311 | AC49 | HEK-293T | C103(1.22) | LDD1550 | [5] |
| LDCM0313 | AC50 | HEK-293T | C103(1.09) | LDD1552 | [5] |
| LDCM0314 | AC51 | HEK-293T | C103(1.05) | LDD1553 | [5] |
| LDCM0315 | AC52 | HEK-293T | C103(1.12) | LDD1554 | [5] |
| LDCM0320 | AC57 | HEK-293T | C103(1.03) | LDD1559 | [5] |
| LDCM0321 | AC58 | HEK-293T | C103(1.12) | LDD1560 | [5] |
| LDCM0322 | AC59 | HEK-293T | C103(1.07) | LDD1561 | [5] |
| LDCM0324 | AC60 | HEK-293T | C103(1.04) | LDD1563 | [5] |
| LDCM0356 | AKOS034007680 | HEK-293T | C103(1.05) | LDD1570 | [5] |
| LDCM0371 | CL102 | HEK-293T | C103(1.10) | LDD1575 | [5] |
| LDCM0375 | CL106 | HEK-293T | C103(1.01) | LDD1579 | [5] |
| LDCM0380 | CL110 | HEK-293T | C103(1.09) | LDD1584 | [5] |
| LDCM0384 | CL114 | HEK-293T | C103(1.29) | LDD1588 | [5] |
| LDCM0388 | CL118 | HEK-293T | C103(1.22) | LDD1592 | [5] |
| LDCM0393 | CL122 | HEK-293T | C103(1.21) | LDD1597 | [5] |
| LDCM0397 | CL126 | HEK-293T | C103(1.15) | LDD1601 | [5] |
| LDCM0401 | CL14 | HEK-293T | C103(0.98) | LDD1605 | [5] |
| LDCM0404 | CL17 | HEK-293T | C103(1.19) | LDD1608 | [5] |
| LDCM0405 | CL18 | HEK-293T | C103(0.92) | LDD1609 | [5] |
| LDCM0406 | CL19 | HEK-293T | C103(0.92) | LDD1610 | [5] |
| LDCM0407 | CL2 | HEK-293T | C103(1.18) | LDD1611 | [5] |
| LDCM0408 | CL20 | HEK-293T | C103(1.08) | LDD1612 | [5] |
| LDCM0414 | CL26 | HEK-293T | C103(1.03) | LDD1618 | [5] |
| LDCM0417 | CL29 | HEK-293T | C103(0.93) | LDD1621 | [5] |
| LDCM0419 | CL30 | HEK-293T | C103(1.07) | LDD1623 | [5] |
| LDCM0420 | CL31 | HEK-293T | C103(1.03) | LDD1624 | [5] |
| LDCM0421 | CL32 | HEK-293T | C103(1.04) | LDD1625 | [5] |
| LDCM0431 | CL41 | HEK-293T | C103(1.23) | LDD1635 | [5] |
| LDCM0432 | CL42 | HEK-293T | C103(1.10) | LDD1636 | [5] |
| LDCM0433 | CL43 | HEK-293T | C103(0.95) | LDD1637 | [5] |
| LDCM0434 | CL44 | HEK-293T | C103(1.13) | LDD1638 | [5] |
| LDCM0440 | CL5 | HEK-293T | C103(1.19) | LDD1644 | [5] |
| LDCM0441 | CL50 | HEK-293T | C103(1.17) | LDD1645 | [5] |
| LDCM0444 | CL53 | HEK-293T | C103(1.17) | LDD1647 | [5] |
| LDCM0445 | CL54 | HEK-293T | C103(1.19) | LDD1648 | [5] |
| LDCM0446 | CL55 | HEK-293T | C103(0.94) | LDD1649 | [5] |
| LDCM0447 | CL56 | HEK-293T | C103(1.07) | LDD1650 | [5] |
| LDCM0451 | CL6 | HEK-293T | C103(1.00) | LDD1654 | [5] |
| LDCM0454 | CL62 | HEK-293T | C103(1.00) | LDD1657 | [5] |
| LDCM0457 | CL65 | HEK-293T | C103(1.10) | LDD1660 | [5] |
| LDCM0458 | CL66 | HEK-293T | C103(0.98) | LDD1661 | [5] |
| LDCM0459 | CL67 | HEK-293T | C103(0.93) | LDD1662 | [5] |
| LDCM0460 | CL68 | HEK-293T | C103(1.09) | LDD1663 | [5] |
| LDCM0462 | CL7 | HEK-293T | C103(0.95) | LDD1665 | [5] |
| LDCM0467 | CL74 | HEK-293T | C103(1.26) | LDD1670 | [5] |
| LDCM0470 | CL77 | HEK-293T | C103(1.16) | LDD1673 | [5] |
| LDCM0471 | CL78 | HEK-293T | C103(0.93) | LDD1674 | [5] |
| LDCM0472 | CL79 | HEK-293T | C103(1.02) | LDD1675 | [5] |
| LDCM0473 | CL8 | HEK-293T | C103(0.98) | LDD1676 | [5] |
| LDCM0474 | CL80 | HEK-293T | C103(1.17) | LDD1677 | [5] |
| LDCM0480 | CL86 | HEK-293T | C103(1.07) | LDD1683 | [5] |
| LDCM0483 | CL89 | HEK-293T | C103(1.02) | LDD1686 | [5] |
| LDCM0485 | CL90 | HEK-293T | C103(1.19) | LDD1688 | [5] |
| LDCM0486 | CL91 | HEK-293T | C103(1.01) | LDD1689 | [5] |
| LDCM0487 | CL92 | HEK-293T | C103(1.18) | LDD1690 | [5] |
| LDCM0493 | CL98 | HEK-293T | C103(1.10) | LDD1696 | [5] |
| LDCM0427 | Fragment51 | HEK-293T | C103(1.09) | LDD1631 | [5] |
| LDCM0022 | KB02 | Hs 936.T | C103(1.54) | LDD2363 | [1] |
| LDCM0023 | KB03 | Hs 936.T | C103(2.18) | LDD2780 | [1] |
| LDCM0024 | KB05 | MOLM-13 | C103(1.57) | LDD3333 | [1] |
References
