Details of the Target
General Information of Target
| Target ID | LDTP03976 | |||||
|---|---|---|---|---|---|---|
| Target Name | Aldo-keto reductase family 1 member C3 (AKR1C3) | |||||
| Gene Name | AKR1C3 | |||||
| Gene ID | 8644 | |||||
| Synonyms |
DDH1; HSD17B5; KIAA0119; PGFS; Aldo-keto reductase family 1 member C3; EC 1.1.1.-; EC 1.1.1.210; EC 1.1.1.53; EC 1.1.1.62; 17-beta-hydroxysteroid dehydrogenase type 5; 17-beta-HSD 5; 3-alpha-HSD type II, brain; 3-alpha-hydroxysteroid dehydrogenase type 2; 3-alpha-HSD type 2; EC 1.1.1.357; Chlordecone reductase homolog HAKRb; Dihydrodiol dehydrogenase 3; DD-3; DD3; Dihydrodiol dehydrogenase type I; HA1753; Prostaglandin F synthase; PGFS; EC 1.1.1.188; Testosterone 17-beta-dehydrogenase 5; EC 1.1.1.239, EC 1.1.1.64
|
|||||
| 3D Structure | ||||||
| Sequence |
MDSKHQCVKLNDGHFMPVLGFGTYAPPEVPRSKALEVTKLAIEAGFRHIDSAHLYNNEEQ
VGLAIRSKIADGSVKREDIFYTSKLWSTFHRPELVRPALENSLKKAQLDYVDLYLIHSPM SLKPGEELSPTDENGKVIFDIVDLCTTWEAMEKCKDAGLAKSIGVSNFNRRQLEMILNKP GLKYKPVCNQVECHPYFNRSKLLDFCKSKDIVLVAYSALGSQRDKRWVDPNSPVLLEDPV LCALAKKHKRTPALIALRYQLQRGVVVLAKSYNEQRIRQNVQVFEFQLTAEDMKAIDGLD RNLHYFNSDSFASHPNYPYSDEY |
|||||
| Target Type |
Successful
|
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Aldo/keto reductase family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Cytosolic aldo-keto reductase that catalyzes the NADH and NADPH-dependent reduction of ketosteroids to hydroxysteroids. Acts as a NAD(P)(H)-dependent 3-, 17- and 20-ketosteroid reductase on the steroid nucleus and side chain and regulates the metabolism of androgens, estrogens and progesterone. Displays the ability to catalyze both oxidation and reduction in vitro, but most probably acts as a reductase in vivo since the oxidase activity measured in vitro is inhibited by physiological concentration of NADPH. Acts preferentially as a 17-ketosteroid reductase and has the highest catalytic efficiency of the AKR1C enzyme for the reduction of delta4-androstenedione to form testosterone. Reduces prostaglandin (PG) D2 to 11beta-prostaglandin F2, progesterone to 20alpha-hydroxyprogesterone and estrone to 17beta-estradiol . Catalyzes the transformation of the potent androgen dihydrotestosterone (DHT) into the less active form, 5-alpha-androstan-3-alpha,17-beta-diol (3-alpha-diol) . Also displays retinaldehyde reductase activity toward 9-cis-retinal.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| C32 | SNV: p.R223Q | . | |||
| ETK1 | SNV: p.C206R | . | |||
| HEPG2 | SNV: p.I69T | NAIA_5 Probe Info | |||
| MFE319 | Insertion: p.K105EfsTer8 | DBIA Probe Info | |||
| NCIH1703 | SNV: p.N198K | DBIA Probe Info | |||
| RKO | SNV: p.A269G | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P1 Probe Info |
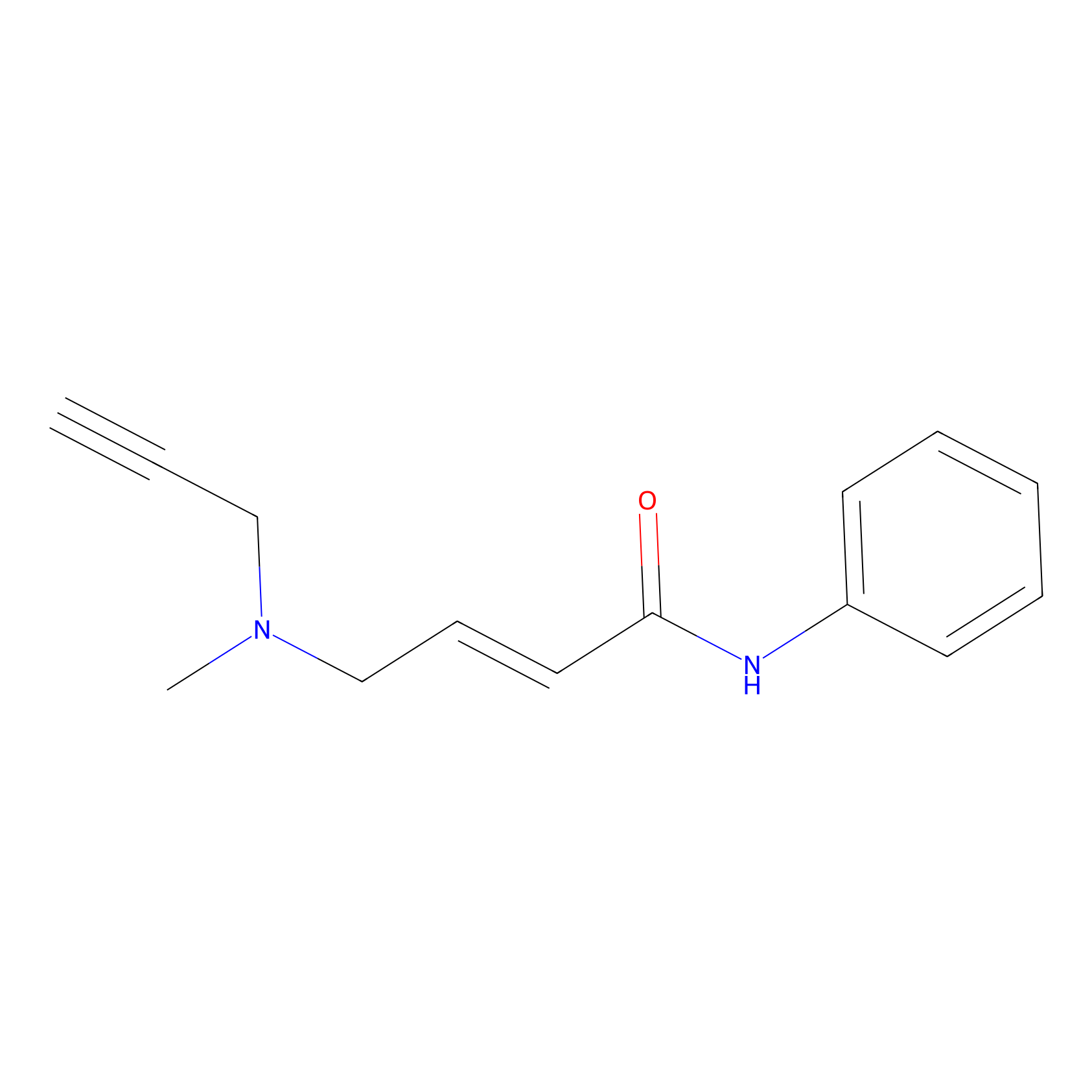 |
2.95 | LDD0452 | [1] | |
|
P2 Probe Info |
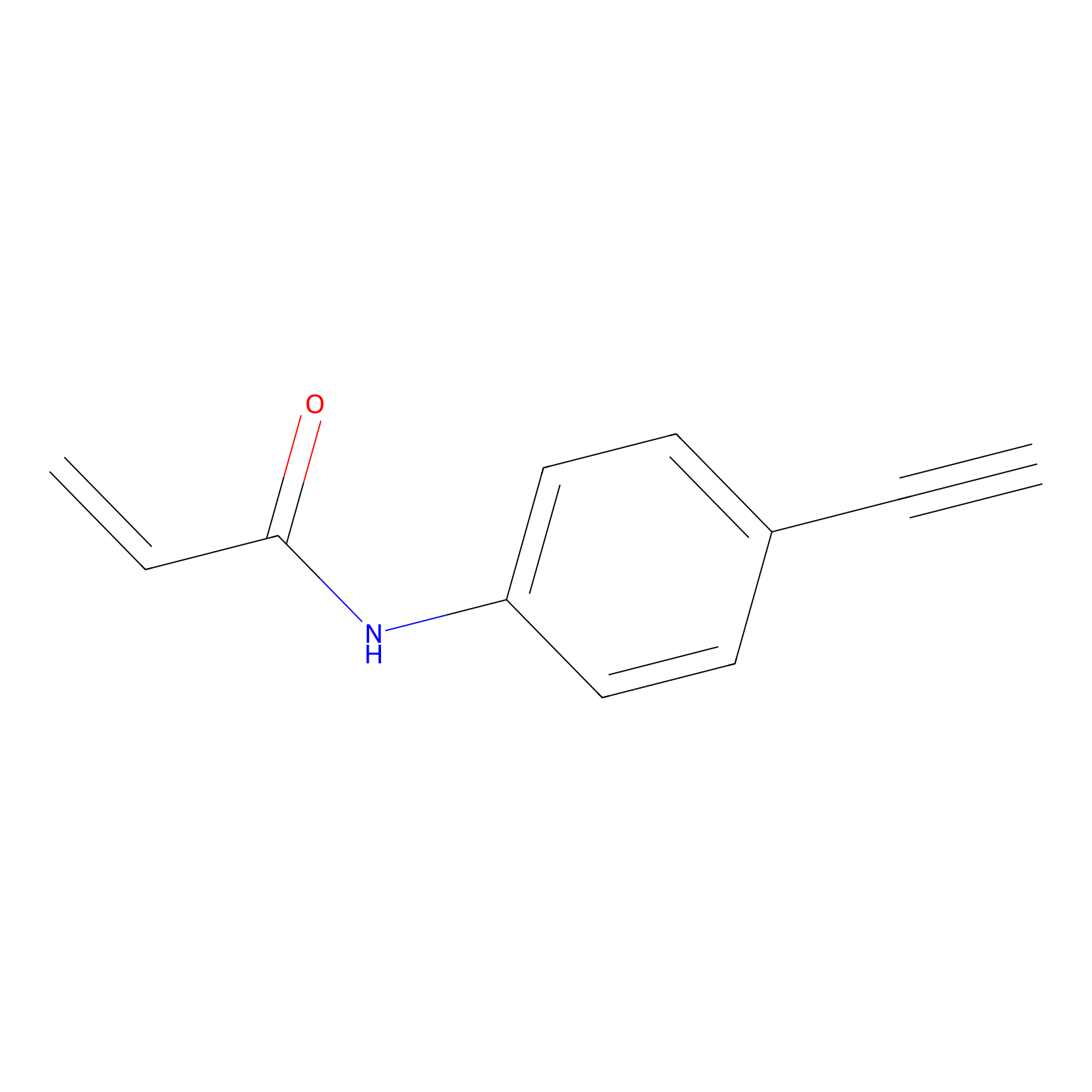 |
1.51 | LDD0453 | [1] | |
|
P8 Probe Info |
 |
2.86 | LDD0455 | [1] | |
|
STPyne Probe Info |
 |
K185(5.26); K201(0.38); K209(8.97); K33(6.92) | LDD0277 | [2] | |
|
OPA-S-S-alkyne Probe Info |
 |
K161(2.19) | LDD3494 | [3] | |
|
DBIA Probe Info |
 |
C26(1.06) | LDD3314 | [4] | |
|
W1 Probe Info |
 |
Q172(1.98); K209(2.12); S208(2.12) | LDD0237 | [5] | |
|
m-APA Probe Info |
 |
N.A. | LDD2231 | [6] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C145(0.00); C242(0.00); C193(0.00); C188(0.00) | LDD0038 | [7] | |
|
IA-alkyne Probe Info |
 |
C206(0.00); C242(0.00); C193(0.00); C188(0.00) | LDD0036 | [7] | |
|
Lodoacetamide azide Probe Info |
 |
C206(0.00); C145(0.00); C242(0.00); C193(0.00) | LDD0037 | [7] | |
|
BTD Probe Info |
 |
N.A. | LDD0004 | [8] | |
|
IPM Probe Info |
 |
C193(0.00); C188(0.00); C242(0.00) | LDD0025 | [9] | |
|
JW-RF-010 Probe Info |
 |
C188(0.00); C242(0.00); C193(0.00) | LDD0026 | [9] | |
|
TFBX Probe Info |
 |
C193(0.00); C206(0.00) | LDD0027 | [9] | |
|
WYneN Probe Info |
 |
C242(0.00); C193(0.00) | LDD0021 | [8] | |
|
Acrolein Probe Info |
 |
H304(0.00); H314(0.00) | LDD0217 | [10] | |
|
NAIA_5 Probe Info |
 |
C206(0.00); C145(0.00); C242(0.00); C7(0.00) | LDD2223 | [11] | |
|
HHS-465 Probe Info |
 |
N.A. | LDD2240 | [12] | |
|
HHS-475 Probe Info |
 |
Y184(1.30) | LDD2238 | [13] | |
|
HHS-482 Probe Info |
 |
Y110(0.80); Y184(1.25); Y216(1.25); Y24(1.22) | LDD2239 | [13] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
A-DA Probe Info |
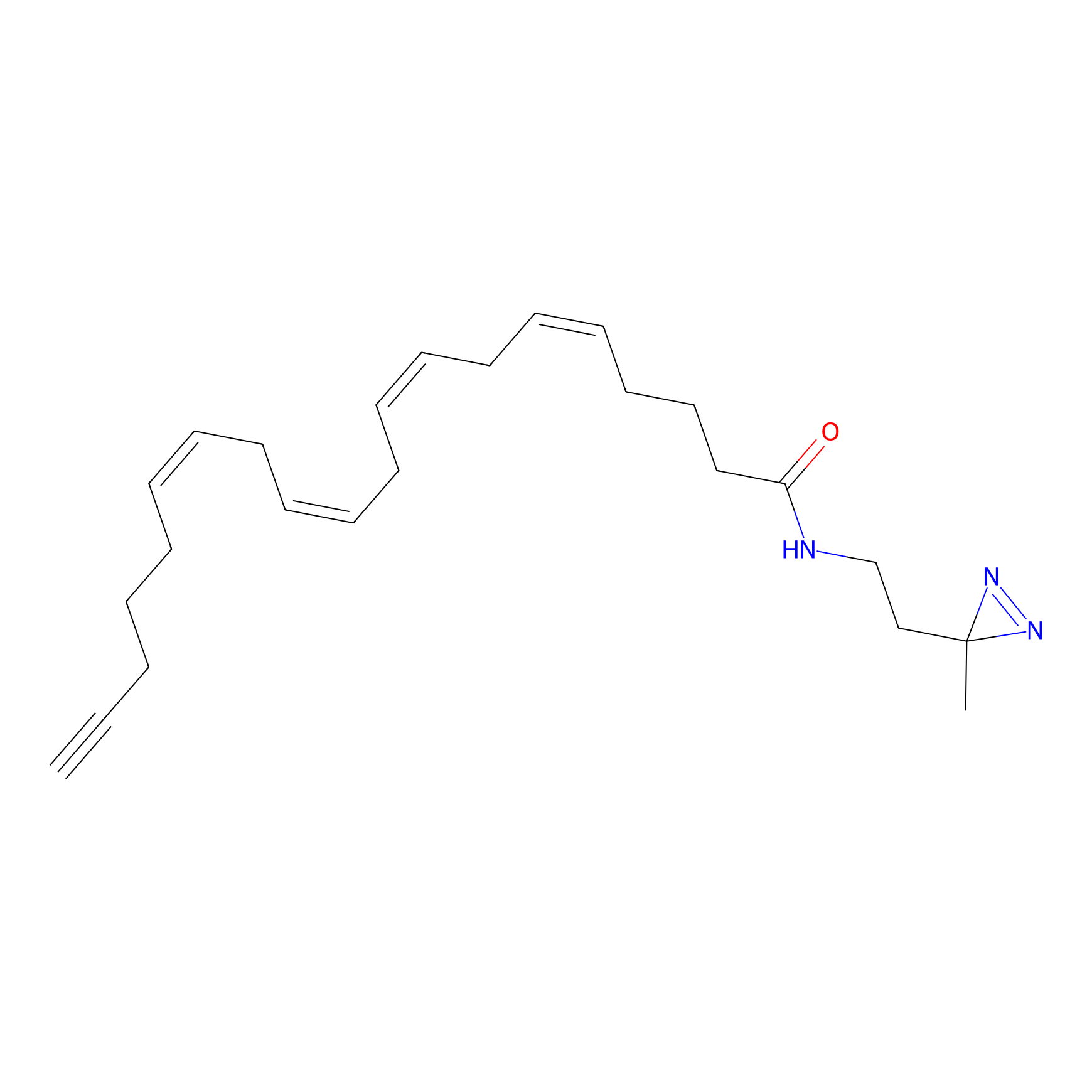 |
2.17 | LDD0140 | [14] | |
|
AEA-DA Probe Info |
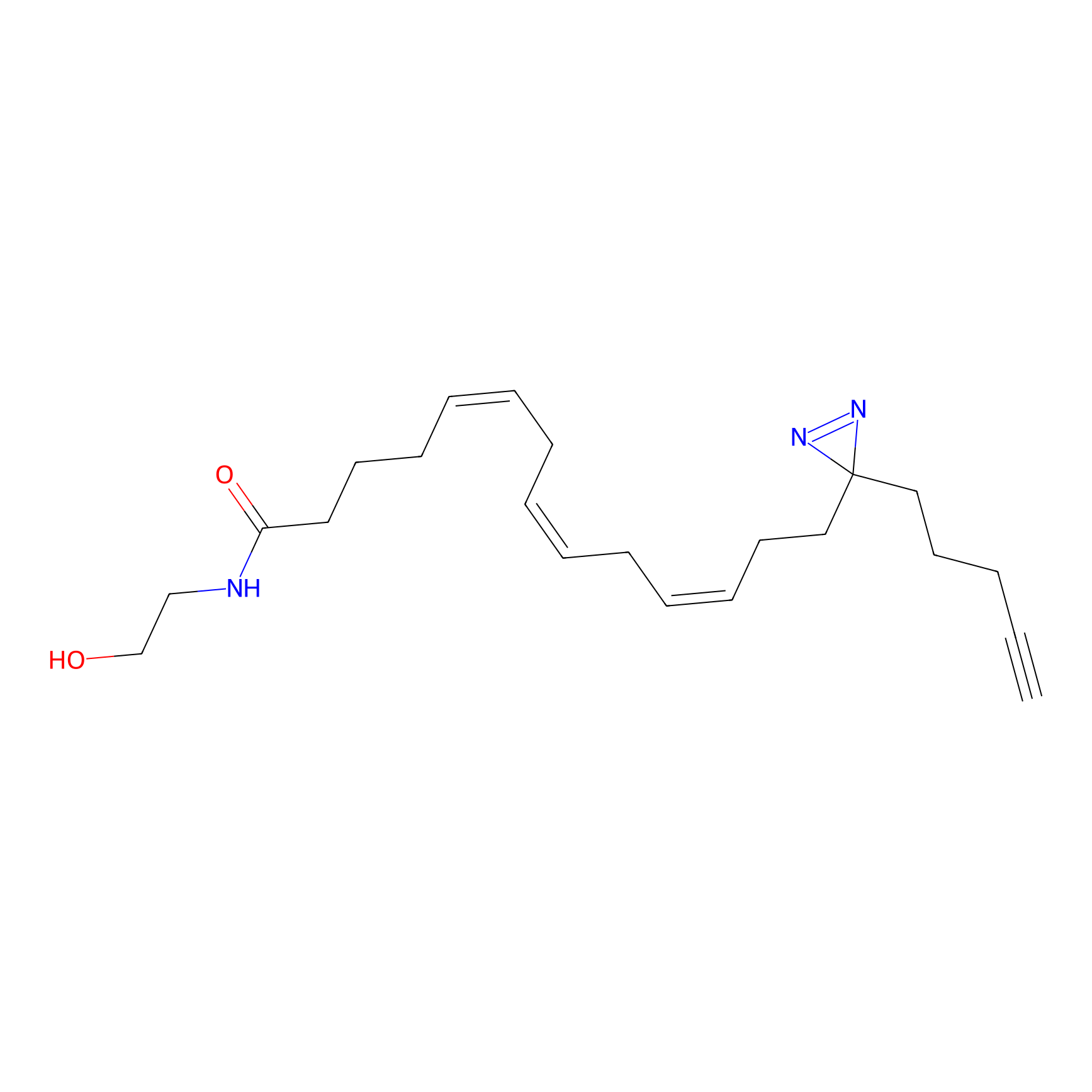 |
6.02 | LDD0146 | [14] | |
|
Kambe_3 Probe Info |
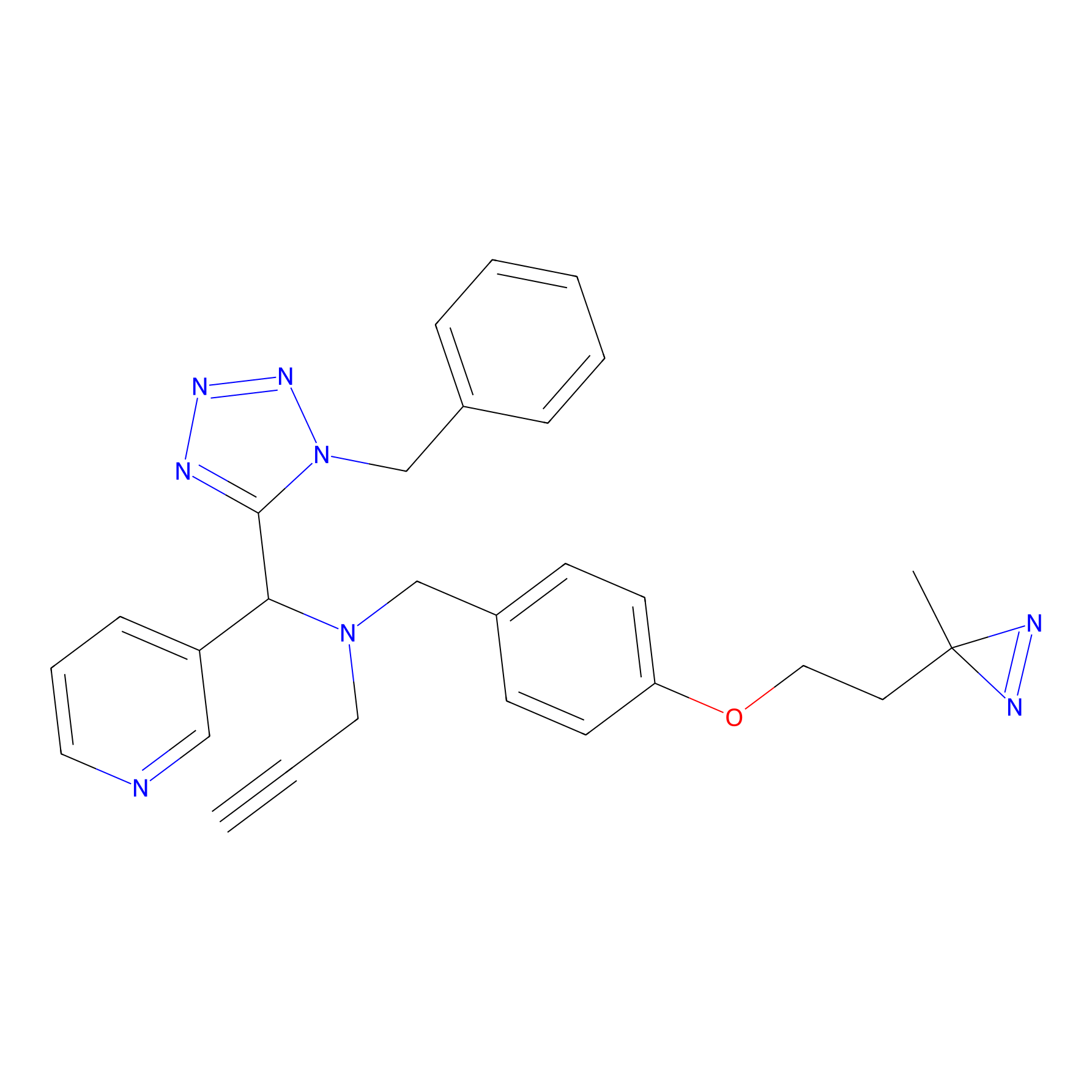 |
20.00 | LDD0130 | [15] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C188(0.79); C193(0.74) | LDD2117 | [16] |
| LDCM0083 | Avasimibe | A-549 | 2.77 | LDD0143 | [14] |
| LDCM0108 | Chloroacetamide | HeLa | H304(0.00); C188(0.00) | LDD0222 | [10] |
| LDCM0632 | CL-Sc | Hep-G2 | C242(3.65); C7(0.63) | LDD2227 | [11] |
| LDCM0082 | FK866 | A-549 | 3.25 | LDD0142 | [14] |
| LDCM0080 | Flurbiprofen | A-549 | 2.17 | LDD0140 | [14] |
| LDCM0107 | IAA | HeLa | H304(0.00); H314(0.00) | LDD0221 | [10] |
| LDCM0074 | Kambe_cp66 | PC-3 | 20.00 | LDD0130 | [15] |
| LDCM0022 | KB02 | 769-P | C242(0.92); C26(0.87) | LDD2246 | [4] |
| LDCM0023 | KB03 | 42-MG-BA | C26(1.29) | LDD2661 | [4] |
| LDCM0024 | KB05 | IGR37 | C26(1.06) | LDD3314 | [4] |
| LDCM0109 | NEM | HeLa | H304(0.00); H48(0.00); H53(0.00); H314(0.00) | LDD0223 | [10] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C188(0.85) | LDD2099 | [16] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C188(1.04) | LDD2107 | [16] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C193(0.83) | LDD2109 | [16] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C193(0.76) | LDD2119 | [16] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C188(1.12); C193(0.95) | LDD2123 | [16] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C193(0.74) | LDD2125 | [16] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C193(1.29) | LDD2136 | [16] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C193(1.97) | LDD2144 | [16] |
| LDCM0081 | Rofecoxib | A-549 | 2.17 | LDD0141 | [14] |
| LDCM0110 | W12 | Hep-G2 | Q172(1.98); K209(2.12); S208(2.12) | LDD0237 | [5] |
| LDCM0111 | W14 | Hep-G2 | S162(0.91); Q271(0.93); R51(1.02); Q208(1.06) | LDD0238 | [5] |
| LDCM0112 | W16 | Hep-G2 | K270(0.96); S271(0.96); K209(1.18); S208(1.22) | LDD0239 | [5] |
| LDCM0113 | W17 | Hep-G2 | K209(1.24); S208(1.24); E174(1.30); Q172(1.30) | LDD0240 | [5] |
The Interaction Atlas With This Target
The Drug(s) Related To This Target
Approved
Phase 2
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Asp-9521 | . | D0H7XJ | |||
Investigative
Patented
Discontinued
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Androstanedione | Small molecular drug | DB01561 | |||
| Androstenedione | Small molecular drug | DB01536 | |||
| Stanolone | Small molecular drug | DB02901 | |||
References
