Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
ONAyne Probe Info |
 |
K232(9.19) | LDD0274 | [1] | |
|
STPyne Probe Info |
 |
K18(9.09); K199(0.69); K210(6.67) | LDD0277 | [1] | |
|
Probe 1 Probe Info |
 |
Y133(13.74); Y233(11.22) | LDD3495 | [2] | |
|
D5yne Probe Info |
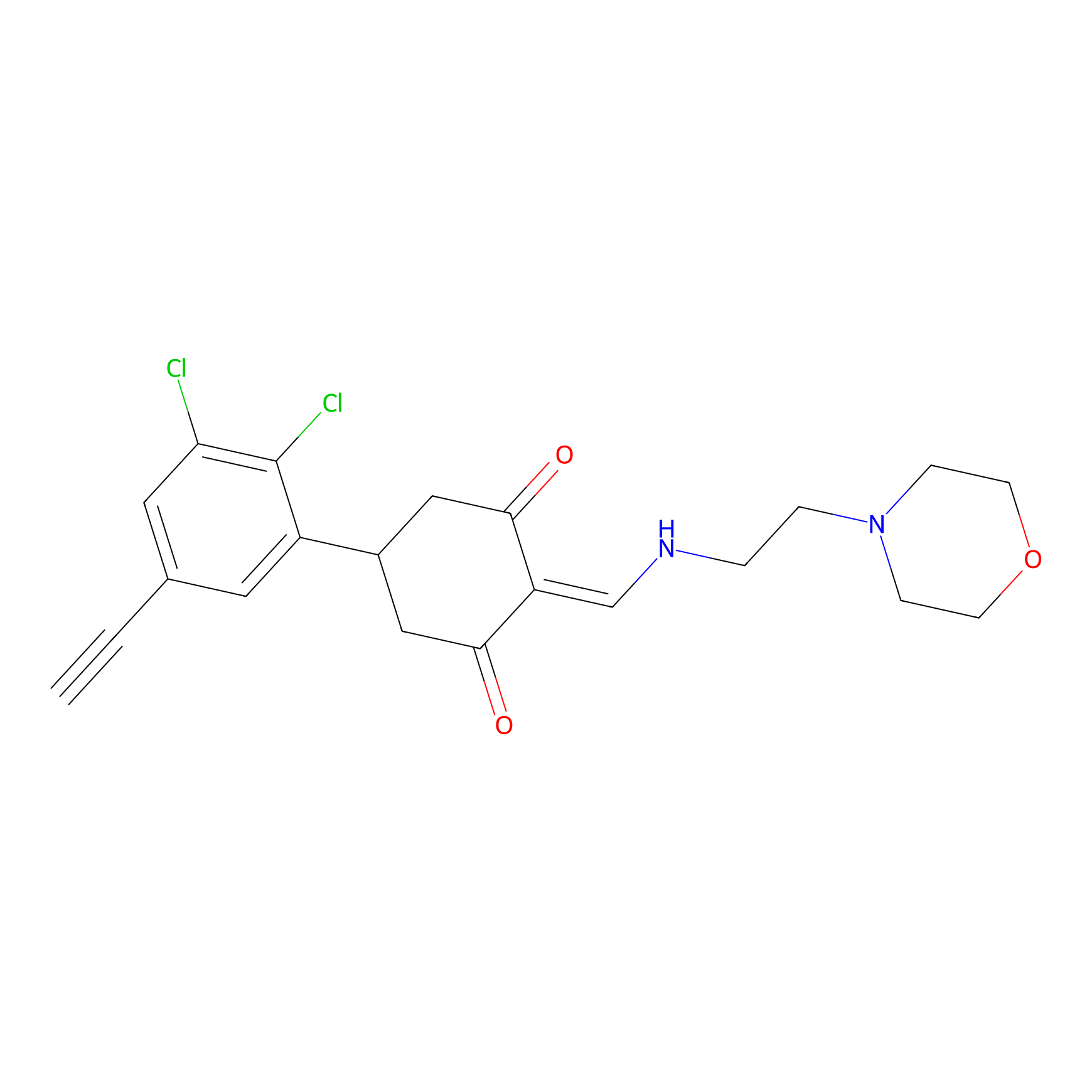 |
1.52 | LDD0055 | [3] | |
|
Jackson_14 Probe Info |
 |
2.13 | LDD0123 | [4] | |
|
ATP probe Probe Info |
 |
K232(0.00); K210(0.00) | LDD0199 | [5] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [6] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [7] | |
|
Ox-W18 Probe Info |
 |
W124(0.00); W151(0.00) | LDD2175 | [8] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [9] | |
|
Acrolein Probe Info |
 |
H215(0.00); H298(0.00) | LDD0217 | [10] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C040 Probe Info |
 |
7.31 | LDD1740 | [11] | |
|
C055 Probe Info |
 |
27.28 | LDD1752 | [11] | |
|
C094 Probe Info |
 |
26.91 | LDD1785 | [11] | |
|
C228 Probe Info |
 |
14.72 | LDD1901 | [11] | |
|
C431 Probe Info |
 |
14.62 | LDD2086 | [11] | |
|
FFF probe11 Probe Info |
 |
18.92 | LDD0471 | [12] | |
|
FFF probe13 Probe Info |
 |
14.77 | LDD0475 | [12] | |
|
FFF probe14 Probe Info |
 |
12.15 | LDD0477 | [12] | |
|
FFF probe2 Probe Info |
 |
18.16 | LDD0463 | [12] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0464 | [12] | |
|
JN0003 Probe Info |
 |
20.00 | LDD0469 | [12] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [13] | |
|
OEA-DA Probe Info |
 |
7.80 | LDD0046 | [14] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | H215(0.00); H90(0.00) | LDD0222 | [10] |
| LDCM0095 | DC-LC3IN-D5 | HeLa | 1.52 | LDD0055 | [3] |
| LDCM0107 | IAA | HeLa | H90(0.00); H215(0.00) | LDD0221 | [10] |
| LDCM0109 | NEM | HeLa | H90(0.00); H215(0.00); H298(0.00) | LDD0223 | [10] |
| LDCM0016 | Ranjitkar_cp1 | MDA-MB-231 | 2.13 | LDD0123 | [4] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Synapse differentiation-inducing gene protein 1 (SYNDIG1) | CD225/Dispanin family | Q9H7V2 | |||
| Huntingtin (HTT) | Huntingtin family | P42858 | |||
Cytokine and receptor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Uteroglobin (SCGB1A1) | Secretoglobin family | P11684 | |||
Other
References
