Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| CHL1 | SNV: p.G128E | DBIA Probe Info | |||
| HEC1 | SNV: p.I282V | DBIA Probe Info | |||
| HEC1B | SNV: p.I282V | . | |||
| KELLY | SNV: p.S14I | . | |||
| MOLT4 | SNV: p.V59I | IA-alkyne Probe Info | |||
| SW1990 | SNV: p.S15W | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C-Sul Probe Info |
 |
5.25 | LDD0066 | [1] | |
|
STPyne Probe Info |
 |
K44(8.33) | LDD0277 | [2] | |
|
IPM Probe Info |
 |
N.A. | LDD0241 | [3] | |
|
Probe 1 Probe Info |
 |
Y115(17.05) | LDD3495 | [4] | |
|
DBIA Probe Info |
 |
C141(1.34) | LDD3311 | [5] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0223 | [6] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [7] | |
|
CY-1 Probe Info |
 |
E76(0.00); E77(0.00); K80(0.00) | LDD0246 | [8] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [9] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [9] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [9] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [10] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [11] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C220 Probe Info |
 |
11.79 | LDD1894 | [12] | |
|
C235 Probe Info |
 |
20.82 | LDD1908 | [12] | |
|
C243 Probe Info |
 |
10.63 | LDD1916 | [12] | |
|
C244 Probe Info |
 |
15.56 | LDD1917 | [12] | |
|
C245 Probe Info |
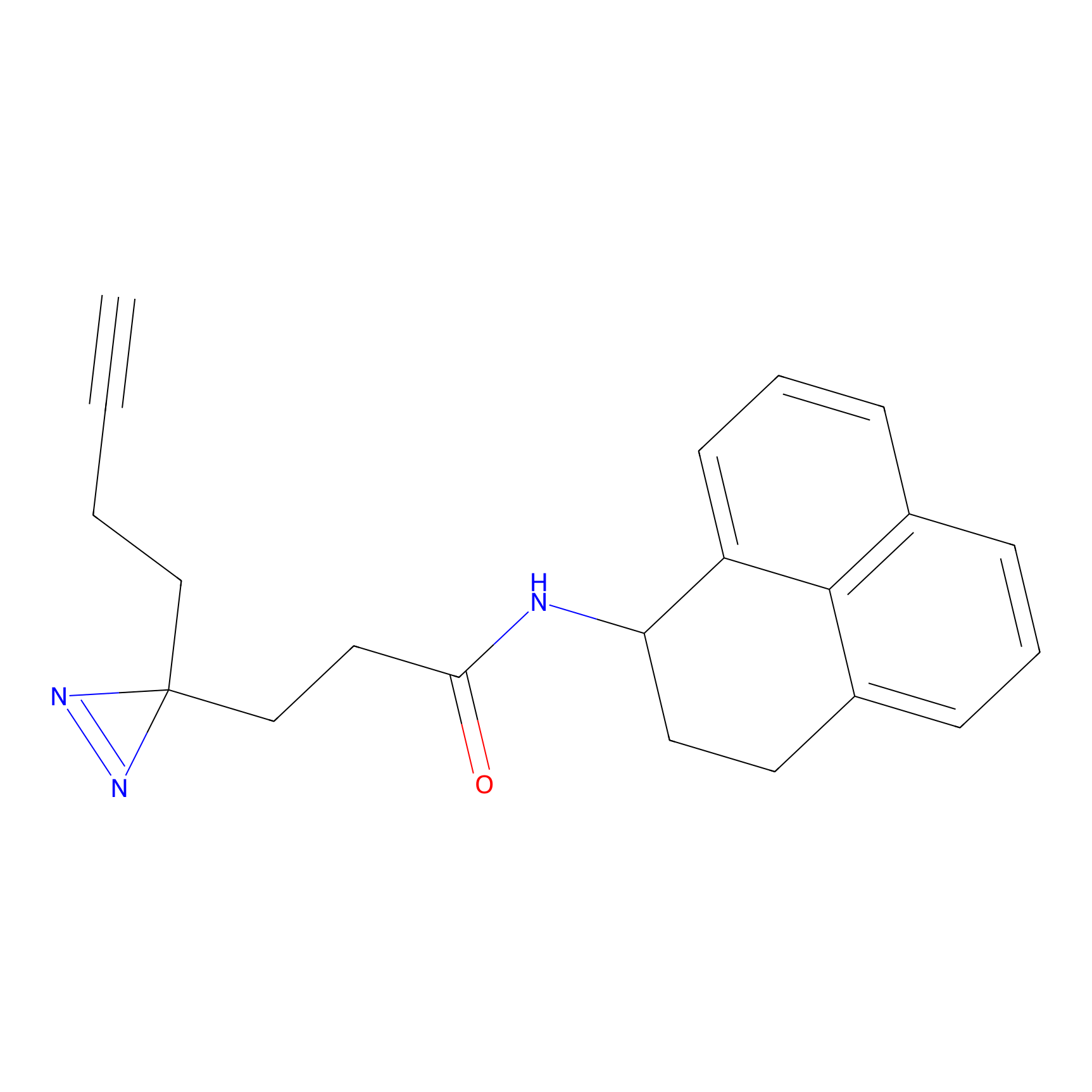 |
5.39 | LDD1918 | [12] | |
|
C246 Probe Info |
 |
15.03 | LDD1919 | [12] | |
|
C249 Probe Info |
 |
12.38 | LDD1922 | [12] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [13] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C141(1.01) | LDD0812 | [14] |
| LDCM0230 | AC113 | HEK-293T | C141(0.96) | LDD0828 | [14] |
| LDCM0231 | AC114 | HEK-293T | C141(0.92) | LDD0829 | [14] |
| LDCM0232 | AC115 | HEK-293T | C141(1.00) | LDD0830 | [14] |
| LDCM0233 | AC116 | HEK-293T | C141(1.00) | LDD0831 | [14] |
| LDCM0234 | AC117 | HEK-293T | C141(1.22) | LDD0832 | [14] |
| LDCM0235 | AC118 | HEK-293T | C141(1.01) | LDD0833 | [14] |
| LDCM0236 | AC119 | HEK-293T | C141(0.95) | LDD0834 | [14] |
| LDCM0238 | AC120 | HEK-293T | C141(1.09) | LDD0836 | [14] |
| LDCM0239 | AC121 | HEK-293T | C141(1.13) | LDD0837 | [14] |
| LDCM0240 | AC122 | HEK-293T | C141(1.05) | LDD0838 | [14] |
| LDCM0241 | AC123 | HEK-293T | C141(1.14) | LDD0839 | [14] |
| LDCM0242 | AC124 | HEK-293T | C141(0.96) | LDD0840 | [14] |
| LDCM0243 | AC125 | HEK-293T | C141(1.12) | LDD0841 | [14] |
| LDCM0244 | AC126 | HEK-293T | C141(1.07) | LDD0842 | [14] |
| LDCM0245 | AC127 | HEK-293T | C141(1.15) | LDD0843 | [14] |
| LDCM0263 | AC143 | HEK-293T | C141(0.84) | LDD0861 | [14] |
| LDCM0264 | AC144 | HEK-293T | C141(1.05) | LDD0862 | [14] |
| LDCM0265 | AC145 | HEK-293T | C141(1.04) | LDD0863 | [14] |
| LDCM0266 | AC146 | HEK-293T | C141(0.96) | LDD0864 | [14] |
| LDCM0267 | AC147 | HEK-293T | C141(1.17) | LDD0865 | [14] |
| LDCM0268 | AC148 | HEK-293T | C141(1.10) | LDD0866 | [14] |
| LDCM0269 | AC149 | HEK-293T | C141(1.16) | LDD0867 | [14] |
| LDCM0271 | AC150 | HEK-293T | C141(1.13) | LDD0869 | [14] |
| LDCM0272 | AC151 | HEK-293T | C141(0.99) | LDD0870 | [14] |
| LDCM0273 | AC152 | HEK-293T | C141(1.04) | LDD0871 | [14] |
| LDCM0274 | AC153 | HEK-293T | C141(1.14) | LDD0872 | [14] |
| LDCM0621 | AC154 | HEK-293T | C141(0.97) | LDD2162 | [14] |
| LDCM0622 | AC155 | HEK-293T | C141(0.96) | LDD2163 | [14] |
| LDCM0623 | AC156 | HEK-293T | C141(1.15) | LDD2164 | [14] |
| LDCM0624 | AC157 | HEK-293T | C141(1.42) | LDD2165 | [14] |
| LDCM0276 | AC17 | HEK-293T | C141(0.93) | LDD0874 | [14] |
| LDCM0277 | AC18 | HEK-293T | C141(0.96) | LDD0875 | [14] |
| LDCM0278 | AC19 | HEK-293T | C141(1.00) | LDD0876 | [14] |
| LDCM0279 | AC2 | HEK-293T | C141(1.11) | LDD0877 | [14] |
| LDCM0280 | AC20 | HEK-293T | C141(0.86) | LDD0878 | [14] |
| LDCM0281 | AC21 | HEK-293T | C141(0.87) | LDD0879 | [14] |
| LDCM0282 | AC22 | HEK-293T | C141(0.89) | LDD0880 | [14] |
| LDCM0283 | AC23 | HEK-293T | C141(0.84) | LDD0881 | [14] |
| LDCM0284 | AC24 | HEK-293T | C141(0.88) | LDD0882 | [14] |
| LDCM0289 | AC29 | HEK-293T | C141(1.04) | LDD1528 | [15] |
| LDCM0290 | AC3 | HEK-293T | C141(1.07) | LDD0888 | [14] |
| LDCM0298 | AC37 | HEK-293T | C141(1.18) | LDD1537 | [15] |
| LDCM0301 | AC4 | HEK-293T | C141(1.23) | LDD0899 | [14] |
| LDCM0307 | AC45 | HEK-293T | C141(1.09) | LDD1546 | [15] |
| LDCM0308 | AC46 | HEK-293T | C141(1.06) | LDD0906 | [14] |
| LDCM0309 | AC47 | HEK-293T | C141(1.04) | LDD0907 | [14] |
| LDCM0310 | AC48 | HEK-293T | C141(0.99) | LDD0908 | [14] |
| LDCM0311 | AC49 | HEK-293T | C141(0.87) | LDD0909 | [14] |
| LDCM0312 | AC5 | HEK-293T | C141(1.00) | LDD0910 | [14] |
| LDCM0313 | AC50 | HEK-293T | C141(0.91) | LDD0911 | [14] |
| LDCM0314 | AC51 | HEK-293T | C141(0.75) | LDD0912 | [14] |
| LDCM0315 | AC52 | HEK-293T | C141(0.76) | LDD0913 | [14] |
| LDCM0316 | AC53 | HEK-293T | C141(0.84) | LDD0914 | [14] |
| LDCM0317 | AC54 | HEK-293T | C141(0.93) | LDD0915 | [14] |
| LDCM0318 | AC55 | HEK-293T | C141(0.97) | LDD0916 | [14] |
| LDCM0319 | AC56 | HEK-293T | C141(1.01) | LDD0917 | [14] |
| LDCM0320 | AC57 | HEK-293T | C141(0.96) | LDD0918 | [14] |
| LDCM0321 | AC58 | HEK-293T | C141(0.87) | LDD0919 | [14] |
| LDCM0322 | AC59 | HEK-293T | C141(0.85) | LDD0920 | [14] |
| LDCM0324 | AC60 | HEK-293T | C141(0.62) | LDD0922 | [14] |
| LDCM0325 | AC61 | HEK-293T | C141(0.92) | LDD0923 | [14] |
| LDCM0326 | AC62 | HEK-293T | C141(0.96) | LDD0924 | [14] |
| LDCM0327 | AC63 | HEK-293T | C141(0.84) | LDD0925 | [14] |
| LDCM0328 | AC64 | HEK-293T | C141(0.88) | LDD0926 | [14] |
| LDCM0329 | AC65 | HEK-293T | C141(0.88) | LDD0927 | [14] |
| LDCM0330 | AC66 | HEK-293T | C141(0.92) | LDD0928 | [14] |
| LDCM0331 | AC67 | HEK-293T | C141(0.83) | LDD0929 | [14] |
| LDCM0332 | AC68 | HEK-293T | C141(1.09) | LDD0930 | [14] |
| LDCM0333 | AC69 | HEK-293T | C141(0.99) | LDD0931 | [14] |
| LDCM0335 | AC70 | HEK-293T | C141(1.09) | LDD0933 | [14] |
| LDCM0336 | AC71 | HEK-293T | C141(1.06) | LDD0934 | [14] |
| LDCM0337 | AC72 | HEK-293T | C141(1.14) | LDD0935 | [14] |
| LDCM0338 | AC73 | HEK-293T | C141(1.20) | LDD0936 | [14] |
| LDCM0339 | AC74 | HEK-293T | C141(1.19) | LDD0937 | [14] |
| LDCM0340 | AC75 | HEK-293T | C141(1.22) | LDD0938 | [14] |
| LDCM0341 | AC76 | HEK-293T | C141(1.08) | LDD0939 | [14] |
| LDCM0342 | AC77 | HEK-293T | C141(1.33) | LDD0940 | [14] |
| LDCM0343 | AC78 | HEK-293T | C141(1.07) | LDD0941 | [14] |
| LDCM0344 | AC79 | HEK-293T | C141(1.13) | LDD0942 | [14] |
| LDCM0346 | AC80 | HEK-293T | C141(1.06) | LDD0944 | [14] |
| LDCM0347 | AC81 | HEK-293T | C141(1.19) | LDD0945 | [14] |
| LDCM0348 | AC82 | HEK-293T | C141(1.20) | LDD0946 | [14] |
| LDCM0248 | AKOS034007472 | HEK-293T | C141(1.14) | LDD1511 | [15] |
| LDCM0369 | CL100 | HEK-293T | C141(1.07) | LDD0967 | [14] |
| LDCM0374 | CL105 | HEK-293T | C141(0.80) | LDD0972 | [14] |
| LDCM0375 | CL106 | HEK-293T | C141(0.95) | LDD0973 | [14] |
| LDCM0376 | CL107 | HEK-293T | C141(0.98) | LDD0974 | [14] |
| LDCM0377 | CL108 | HEK-293T | C141(0.98) | LDD0975 | [14] |
| LDCM0378 | CL109 | HEK-293T | C141(0.91) | LDD0976 | [14] |
| LDCM0380 | CL110 | HEK-293T | C141(0.91) | LDD0978 | [14] |
| LDCM0381 | CL111 | HEK-293T | C141(0.79) | LDD0979 | [14] |
| LDCM0392 | CL121 | HEK-293T | C141(0.99) | LDD0990 | [14] |
| LDCM0393 | CL122 | HEK-293T | C141(0.95) | LDD0991 | [14] |
| LDCM0394 | CL123 | HEK-293T | C141(0.92) | LDD0992 | [14] |
| LDCM0395 | CL124 | HEK-293T | C141(1.09) | LDD0993 | [14] |
| LDCM0396 | CL125 | HEK-293T | C141(0.86) | LDD0994 | [14] |
| LDCM0397 | CL126 | HEK-293T | C141(0.85) | LDD0995 | [14] |
| LDCM0398 | CL127 | HEK-293T | C141(0.85) | LDD0996 | [14] |
| LDCM0399 | CL128 | HEK-293T | C141(1.05) | LDD0997 | [14] |
| LDCM0409 | CL21 | HEK-293T | C141(1.01) | LDD1613 | [15] |
| LDCM0422 | CL33 | HEK-293T | C141(1.07) | LDD1626 | [15] |
| LDCM0435 | CL45 | HEK-293T | C141(1.00) | LDD1639 | [15] |
| LDCM0448 | CL57 | HEK-293T | C141(1.13) | LDD1651 | [15] |
| LDCM0453 | CL61 | HEK-293T | C141(1.15) | LDD1051 | [14] |
| LDCM0454 | CL62 | HEK-293T | C141(0.96) | LDD1052 | [14] |
| LDCM0455 | CL63 | HEK-293T | C141(0.79) | LDD1053 | [14] |
| LDCM0456 | CL64 | HEK-293T | C141(0.78) | LDD1054 | [14] |
| LDCM0457 | CL65 | HEK-293T | C141(0.82) | LDD1055 | [14] |
| LDCM0458 | CL66 | HEK-293T | C141(0.80) | LDD1056 | [14] |
| LDCM0459 | CL67 | HEK-293T | C141(0.97) | LDD1057 | [14] |
| LDCM0460 | CL68 | HEK-293T | C141(0.95) | LDD1058 | [14] |
| LDCM0461 | CL69 | HEK-293T | C141(0.95) | LDD1059 | [14] |
| LDCM0463 | CL70 | HEK-293T | C141(0.97) | LDD1061 | [14] |
| LDCM0464 | CL71 | HEK-293T | C141(0.95) | LDD1062 | [14] |
| LDCM0465 | CL72 | HEK-293T | C141(0.86) | LDD1063 | [14] |
| LDCM0466 | CL73 | HEK-293T | C141(0.93) | LDD1064 | [14] |
| LDCM0467 | CL74 | HEK-293T | C141(0.97) | LDD1065 | [14] |
| LDCM0475 | CL81 | HEK-293T | C141(1.48) | LDD1678 | [15] |
| LDCM0484 | CL9 | HEK-293T | C141(1.12) | LDD1687 | [15] |
| LDCM0486 | CL91 | HEK-293T | C141(0.88) | LDD1084 | [14] |
| LDCM0487 | CL92 | HEK-293T | C141(0.93) | LDD1085 | [14] |
| LDCM0488 | CL93 | HEK-293T | C141(0.90) | LDD1086 | [14] |
| LDCM0489 | CL94 | HEK-293T | C141(0.89) | LDD1087 | [14] |
| LDCM0490 | CL95 | HEK-293T | C141(0.91) | LDD1088 | [14] |
| LDCM0491 | CL96 | HEK-293T | C141(0.98) | LDD1089 | [14] |
| LDCM0492 | CL97 | HEK-293T | C141(0.96) | LDD1090 | [14] |
| LDCM0493 | CL98 | HEK-293T | C141(0.95) | LDD1091 | [14] |
| LDCM0494 | CL99 | HEK-293T | C141(1.03) | LDD1092 | [14] |
| LDCM0468 | Fragment33 | HEK-293T | C141(0.67) | LDD1066 | [14] |
| LDCM0022 | KB02 | HEK-293T | C141(0.98) | LDD1492 | [15] |
| LDCM0023 | KB03 | HEK-293T | C141(1.09) | LDD1497 | [15] |
| LDCM0024 | KB05 | G361 | C141(1.34) | LDD3311 | [5] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [6] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
GPCR
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Probable G-protein coupled receptor 152 (GPR152) | G-protein coupled receptor 1 family | Q8TDT2 | |||
Other
References
