Details of the Target
General Information of Target
| Target ID | LDTP03170 | |||||
|---|---|---|---|---|---|---|
| Target Name | Ectonucleotide pyrophosphatase/phosphodiesterase family member 1 (ENPP1) | |||||
| Gene Name | ENPP1 | |||||
| Gene ID | 5167 | |||||
| Synonyms |
M6S1; NPPS; PC1; PDNP1; Ectonucleotide pyrophosphatase/phosphodiesterase family member 1; E-NPP 1; Membrane component chromosome 6 surface marker 1; Phosphodiesterase I/nucleotide pyrophosphatase 1; Plasma-cell membrane glycoprotein PC-1) [Cleaved into: Ectonucleotide pyrophosphatase/phosphodiesterase family member 1, secreted form] [Includes: Alkaline phosphodiesterase I; EC 3.1.4.1; Nucleotide pyrophosphatase; NPPase; EC 3.6.1.9; Nucleotide diphosphatase)]
|
|||||
| 3D Structure | ||||||
| Sequence |
MERDGCAGGGSRGGEGGRAPREGPAGNGRDRGRSHAAEAPGDPQAAASLLAPMDVGEEPL
EKAARARTAKDPNTYKVLSLVLSVCVLTTILGCIFGLKPSCAKEVKSCKGRCFERTFGNC RCDAACVELGNCCLDYQETCIEPEHIWTCNKFRCGEKRLTRSLCACSDDCKDKGDCCINY SSVCQGEKSWVEEPCESINEPQCPAGFETPPTLLFSLDGFRAEYLHTWGGLLPVISKLKK CGTYTKNMRPVYPTKTFPNHYSIVTGLYPESHGIIDNKMYDPKMNASFSLKSKEKFNPEW YKGEPIWVTAKYQGLKSGTFFWPGSDVEINGIFPDIYKMYNGSVPFEERILAVLQWLQLP KDERPHFYTLYLEEPDSSGHSYGPVSSEVIKALQRVDGMVGMLMDGLKELNLHRCLNLIL ISDHGMEQGSCKKYIYLNKYLGDVKNIKVIYGPAARLRPSDVPDKYYSFNYEGIARNLSC REPNQHFKPYLKHFLPKRLHFAKSDRIEPLTFYLDPQWQLALNPSERKYCGSGFHGSDNV FSNMQALFVGYGPGFKHGIEADTFENIEVYNLMCDLLNLTPAPNNGTHGSLNHLLKNPVY TPKHPKEVHPLVQCPFTRNPRDNLGCSCNPSILPIEDFQTQFNLTVAEEKIIKHETLPYG RPRVLQKENTICLLSQHQFMSGYSQDILMPLWTSYTVDRNDSFSTEDFSNCLYQDFRIPL SPVHKCSFYKNNTKVSYGFLSPPQLNKNSSGIYSEALLTTNIVPMYQSFQVIWRYFHDTL LRKYAEERNGVNVVSGPVFDFDYDGRCDSLENLRQKRRVIRNQEILIPTHFFIVLTSCKD TSQTPLHCENLDTLAFILPHRTDNSESCVHGKHDSSWVEELLMLHRARITDVEHITGLSF YQQRKEPVSDILKLKTHLPTFSQED |
|||||
| Target Type |
Clinical trial
|
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Nucleotide pyrophosphatase/phosphodiesterase family
|
|||||
| Subcellular location |
Secreted; Cell membrane
|
|||||
| Function |
Nucleotide pyrophosphatase that generates diphosphate (PPi) and functions in bone mineralization and soft tissue calcification by regulating pyrophosphate levels. PPi inhibits bone mineralization and soft tissue calcification by binding to nascent hydroxyapatite crystals, thereby preventing further growth of these crystals. Preferentially hydrolyzes ATP, but can also hydrolyze other nucleoside 5' triphosphates such as GTP, CTP and UTP to their corresponding monophosphates with release of pyrophosphate, as well as diadenosine polyphosphates, and also 3',5'-cAMP to AMP. May also be involved in the regulation of the availability of nucleotide sugars in the endoplasmic reticulum and Golgi, and the regulation of purinergic signaling. Inhibits ectopic joint calcification and maintains articular chondrocytes by repressing hedgehog signaling; it is however unclear whether hedgehog inhibition is direct or indirect. Appears to modulate insulin sensitivity and function. Also involved in melanogenesis. Also able to hydrolyze 2',3'-cGAMP (cyclic GMP-AMP), a second messenger that activates TMEM173/STING and triggers type-I interferon production. 2',3'-cGAMP degradation takes place in the lumen or extracellular space, and not in the cytosol where it is produced; the role of 2',3'-cGAMP hydrolysis is therefore unclear. Not able to hydrolyze the 2',3'-cGAMP linkage isomer 3'-3'-cGAMP.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [1] | |
|
STPyne Probe Info |
 |
K653(4.47) | LDD0277 | [2] | |
|
IPM Probe Info |
 |
C807(0.00); C6(0.00); C868(0.00) | LDD0241 | [3] | |
|
OPA-S-S-alkyne Probe Info |
 |
K448(0.53); K465(1.52); K503(1.89) | LDD3494 | [4] | |
|
Probe 1 Probe Info |
 |
Y803(2.80) | LDD3495 | [5] | |
|
DBIA Probe Info |
 |
C6(1.55) | LDD0080 | [6] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [7] | |
|
ATP probe Probe Info |
 |
K278(0.00); K283(0.00); K653(0.00) | LDD0199 | [8] | |
|
m-APA Probe Info |
 |
H847(0.00); H894(0.00) | LDD2231 | [7] | |
|
IA-alkyne Probe Info |
 |
C868(0.00); C838(0.00); C848(0.00); C614(0.00) | LDD0165 | [9] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2224 | [10] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [11] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [12] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [13] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C027 Probe Info |
 |
11.31 | LDD1733 | [14] | |
|
C158 Probe Info |
 |
58.49 | LDD1838 | [14] | |
|
C364 Probe Info |
 |
17.15 | LDD2025 | [14] | |
|
Alk-rapa Probe Info |
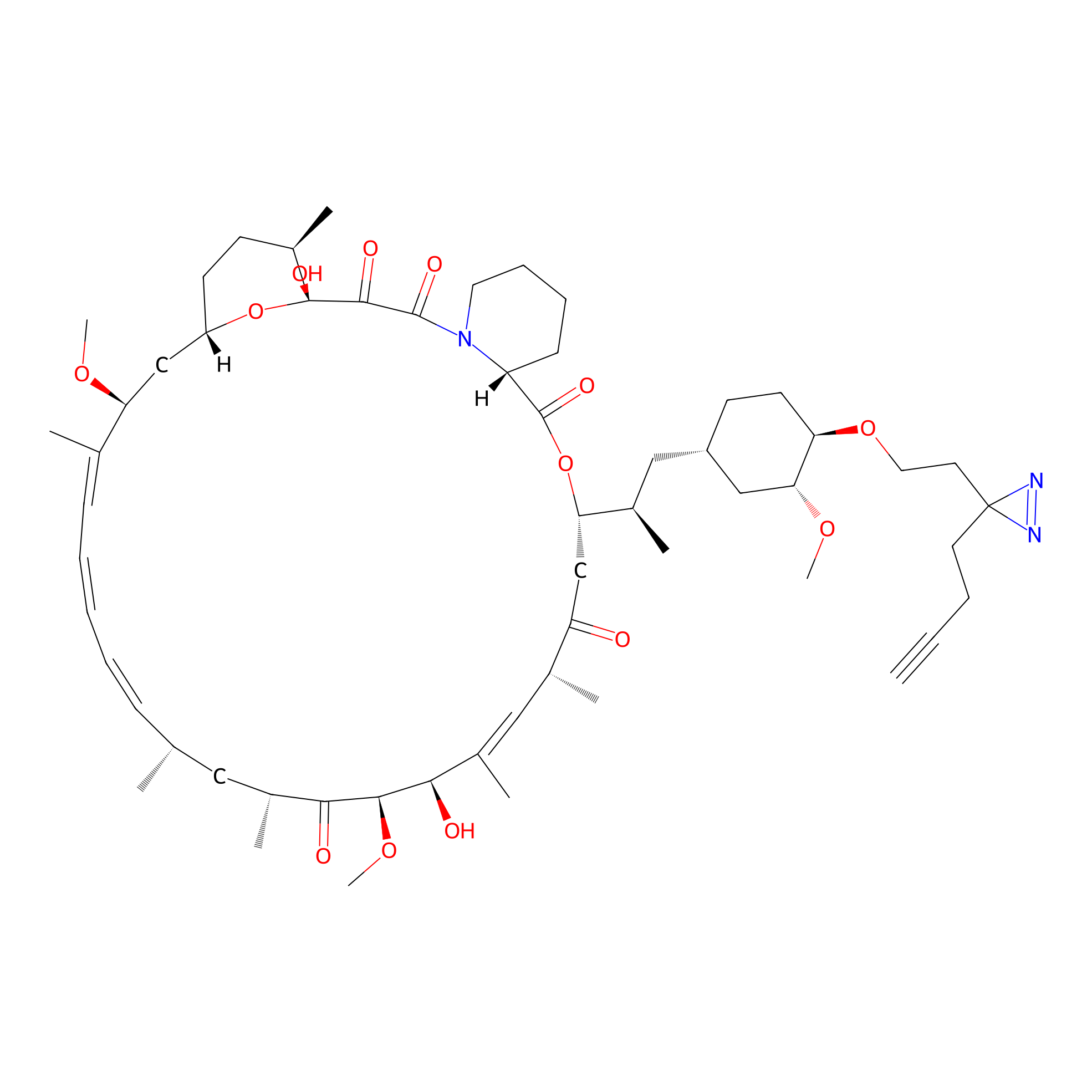 |
4.30 | LDD0213 | [15] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0215 | AC10 | HEK-293T | C6(1.06) | LDD0813 | [6] |
| LDCM0216 | AC100 | HEK-293T | C6(0.83) | LDD0814 | [6] |
| LDCM0217 | AC101 | HEK-293T | C6(0.82) | LDD0815 | [6] |
| LDCM0218 | AC102 | HEK-293T | C6(1.27) | LDD0816 | [6] |
| LDCM0219 | AC103 | HEK-293T | C6(0.61) | LDD0817 | [6] |
| LDCM0220 | AC104 | HEK-293T | C6(1.19) | LDD0818 | [6] |
| LDCM0221 | AC105 | HEK-293T | C6(0.82) | LDD0819 | [6] |
| LDCM0222 | AC106 | HEK-293T | C6(0.98) | LDD0820 | [6] |
| LDCM0223 | AC107 | HEK-293T | C6(0.94) | LDD0821 | [6] |
| LDCM0224 | AC108 | HEK-293T | C6(1.17) | LDD0822 | [6] |
| LDCM0225 | AC109 | HEK-293T | C6(0.90) | LDD0823 | [6] |
| LDCM0226 | AC11 | HEK-293T | C6(0.94) | LDD0824 | [6] |
| LDCM0227 | AC110 | HEK-293T | C6(0.86) | LDD0825 | [6] |
| LDCM0228 | AC111 | HEK-293T | C6(0.84) | LDD0826 | [6] |
| LDCM0229 | AC112 | HEK-293T | C6(0.96) | LDD0827 | [6] |
| LDCM0237 | AC12 | HEK-293T | C6(0.82) | LDD0835 | [6] |
| LDCM0259 | AC14 | HEK-293T | C6(0.87) | LDD0857 | [6] |
| LDCM0263 | AC143 | HEK-293T | C6(0.74) | LDD0861 | [6] |
| LDCM0264 | AC144 | HEK-293T | C6(0.76) | LDD0862 | [6] |
| LDCM0265 | AC145 | HEK-293T | C6(0.97) | LDD0863 | [6] |
| LDCM0266 | AC146 | HEK-293T | C6(0.89) | LDD0864 | [6] |
| LDCM0267 | AC147 | HEK-293T | C6(0.65) | LDD0865 | [6] |
| LDCM0268 | AC148 | HEK-293T | C6(0.74) | LDD0866 | [6] |
| LDCM0269 | AC149 | HEK-293T | C6(0.59) | LDD0867 | [6] |
| LDCM0270 | AC15 | HEK-293T | C6(0.94) | LDD0868 | [6] |
| LDCM0271 | AC150 | HEK-293T | C6(0.77) | LDD0869 | [6] |
| LDCM0272 | AC151 | HEK-293T | C6(0.75) | LDD0870 | [6] |
| LDCM0273 | AC152 | HEK-293T | C6(0.70) | LDD0871 | [6] |
| LDCM0274 | AC153 | HEK-293T | C6(0.53) | LDD0872 | [6] |
| LDCM0621 | AC154 | HEK-293T | C6(0.60) | LDD2162 | [6] |
| LDCM0622 | AC155 | HEK-293T | C6(0.57) | LDD2163 | [6] |
| LDCM0623 | AC156 | HEK-293T | C6(0.74) | LDD2164 | [6] |
| LDCM0624 | AC157 | HEK-293T | C6(0.59) | LDD2165 | [6] |
| LDCM0276 | AC17 | HEK-293T | C6(1.87) | LDD0874 | [6] |
| LDCM0277 | AC18 | HEK-293T | C6(1.69) | LDD0875 | [6] |
| LDCM0278 | AC19 | HEK-293T | C6(1.42) | LDD0876 | [6] |
| LDCM0280 | AC20 | HEK-293T | C6(1.06) | LDD0878 | [6] |
| LDCM0281 | AC21 | HEK-293T | C6(1.18) | LDD0879 | [6] |
| LDCM0282 | AC22 | HEK-293T | C6(0.90) | LDD0880 | [6] |
| LDCM0283 | AC23 | HEK-293T | C6(1.19) | LDD0881 | [6] |
| LDCM0284 | AC24 | HEK-293T | C6(1.35) | LDD0882 | [6] |
| LDCM0285 | AC25 | HEK-293T | C6(0.55) | LDD0883 | [6] |
| LDCM0286 | AC26 | HEK-293T | C6(1.80) | LDD0884 | [6] |
| LDCM0287 | AC27 | HEK-293T | C6(1.19) | LDD0885 | [6] |
| LDCM0288 | AC28 | HEK-293T | C6(0.73) | LDD0886 | [6] |
| LDCM0289 | AC29 | HEK-293T | C6(0.68) | LDD0887 | [6] |
| LDCM0291 | AC30 | HEK-293T | C6(0.97) | LDD0889 | [6] |
| LDCM0292 | AC31 | HEK-293T | C6(0.67) | LDD0890 | [6] |
| LDCM0293 | AC32 | HEK-293T | C6(0.97) | LDD0891 | [6] |
| LDCM0294 | AC33 | HEK-293T | C6(0.86) | LDD0892 | [6] |
| LDCM0295 | AC34 | HEK-293T | C6(0.78) | LDD0893 | [6] |
| LDCM0297 | AC36 | HEK-293T | C807(1.07) | LDD1536 | [16] |
| LDCM0298 | AC37 | HEK-293T | C807(1.13) | LDD1537 | [16] |
| LDCM0299 | AC38 | HEK-293T | C807(0.77) | LDD1538 | [16] |
| LDCM0300 | AC39 | HEK-293T | C807(1.09) | LDD1539 | [16] |
| LDCM0301 | AC4 | HEK-293T | C807(0.84) | LDD1540 | [16] |
| LDCM0302 | AC40 | HEK-293T | C807(0.93); C431(0.70) | LDD1541 | [16] |
| LDCM0306 | AC44 | HEK-293T | C807(1.09) | LDD1545 | [16] |
| LDCM0307 | AC45 | HEK-293T | C807(1.10) | LDD1546 | [16] |
| LDCM0308 | AC46 | HEK-293T | C6(0.89) | LDD0906 | [6] |
| LDCM0309 | AC47 | HEK-293T | C6(0.73) | LDD0907 | [6] |
| LDCM0310 | AC48 | HEK-293T | C6(1.69) | LDD0908 | [6] |
| LDCM0311 | AC49 | HEK-293T | C6(1.21) | LDD0909 | [6] |
| LDCM0312 | AC5 | HEK-293T | C807(1.06) | LDD1551 | [16] |
| LDCM0313 | AC50 | HEK-293T | C6(0.92) | LDD0911 | [6] |
| LDCM0314 | AC51 | HEK-293T | C6(1.09) | LDD0912 | [6] |
| LDCM0315 | AC52 | HEK-293T | C6(1.45) | LDD0913 | [6] |
| LDCM0316 | AC53 | HEK-293T | C6(0.99) | LDD0914 | [6] |
| LDCM0317 | AC54 | HEK-293T | C6(0.98) | LDD0915 | [6] |
| LDCM0318 | AC55 | HEK-293T | C6(1.46) | LDD0916 | [6] |
| LDCM0319 | AC56 | HEK-293T | C6(0.97) | LDD0917 | [6] |
| LDCM0323 | AC6 | HEK-293T | C6(0.86) | LDD0921 | [6] |
| LDCM0324 | AC60 | HEK-293T | C807(0.93) | LDD1563 | [16] |
| LDCM0325 | AC61 | HEK-293T | C807(1.08) | LDD1564 | [16] |
| LDCM0326 | AC62 | HEK-293T | C807(0.78) | LDD1565 | [16] |
| LDCM0327 | AC63 | HEK-293T | C807(0.92) | LDD1566 | [16] |
| LDCM0328 | AC64 | HEK-293T | C807(1.01); C431(0.83) | LDD1567 | [16] |
| LDCM0332 | AC68 | HEK-293T | C6(1.20) | LDD0930 | [6] |
| LDCM0333 | AC69 | HEK-293T | C6(1.30) | LDD0931 | [6] |
| LDCM0334 | AC7 | HEK-293T | C6(0.81) | LDD0932 | [6] |
| LDCM0335 | AC70 | HEK-293T | C6(1.02) | LDD0933 | [6] |
| LDCM0336 | AC71 | HEK-293T | C6(0.93) | LDD0934 | [6] |
| LDCM0337 | AC72 | HEK-293T | C6(1.07) | LDD0935 | [6] |
| LDCM0338 | AC73 | HEK-293T | C6(0.83) | LDD0936 | [6] |
| LDCM0339 | AC74 | HEK-293T | C6(0.74) | LDD0937 | [6] |
| LDCM0340 | AC75 | HEK-293T | C6(0.87) | LDD0938 | [6] |
| LDCM0341 | AC76 | HEK-293T | C6(1.16) | LDD0939 | [6] |
| LDCM0342 | AC77 | HEK-293T | C6(1.63) | LDD0940 | [6] |
| LDCM0343 | AC78 | HEK-293T | C6(1.42) | LDD0941 | [6] |
| LDCM0344 | AC79 | HEK-293T | C6(1.32) | LDD0942 | [6] |
| LDCM0345 | AC8 | HEK-293T | C6(0.69) | LDD0943 | [6] |
| LDCM0346 | AC80 | HEK-293T | C6(1.07) | LDD0944 | [6] |
| LDCM0347 | AC81 | HEK-293T | C6(1.30) | LDD0945 | [6] |
| LDCM0348 | AC82 | HEK-293T | C6(2.14) | LDD0946 | [6] |
| LDCM0365 | AC98 | HEK-293T | C6(1.03) | LDD0963 | [6] |
| LDCM0366 | AC99 | HEK-293T | C6(0.86) | LDD0964 | [6] |
| LDCM0248 | AKOS034007472 | HEK-293T | C6(0.82) | LDD0846 | [6] |
| LDCM0356 | AKOS034007680 | HEK-293T | C6(0.95) | LDD0954 | [6] |
| LDCM0275 | AKOS034007705 | HEK-293T | C6(0.99) | LDD0873 | [6] |
| LDCM0020 | ARS-1620 | HCC44 | C6(0.80) | LDD2171 | [6] |
| LDCM0367 | CL1 | HEK-293T | C6(1.17) | LDD0965 | [6] |
| LDCM0368 | CL10 | HEK-293T | C6(1.20) | LDD0966 | [6] |
| LDCM0369 | CL100 | HEK-293T | C6(0.94) | LDD1573 | [16] |
| LDCM0370 | CL101 | HEK-293T | C6(0.75) | LDD0968 | [6] |
| LDCM0371 | CL102 | HEK-293T | C6(0.95) | LDD0969 | [6] |
| LDCM0372 | CL103 | HEK-293T | C6(0.69) | LDD0970 | [6] |
| LDCM0373 | CL104 | HEK-293T | C6(0.81) | LDD0971 | [6] |
| LDCM0374 | CL105 | HEK-293T | C6(0.93) | LDD0972 | [6] |
| LDCM0375 | CL106 | HEK-293T | C6(1.24) | LDD0973 | [6] |
| LDCM0376 | CL107 | HEK-293T | C6(1.33) | LDD0974 | [6] |
| LDCM0377 | CL108 | HEK-293T | C6(1.24) | LDD0975 | [6] |
| LDCM0378 | CL109 | HEK-293T | C6(2.02) | LDD0976 | [6] |
| LDCM0379 | CL11 | HEK-293T | C6(1.23) | LDD0977 | [6] |
| LDCM0380 | CL110 | HEK-293T | C6(1.03) | LDD0978 | [6] |
| LDCM0381 | CL111 | HEK-293T | C6(1.09) | LDD0979 | [6] |
| LDCM0382 | CL112 | HEK-293T | C6(0.79) | LDD0980 | [6] |
| LDCM0383 | CL113 | HEK-293T | C6(0.95) | LDD0981 | [6] |
| LDCM0384 | CL114 | HEK-293T | C6(0.73) | LDD0982 | [6] |
| LDCM0385 | CL115 | HEK-293T | C6(1.16) | LDD0983 | [6] |
| LDCM0386 | CL116 | HEK-293T | C6(1.13) | LDD0984 | [6] |
| LDCM0387 | CL117 | HEK-293T | C807(1.19) | LDD1591 | [16] |
| LDCM0388 | CL118 | HEK-293T | C6(1.03) | LDD1592 | [16] |
| LDCM0390 | CL12 | HEK-293T | C6(1.40) | LDD0988 | [6] |
| LDCM0391 | CL120 | HEK-293T | C6(0.98) | LDD1595 | [16] |
| LDCM0392 | CL121 | HEK-293T | C6(0.96) | LDD0990 | [6] |
| LDCM0393 | CL122 | HEK-293T | C6(1.11) | LDD0991 | [6] |
| LDCM0394 | CL123 | HEK-293T | C6(1.08) | LDD0992 | [6] |
| LDCM0395 | CL124 | HEK-293T | C6(1.14) | LDD0993 | [6] |
| LDCM0396 | CL125 | HEK-293T | C807(1.18) | LDD1600 | [16] |
| LDCM0397 | CL126 | HEK-293T | C6(0.91) | LDD1601 | [16] |
| LDCM0399 | CL128 | HEK-293T | C6(0.97) | LDD1603 | [16] |
| LDCM0400 | CL13 | HEK-293T | C6(1.10) | LDD0998 | [6] |
| LDCM0401 | CL14 | HEK-293T | C6(1.29) | LDD0999 | [6] |
| LDCM0402 | CL15 | HEK-293T | C6(1.33) | LDD1000 | [6] |
| LDCM0403 | CL16 | HEK-293T | C6(1.37) | LDD1001 | [6] |
| LDCM0404 | CL17 | HEK-293T | C6(0.97) | LDD1002 | [6] |
| LDCM0405 | CL18 | HEK-293T | C6(1.30) | LDD1003 | [6] |
| LDCM0406 | CL19 | HEK-293T | C6(1.04) | LDD1004 | [6] |
| LDCM0407 | CL2 | HEK-293T | C6(1.51) | LDD1005 | [6] |
| LDCM0408 | CL20 | HEK-293T | C6(1.05) | LDD1006 | [6] |
| LDCM0409 | CL21 | HEK-293T | C6(1.60) | LDD1007 | [6] |
| LDCM0410 | CL22 | HEK-293T | C6(1.08) | LDD1008 | [6] |
| LDCM0411 | CL23 | HEK-293T | C6(0.73) | LDD1009 | [6] |
| LDCM0412 | CL24 | HEK-293T | C6(0.95) | LDD1010 | [6] |
| LDCM0413 | CL25 | HEK-293T | C6(0.92) | LDD1011 | [6] |
| LDCM0414 | CL26 | HEK-293T | C6(0.63) | LDD1012 | [6] |
| LDCM0415 | CL27 | HEK-293T | C6(1.07) | LDD1013 | [6] |
| LDCM0416 | CL28 | HEK-293T | C6(0.78) | LDD1014 | [6] |
| LDCM0417 | CL29 | HEK-293T | C6(1.07) | LDD1015 | [6] |
| LDCM0418 | CL3 | HEK-293T | C6(1.33) | LDD1016 | [6] |
| LDCM0419 | CL30 | HEK-293T | C6(0.89) | LDD1017 | [6] |
| LDCM0420 | CL31 | HEK-293T | C6(1.33) | LDD1018 | [6] |
| LDCM0421 | CL32 | HEK-293T | C6(1.62) | LDD1019 | [6] |
| LDCM0422 | CL33 | HEK-293T | C6(2.99) | LDD1020 | [6] |
| LDCM0423 | CL34 | HEK-293T | C6(1.24) | LDD1021 | [6] |
| LDCM0424 | CL35 | HEK-293T | C6(1.41) | LDD1022 | [6] |
| LDCM0425 | CL36 | HEK-293T | C6(1.42) | LDD1023 | [6] |
| LDCM0426 | CL37 | HEK-293T | C6(1.40) | LDD1024 | [6] |
| LDCM0428 | CL39 | HEK-293T | C6(1.73) | LDD1026 | [6] |
| LDCM0429 | CL4 | HEK-293T | C6(1.40) | LDD1027 | [6] |
| LDCM0430 | CL40 | HEK-293T | C6(0.97) | LDD1028 | [6] |
| LDCM0431 | CL41 | HEK-293T | C6(1.31) | LDD1029 | [6] |
| LDCM0432 | CL42 | HEK-293T | C6(1.15) | LDD1030 | [6] |
| LDCM0433 | CL43 | HEK-293T | C6(1.29) | LDD1031 | [6] |
| LDCM0434 | CL44 | HEK-293T | C6(1.27) | LDD1032 | [6] |
| LDCM0435 | CL45 | HEK-293T | C6(1.12) | LDD1033 | [6] |
| LDCM0436 | CL46 | HEK-293T | C6(0.64) | LDD1034 | [6] |
| LDCM0437 | CL47 | HEK-293T | C6(0.65) | LDD1035 | [6] |
| LDCM0438 | CL48 | HEK-293T | C6(2.31) | LDD1036 | [6] |
| LDCM0439 | CL49 | HEK-293T | C6(0.85) | LDD1037 | [6] |
| LDCM0440 | CL5 | HEK-293T | C6(1.49) | LDD1038 | [6] |
| LDCM0441 | CL50 | HEK-293T | C6(0.89) | LDD1039 | [6] |
| LDCM0442 | CL51 | HEK-293T | C6(0.93) | LDD1040 | [6] |
| LDCM0443 | CL52 | HEK-293T | C6(0.59) | LDD1041 | [6] |
| LDCM0444 | CL53 | HEK-293T | C6(0.88) | LDD1042 | [6] |
| LDCM0445 | CL54 | HEK-293T | C6(0.85) | LDD1043 | [6] |
| LDCM0446 | CL55 | HEK-293T | C6(0.45) | LDD1044 | [6] |
| LDCM0447 | CL56 | HEK-293T | C6(0.65) | LDD1045 | [6] |
| LDCM0448 | CL57 | HEK-293T | C6(1.03) | LDD1046 | [6] |
| LDCM0449 | CL58 | HEK-293T | C6(0.88) | LDD1047 | [6] |
| LDCM0450 | CL59 | HEK-293T | C6(0.75) | LDD1048 | [6] |
| LDCM0451 | CL6 | HEK-293T | C6(1.43) | LDD1049 | [6] |
| LDCM0452 | CL60 | HEK-293T | C6(0.61) | LDD1050 | [6] |
| LDCM0453 | CL61 | HEK-293T | C6(1.08) | LDD1051 | [6] |
| LDCM0454 | CL62 | HEK-293T | C6(0.69) | LDD1052 | [6] |
| LDCM0455 | CL63 | HEK-293T | C6(0.53) | LDD1053 | [6] |
| LDCM0456 | CL64 | HEK-293T | C6(1.47) | LDD1054 | [6] |
| LDCM0457 | CL65 | HEK-293T | C6(0.88) | LDD1055 | [6] |
| LDCM0458 | CL66 | HEK-293T | C6(0.54) | LDD1056 | [6] |
| LDCM0459 | CL67 | HEK-293T | C6(0.97) | LDD1057 | [6] |
| LDCM0460 | CL68 | HEK-293T | C6(0.79) | LDD1058 | [6] |
| LDCM0461 | CL69 | HEK-293T | C6(0.54) | LDD1059 | [6] |
| LDCM0462 | CL7 | HEK-293T | C6(1.14) | LDD1060 | [6] |
| LDCM0463 | CL70 | HEK-293T | C6(0.87) | LDD1061 | [6] |
| LDCM0464 | CL71 | HEK-293T | C6(0.67) | LDD1062 | [6] |
| LDCM0465 | CL72 | HEK-293T | C6(0.66) | LDD1063 | [6] |
| LDCM0466 | CL73 | HEK-293T | C6(0.88) | LDD1064 | [6] |
| LDCM0467 | CL74 | HEK-293T | C6(0.47) | LDD1065 | [6] |
| LDCM0469 | CL76 | HEK-293T | C6(0.89) | LDD1067 | [6] |
| LDCM0470 | CL77 | HEK-293T | C6(1.11) | LDD1068 | [6] |
| LDCM0471 | CL78 | HEK-293T | C6(0.78) | LDD1069 | [6] |
| LDCM0472 | CL79 | HEK-293T | C6(1.02) | LDD1070 | [6] |
| LDCM0473 | CL8 | HEK-293T | C807(0.73) | LDD1676 | [16] |
| LDCM0474 | CL80 | HEK-293T | C6(0.95) | LDD1072 | [6] |
| LDCM0475 | CL81 | HEK-293T | C6(1.06) | LDD1073 | [6] |
| LDCM0476 | CL82 | HEK-293T | C6(0.64) | LDD1074 | [6] |
| LDCM0477 | CL83 | HEK-293T | C6(0.61) | LDD1075 | [6] |
| LDCM0478 | CL84 | HEK-293T | C6(1.01) | LDD1076 | [6] |
| LDCM0479 | CL85 | HEK-293T | C6(0.83) | LDD1077 | [6] |
| LDCM0480 | CL86 | HEK-293T | C6(0.91) | LDD1078 | [6] |
| LDCM0481 | CL87 | HEK-293T | C6(1.01) | LDD1079 | [6] |
| LDCM0482 | CL88 | HEK-293T | C6(1.09) | LDD1080 | [6] |
| LDCM0483 | CL89 | HEK-293T | C6(0.90) | LDD1081 | [6] |
| LDCM0484 | CL9 | HEK-293T | C807(1.00) | LDD1687 | [16] |
| LDCM0485 | CL90 | HEK-293T | C6(0.87) | LDD1083 | [6] |
| LDCM0487 | CL92 | HEK-293T | C807(0.91) | LDD1690 | [16] |
| LDCM0488 | CL93 | HEK-293T | C807(1.10) | LDD1691 | [16] |
| LDCM0489 | CL94 | HEK-293T | C807(0.90) | LDD1692 | [16] |
| LDCM0490 | CL95 | HEK-293T | C807(0.93) | LDD1693 | [16] |
| LDCM0491 | CL96 | HEK-293T | C807(1.04); C431(0.72) | LDD1694 | [16] |
| LDCM0492 | CL97 | HEK-293T | C807(0.96) | LDD1695 | [16] |
| LDCM0493 | CL98 | HEK-293T | C6(1.00) | LDD1696 | [16] |
| LDCM0468 | Fragment33 | HEK-293T | C6(0.78) | LDD1066 | [6] |
| LDCM0427 | Fragment51 | HEK-293T | C6(1.72) | LDD1025 | [6] |
| LDCM0022 | KB02 | HCT 116 | C6(1.55) | LDD0080 | [6] |
| LDCM0023 | KB03 | HCT 116 | C6(1.19) | LDD0081 | [6] |
| LDCM0024 | KB05 | HCT 116 | C6(1.46) | LDD0082 | [6] |
| LDCM0090 | Rapamycin | JHH-7 | 4.30 | LDD0213 | [15] |
| LDCM0021 | THZ1 | HCT 116 | C6(0.80) | LDD2173 | [6] |
The Interaction Atlas With This Target
References
