Details of the Target
General Information of Target
| Target ID | LDTP00558 | |||||
|---|---|---|---|---|---|---|
| Target Name | Phospholipid scramblase 1 (PLSCR1) | |||||
| Gene Name | PLSCR1 | |||||
| Gene ID | 5359 | |||||
| Synonyms |
Phospholipid scramblase 1; PL scramblase 1; Ca(2+)-dependent phospholipid scramblase 1; Erythrocyte phospholipid scramblase; Mg(2+)-dependent nuclease; EC 3.1.-.-; MmTRA1b |
|||||
| 3D Structure | ||||||
| Sequence |
MDKQNSQMNASHPETNLPVGYPPQYPPTAFQGPPGYSGYPGPQVSYPPPPAGHSGPGPAG
FPVPNQPVYNQPVYNQPVGAAGVPWMPAPQPPLNCPPGLEYLSQIDQILIHQQIELLEVL TGFETNNKYEIKNSFGQRVYFAAEDTDCCTRNCCGPSRPFTLRIIDNMGQEVITLERPLR CSSCCCPCCLQEIEIQAPPGVPIGYVIQTWHPCLPKFTIQNEKREDVLKISGPCVVCSCC GDVDFEIKSLDEQCVVGKISKHWTGILREAFTDADNFGIQFPLDLDVKMKAVMIGACFLI DFMFFESTGSQEQKSGVW |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Phospholipid scramblase family
|
|||||
| Subcellular location |
Cell membrane
|
|||||
| Function |
Catalyzes calcium-induced ATP-independent rapid bidirectional and non-specific movement of phospholipids (lipid scrambling or lipid flip-flop) between the inner and outer leaflet of the plasma membrane resulting in collapse of the phospholipid asymmetry which leads to phosphatidylserine externalization on the cell surface. Mediates calcium-dependent phosphatidylserine externalization and apoptosis in neurons via its association with TRPC5. Also exhibits magnesium-dependent nuclease activity against double-stranded DNA and RNA but not single-stranded DNA and can enhance DNA decatenation mediated by TOP2A. Negatively regulates FcR-mediated phagocytosis in differentiated macrophages. May contribute to cytokine-regulated cell proliferation and differentiation. May play a role in the antiviral response of interferon (IFN) by amplifying and enhancing the IFN response through increased expression of select subset of potent antiviral genes. Inhibits the functions of viral transactivators, including human T-cell leukemia virus (HTLV)-1 protein Tax, human immunodeficiency virus (HIV)-1 Tat, human hepatitis B virus (HBV) HBx, Epstein-Barr virus (EBV) BZLF1 and human cytomegalovirus IE1 and IE2 proteins through direct interactions. Mediates also the inhibition of influenza virus infection by preventing nuclear import of the viral nucleoprotein/NP. Plays a crucial role as a defense factor against SARS-CoV-2 independently of its scramblase activity by directly targeting nascent viral vesicles to prevent virus-membrane fusion and the release of viral RNA into the host-cell cytosol.; (Microbial infection) Acts as an attachment receptor for HCV.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| CCK81 | SNV: p.F217L | DBIA Probe Info | |||
| DU145 | SNV: p.A29V | . | |||
| HG3 | SNV: p.Y129C | . | |||
| HT115 | Insertion: p.T161YfsTer6 | DBIA Probe Info | |||
| MCC26 | SNV: p.L175V | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
IPM Probe Info |
 |
N.A. | LDD0241 | [1] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [2] | |
|
THZ1-DTB Probe Info |
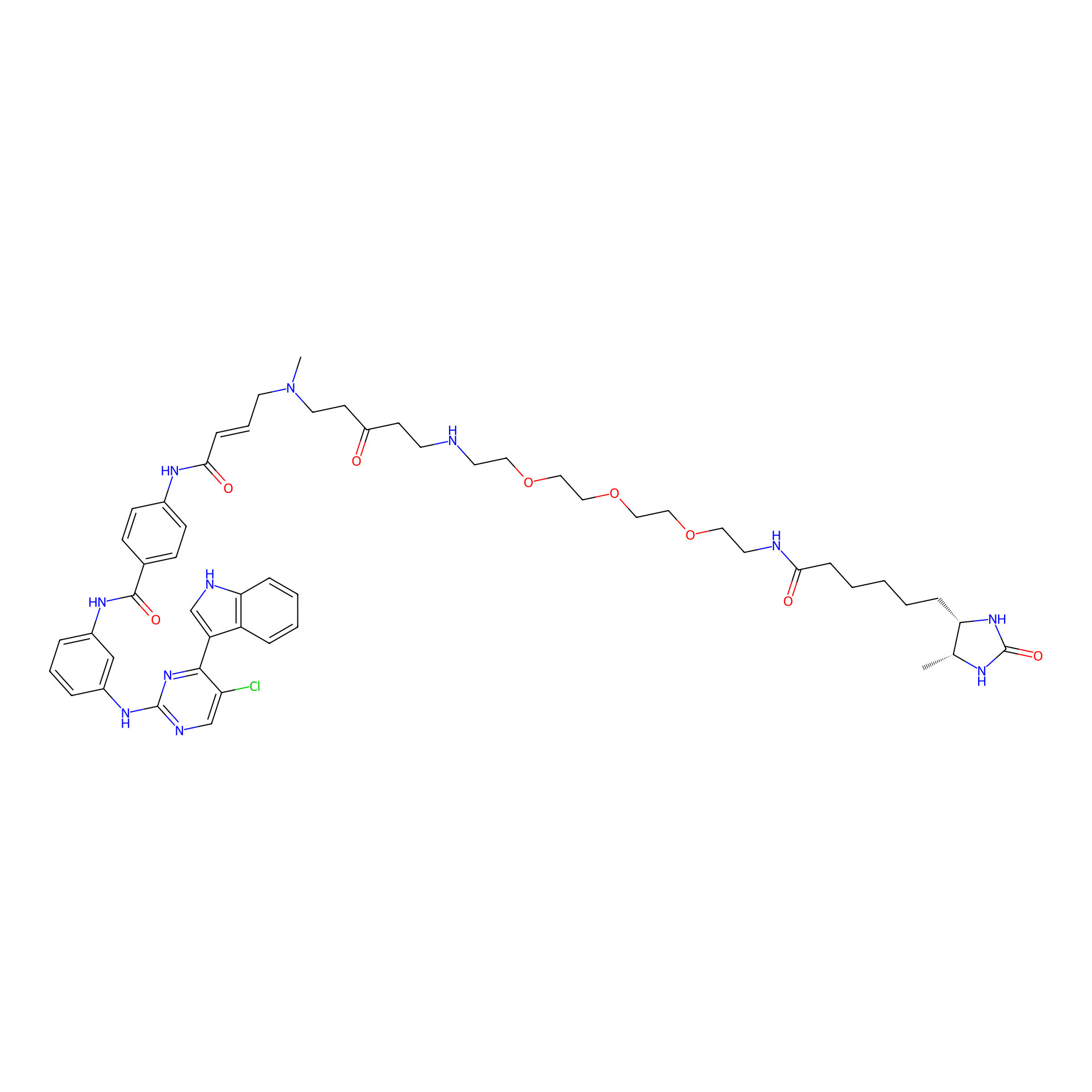 |
C148(1.05) | LDD0460 | [2] | |
|
BTD Probe Info |
 |
C254(1.52) | LDD2090 | [3] | |
|
DBIA Probe Info |
 |
C254(1.11) | LDD0080 | [4] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [5] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [5] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [5] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [6] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [6] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0017 | [7] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [8] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C254(0.58) | LDD2142 | [3] |
| LDCM0214 | AC1 | HEK-293T | C254(0.92); C234(0.95) | LDD1507 | [9] |
| LDCM0215 | AC10 | HEK-293T | C254(0.91); C148(1.03) | LDD1508 | [9] |
| LDCM0226 | AC11 | HEK-293T | C254(0.94) | LDD1509 | [9] |
| LDCM0237 | AC12 | HEK-293T | C254(0.89) | LDD1510 | [9] |
| LDCM0259 | AC14 | HEK-293T | C254(0.92) | LDD1512 | [9] |
| LDCM0270 | AC15 | HEK-293T | C254(0.73); C234(1.11); C148(1.16) | LDD1513 | [9] |
| LDCM0276 | AC17 | HEK-293T | C254(1.03); C234(0.94) | LDD1515 | [9] |
| LDCM0277 | AC18 | HEK-293T | C254(0.98); C148(1.00) | LDD1516 | [9] |
| LDCM0278 | AC19 | HEK-293T | C254(0.82) | LDD1517 | [9] |
| LDCM0279 | AC2 | HEK-293T | C254(0.96); C148(0.60) | LDD1518 | [9] |
| LDCM0280 | AC20 | HEK-293T | C254(0.98) | LDD1519 | [9] |
| LDCM0281 | AC21 | HEK-293T | C254(1.21) | LDD1520 | [9] |
| LDCM0282 | AC22 | HEK-293T | C254(0.94) | LDD1521 | [9] |
| LDCM0283 | AC23 | HEK-293T | C254(0.98); C234(1.00); C148(0.83) | LDD1522 | [9] |
| LDCM0284 | AC24 | HEK-293T | C254(1.07) | LDD1523 | [9] |
| LDCM0285 | AC25 | HEK-293T | C254(0.97); C234(1.00) | LDD1524 | [9] |
| LDCM0286 | AC26 | HEK-293T | C254(0.98); C148(1.31) | LDD1525 | [9] |
| LDCM0287 | AC27 | HEK-293T | C254(0.97) | LDD1526 | [9] |
| LDCM0288 | AC28 | HEK-293T | C254(0.93) | LDD1527 | [9] |
| LDCM0289 | AC29 | HEK-293T | C254(1.12) | LDD1528 | [9] |
| LDCM0290 | AC3 | HEK-293T | C254(0.84) | LDD1529 | [9] |
| LDCM0291 | AC30 | HEK-293T | C254(1.04) | LDD1530 | [9] |
| LDCM0292 | AC31 | HEK-293T | C254(0.94); C234(0.97); C148(0.89) | LDD1531 | [9] |
| LDCM0293 | AC32 | HEK-293T | C254(0.95) | LDD1532 | [9] |
| LDCM0294 | AC33 | HEK-293T | C254(0.92); C234(0.97) | LDD1533 | [9] |
| LDCM0295 | AC34 | HEK-293T | C254(0.91); C148(1.07) | LDD1534 | [9] |
| LDCM0296 | AC35 | HEK-293T | C254(0.91) | LDD1535 | [9] |
| LDCM0297 | AC36 | HEK-293T | C254(0.91) | LDD1536 | [9] |
| LDCM0298 | AC37 | HEK-293T | C254(0.99) | LDD1537 | [9] |
| LDCM0299 | AC38 | HEK-293T | C254(0.96) | LDD1538 | [9] |
| LDCM0300 | AC39 | HEK-293T | C254(0.77); C234(0.92); C148(1.10) | LDD1539 | [9] |
| LDCM0301 | AC4 | HEK-293T | C254(0.90) | LDD1540 | [9] |
| LDCM0302 | AC40 | HEK-293T | C254(0.91) | LDD1541 | [9] |
| LDCM0303 | AC41 | HEK-293T | C254(0.89); C234(1.16) | LDD1542 | [9] |
| LDCM0304 | AC42 | HEK-293T | C254(0.89); C148(1.05) | LDD1543 | [9] |
| LDCM0305 | AC43 | HEK-293T | C254(0.84) | LDD1544 | [9] |
| LDCM0306 | AC44 | HEK-293T | C254(0.92) | LDD1545 | [9] |
| LDCM0307 | AC45 | HEK-293T | C254(1.00) | LDD1546 | [9] |
| LDCM0308 | AC46 | HEK-293T | C254(0.96) | LDD1547 | [9] |
| LDCM0309 | AC47 | HEK-293T | C254(0.79); C234(1.04); C148(1.08) | LDD1548 | [9] |
| LDCM0310 | AC48 | HEK-293T | C254(0.92) | LDD1549 | [9] |
| LDCM0311 | AC49 | HEK-293T | C254(0.92); C234(0.91) | LDD1550 | [9] |
| LDCM0312 | AC5 | HEK-293T | C254(1.12) | LDD1551 | [9] |
| LDCM0313 | AC50 | HEK-293T | C254(1.01); C148(0.85) | LDD1552 | [9] |
| LDCM0314 | AC51 | HEK-293T | C254(0.98) | LDD1553 | [9] |
| LDCM0315 | AC52 | HEK-293T | C254(0.96) | LDD1554 | [9] |
| LDCM0316 | AC53 | HEK-293T | C254(1.07) | LDD1555 | [9] |
| LDCM0317 | AC54 | HEK-293T | C254(0.92) | LDD1556 | [9] |
| LDCM0318 | AC55 | HEK-293T | C254(0.79); C234(1.04); C148(1.25) | LDD1557 | [9] |
| LDCM0319 | AC56 | HEK-293T | C254(0.93) | LDD1558 | [9] |
| LDCM0320 | AC57 | HEK-293T | C254(0.95); C234(0.88) | LDD1559 | [9] |
| LDCM0321 | AC58 | HEK-293T | C254(1.03); C148(0.84) | LDD1560 | [9] |
| LDCM0322 | AC59 | HEK-293T | C254(0.94) | LDD1561 | [9] |
| LDCM0323 | AC6 | HEK-293T | C254(0.94) | LDD1562 | [9] |
| LDCM0324 | AC60 | HEK-293T | C254(0.94) | LDD1563 | [9] |
| LDCM0325 | AC61 | HEK-293T | C254(1.03) | LDD1564 | [9] |
| LDCM0326 | AC62 | HEK-293T | C254(0.94) | LDD1565 | [9] |
| LDCM0327 | AC63 | HEK-293T | C254(0.90); C234(1.05); C148(1.08) | LDD1566 | [9] |
| LDCM0328 | AC64 | HEK-293T | C254(0.95) | LDD1567 | [9] |
| LDCM0334 | AC7 | HEK-293T | C254(1.09); C234(0.98); C148(0.88) | LDD1568 | [9] |
| LDCM0345 | AC8 | HEK-293T | C254(0.90) | LDD1569 | [9] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C254(0.98) | LDD2113 | [3] |
| LDCM0248 | AKOS034007472 | HEK-293T | C254(1.06) | LDD1511 | [9] |
| LDCM0356 | AKOS034007680 | HEK-293T | C254(0.90); C234(0.98) | LDD1570 | [9] |
| LDCM0275 | AKOS034007705 | HEK-293T | C254(1.03) | LDD1514 | [9] |
| LDCM0367 | CL1 | HEK-293T | C254(1.01) | LDD1571 | [9] |
| LDCM0368 | CL10 | HEK-293T | C254(0.94) | LDD1572 | [9] |
| LDCM0369 | CL100 | HEK-293T | C254(0.97) | LDD1573 | [9] |
| LDCM0370 | CL101 | HEK-293T | C254(0.98) | LDD1574 | [9] |
| LDCM0371 | CL102 | HEK-293T | C254(0.86) | LDD1575 | [9] |
| LDCM0372 | CL103 | HEK-293T | C254(0.96) | LDD1576 | [9] |
| LDCM0373 | CL104 | HEK-293T | C254(0.91) | LDD1577 | [9] |
| LDCM0374 | CL105 | HEK-293T | C254(0.97) | LDD1578 | [9] |
| LDCM0375 | CL106 | HEK-293T | C254(0.95) | LDD1579 | [9] |
| LDCM0376 | CL107 | HEK-293T | C254(1.01) | LDD1580 | [9] |
| LDCM0377 | CL108 | HEK-293T | C254(0.90) | LDD1581 | [9] |
| LDCM0378 | CL109 | HEK-293T | C254(0.95) | LDD1582 | [9] |
| LDCM0379 | CL11 | HEK-293T | C254(0.80); C234(0.99); C148(1.72) | LDD1583 | [9] |
| LDCM0380 | CL110 | HEK-293T | C254(0.88) | LDD1584 | [9] |
| LDCM0381 | CL111 | HEK-293T | C254(0.98) | LDD1585 | [9] |
| LDCM0382 | CL112 | HEK-293T | C254(0.90) | LDD1586 | [9] |
| LDCM0383 | CL113 | HEK-293T | C254(1.07) | LDD1587 | [9] |
| LDCM0384 | CL114 | HEK-293T | C254(0.84) | LDD1588 | [9] |
| LDCM0385 | CL115 | HEK-293T | C254(1.02) | LDD1589 | [9] |
| LDCM0386 | CL116 | HEK-293T | C254(0.96) | LDD1590 | [9] |
| LDCM0387 | CL117 | HEK-293T | C254(0.99) | LDD1591 | [9] |
| LDCM0388 | CL118 | HEK-293T | C254(0.96) | LDD1592 | [9] |
| LDCM0389 | CL119 | HEK-293T | C254(1.02) | LDD1593 | [9] |
| LDCM0390 | CL12 | HEK-293T | C254(0.89) | LDD1594 | [9] |
| LDCM0391 | CL120 | HEK-293T | C254(0.96) | LDD1595 | [9] |
| LDCM0392 | CL121 | HEK-293T | C254(0.94) | LDD1596 | [9] |
| LDCM0393 | CL122 | HEK-293T | C254(0.98) | LDD1597 | [9] |
| LDCM0394 | CL123 | HEK-293T | C254(0.93) | LDD1598 | [9] |
| LDCM0395 | CL124 | HEK-293T | C254(0.89) | LDD1599 | [9] |
| LDCM0396 | CL125 | HEK-293T | C254(1.01) | LDD1600 | [9] |
| LDCM0397 | CL126 | HEK-293T | C254(0.95) | LDD1601 | [9] |
| LDCM0398 | CL127 | HEK-293T | C254(1.02) | LDD1602 | [9] |
| LDCM0399 | CL128 | HEK-293T | C254(0.98) | LDD1603 | [9] |
| LDCM0400 | CL13 | HEK-293T | C254(0.98) | LDD1604 | [9] |
| LDCM0401 | CL14 | HEK-293T | C254(0.93) | LDD1605 | [9] |
| LDCM0402 | CL15 | HEK-293T | C254(0.90) | LDD1606 | [9] |
| LDCM0403 | CL16 | HEK-293T | C254(0.98) | LDD1607 | [9] |
| LDCM0404 | CL17 | HEK-293T | C254(0.88); C234(1.17) | LDD1608 | [9] |
| LDCM0405 | CL18 | HEK-293T | C254(0.89); C148(1.14) | LDD1609 | [9] |
| LDCM0406 | CL19 | HEK-293T | C254(0.91) | LDD1610 | [9] |
| LDCM0407 | CL2 | HEK-293T | C254(0.86) | LDD1611 | [9] |
| LDCM0408 | CL20 | HEK-293T | C254(0.96) | LDD1612 | [9] |
| LDCM0409 | CL21 | HEK-293T | C254(0.98) | LDD1613 | [9] |
| LDCM0410 | CL22 | HEK-293T | C254(1.02) | LDD1614 | [9] |
| LDCM0411 | CL23 | HEK-293T | C254(1.02); C234(1.12); C148(0.82) | LDD1615 | [9] |
| LDCM0412 | CL24 | HEK-293T | C254(1.12) | LDD1616 | [9] |
| LDCM0413 | CL25 | HEK-293T | C254(1.15) | LDD1617 | [9] |
| LDCM0414 | CL26 | HEK-293T | C254(0.92) | LDD1618 | [9] |
| LDCM0415 | CL27 | HEK-293T | C254(0.95) | LDD1619 | [9] |
| LDCM0416 | CL28 | HEK-293T | C254(1.04) | LDD1620 | [9] |
| LDCM0417 | CL29 | HEK-293T | C254(0.93); C234(1.15) | LDD1621 | [9] |
| LDCM0418 | CL3 | HEK-293T | C254(1.03) | LDD1622 | [9] |
| LDCM0419 | CL30 | HEK-293T | C254(1.00); C148(0.91) | LDD1623 | [9] |
| LDCM0420 | CL31 | HEK-293T | C254(0.75) | LDD1624 | [9] |
| LDCM0421 | CL32 | HEK-293T | C254(0.86) | LDD1625 | [9] |
| LDCM0422 | CL33 | HEK-293T | C254(0.89) | LDD1626 | [9] |
| LDCM0423 | CL34 | HEK-293T | C254(0.98) | LDD1627 | [9] |
| LDCM0424 | CL35 | HEK-293T | C254(1.44); C234(0.99); C148(0.83) | LDD1628 | [9] |
| LDCM0425 | CL36 | HEK-293T | C254(1.05) | LDD1629 | [9] |
| LDCM0426 | CL37 | HEK-293T | C254(1.02) | LDD1630 | [9] |
| LDCM0428 | CL39 | HEK-293T | C254(0.87) | LDD1632 | [9] |
| LDCM0429 | CL4 | HEK-293T | C254(0.88) | LDD1633 | [9] |
| LDCM0430 | CL40 | HEK-293T | C254(1.00) | LDD1634 | [9] |
| LDCM0431 | CL41 | HEK-293T | C254(0.91); C234(1.10) | LDD1635 | [9] |
| LDCM0432 | CL42 | HEK-293T | C254(1.02); C148(1.21) | LDD1636 | [9] |
| LDCM0433 | CL43 | HEK-293T | C254(0.86) | LDD1637 | [9] |
| LDCM0434 | CL44 | HEK-293T | C254(0.90) | LDD1638 | [9] |
| LDCM0435 | CL45 | HEK-293T | C254(0.90) | LDD1639 | [9] |
| LDCM0436 | CL46 | HEK-293T | C254(1.02) | LDD1640 | [9] |
| LDCM0437 | CL47 | HEK-293T | C254(1.36); C234(1.05); C148(1.31) | LDD1641 | [9] |
| LDCM0438 | CL48 | HEK-293T | C254(1.02) | LDD1642 | [9] |
| LDCM0439 | CL49 | HEK-293T | C254(0.97) | LDD1643 | [9] |
| LDCM0440 | CL5 | HEK-293T | C254(0.96); C234(1.24) | LDD1644 | [9] |
| LDCM0441 | CL50 | HEK-293T | C254(0.90) | LDD1645 | [9] |
| LDCM0443 | CL52 | HEK-293T | C254(0.92) | LDD1646 | [9] |
| LDCM0444 | CL53 | HEK-293T | C254(0.88); C234(1.30) | LDD1647 | [9] |
| LDCM0445 | CL54 | HEK-293T | C254(0.82); C148(1.26) | LDD1648 | [9] |
| LDCM0446 | CL55 | HEK-293T | C254(0.83) | LDD1649 | [9] |
| LDCM0447 | CL56 | HEK-293T | C254(0.87) | LDD1650 | [9] |
| LDCM0448 | CL57 | HEK-293T | C254(0.94) | LDD1651 | [9] |
| LDCM0449 | CL58 | HEK-293T | C254(0.94) | LDD1652 | [9] |
| LDCM0450 | CL59 | HEK-293T | C254(0.81); C234(1.10); C148(0.81) | LDD1653 | [9] |
| LDCM0451 | CL6 | HEK-293T | C254(0.86); C148(1.44) | LDD1654 | [9] |
| LDCM0452 | CL60 | HEK-293T | C254(0.97) | LDD1655 | [9] |
| LDCM0453 | CL61 | HEK-293T | C254(1.00) | LDD1656 | [9] |
| LDCM0454 | CL62 | HEK-293T | C254(0.92) | LDD1657 | [9] |
| LDCM0455 | CL63 | HEK-293T | C254(0.97) | LDD1658 | [9] |
| LDCM0456 | CL64 | HEK-293T | C254(0.84) | LDD1659 | [9] |
| LDCM0457 | CL65 | HEK-293T | C254(0.93); C234(1.22) | LDD1660 | [9] |
| LDCM0458 | CL66 | HEK-293T | C254(0.92); C148(1.30) | LDD1661 | [9] |
| LDCM0459 | CL67 | HEK-293T | C254(0.73) | LDD1662 | [9] |
| LDCM0460 | CL68 | HEK-293T | C254(0.89) | LDD1663 | [9] |
| LDCM0461 | CL69 | HEK-293T | C254(1.08) | LDD1664 | [9] |
| LDCM0462 | CL7 | HEK-293T | C254(0.88) | LDD1665 | [9] |
| LDCM0463 | CL70 | HEK-293T | C254(0.94) | LDD1666 | [9] |
| LDCM0464 | CL71 | HEK-293T | C254(0.94); C234(1.12); C148(1.12) | LDD1667 | [9] |
| LDCM0465 | CL72 | HEK-293T | C254(1.13) | LDD1668 | [9] |
| LDCM0466 | CL73 | HEK-293T | C254(0.94) | LDD1669 | [9] |
| LDCM0467 | CL74 | HEK-293T | C254(0.96) | LDD1670 | [9] |
| LDCM0469 | CL76 | HEK-293T | C254(0.89) | LDD1672 | [9] |
| LDCM0470 | CL77 | HEK-293T | C254(0.82); C234(1.34) | LDD1673 | [9] |
| LDCM0471 | CL78 | HEK-293T | C254(0.92); C148(0.84) | LDD1674 | [9] |
| LDCM0472 | CL79 | HEK-293T | C254(0.98) | LDD1675 | [9] |
| LDCM0473 | CL8 | HEK-293T | C254(0.82) | LDD1676 | [9] |
| LDCM0474 | CL80 | HEK-293T | C254(0.92) | LDD1677 | [9] |
| LDCM0475 | CL81 | HEK-293T | C254(0.93) | LDD1678 | [9] |
| LDCM0476 | CL82 | HEK-293T | C254(0.96) | LDD1679 | [9] |
| LDCM0477 | CL83 | HEK-293T | C254(0.83); C234(0.97); C148(1.46) | LDD1680 | [9] |
| LDCM0478 | CL84 | HEK-293T | C254(0.90) | LDD1681 | [9] |
| LDCM0479 | CL85 | HEK-293T | C254(1.01) | LDD1682 | [9] |
| LDCM0480 | CL86 | HEK-293T | C254(0.98) | LDD1683 | [9] |
| LDCM0481 | CL87 | HEK-293T | C254(1.06) | LDD1684 | [9] |
| LDCM0482 | CL88 | HEK-293T | C254(0.88) | LDD1685 | [9] |
| LDCM0483 | CL89 | HEK-293T | C254(0.97); C234(0.95) | LDD1686 | [9] |
| LDCM0484 | CL9 | HEK-293T | C254(1.00) | LDD1687 | [9] |
| LDCM0485 | CL90 | HEK-293T | C254(0.79); C148(0.81) | LDD1688 | [9] |
| LDCM0486 | CL91 | HEK-293T | C254(0.95) | LDD1689 | [9] |
| LDCM0487 | CL92 | HEK-293T | C254(0.86) | LDD1690 | [9] |
| LDCM0488 | CL93 | HEK-293T | C254(0.92) | LDD1691 | [9] |
| LDCM0489 | CL94 | HEK-293T | C254(0.91) | LDD1692 | [9] |
| LDCM0490 | CL95 | HEK-293T | C254(0.58); C234(1.27); C148(1.46) | LDD1693 | [9] |
| LDCM0491 | CL96 | HEK-293T | C254(1.06) | LDD1694 | [9] |
| LDCM0492 | CL97 | HEK-293T | C254(0.95) | LDD1695 | [9] |
| LDCM0493 | CL98 | HEK-293T | C254(0.91) | LDD1696 | [9] |
| LDCM0494 | CL99 | HEK-293T | C254(0.92) | LDD1697 | [9] |
| LDCM0495 | E2913 | HEK-293T | C254(0.92) | LDD1698 | [9] |
| LDCM0468 | Fragment33 | HEK-293T | C254(0.98) | LDD1671 | [9] |
| LDCM0427 | Fragment51 | HEK-293T | C254(0.98) | LDD1631 | [9] |
| LDCM0179 | JZ128 | PC-3 | N.A. | LDD0462 | [2] |
| LDCM0022 | KB02 | HCT 116 | C254(1.11) | LDD0080 | [4] |
| LDCM0023 | KB03 | HCT 116 | C254(1.29) | LDD0081 | [4] |
| LDCM0024 | KB05 | HCT 116 | C254(1.30) | LDD0082 | [4] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C254(1.52) | LDD2090 | [3] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C254(0.92) | LDD2092 | [3] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C254(0.90) | LDD2093 | [3] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C254(0.65) | LDD2097 | [3] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C254(0.86) | LDD2099 | [3] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C254(0.45) | LDD2100 | [3] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C254(0.61) | LDD2101 | [3] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C254(0.54) | LDD2104 | [3] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C254(1.04) | LDD2105 | [3] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C254(0.34) | LDD2106 | [3] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C254(0.99) | LDD2107 | [3] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C254(0.70) | LDD2108 | [3] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C254(1.00) | LDD2111 | [3] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C254(0.36) | LDD2114 | [3] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C254(0.61) | LDD2115 | [3] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C254(1.18) | LDD2119 | [3] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C254(1.01) | LDD2125 | [3] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C254(0.60) | LDD2127 | [3] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C254(0.93) | LDD2129 | [3] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C254(0.86) | LDD2133 | [3] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C254(1.14) | LDD2135 | [3] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C254(0.75) | LDD2136 | [3] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C254(1.10) | LDD2137 | [3] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C254(0.75) | LDD2140 | [3] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C254(0.60) | LDD2141 | [3] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C254(1.49) | LDD2144 | [3] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C254(1.37) | LDD2145 | [3] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C254(1.05) | LDD2153 | [3] |
| LDCM0021 | THZ1 | HeLa S3 | C148(1.05) | LDD0460 | [2] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
GPCR
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| P2Y purinoceptor 6 (P2RY6) | G-protein coupled receptor 1 family | Q15077 | |||
Other
References
